Proteomic Analysis of the Rad18 Interaction Network in DT40 – a Chicken B Cell Line
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ubiquitination Is Not Omnipresent in Myeloid Leukemia Ramesh C
Editorials Ubiquitination is not omnipresent in myeloid leukemia Ramesh C. Nayak1 and Jose A. Cancelas1,2 1Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital Medical Center and 2Hoxworth Blood Center, University of Cincinnati Academic Health Center, Cincinnati, OH, USA E-mail: JOSE A. CANCELAS - [email protected] / [email protected] doi:10.3324/haematol.2019.224162 hronic myelogenous leukemia (CML) is a clonal tination of target proteins through their cognate E3 ubiq- biphasic hematopoietic disorder most frequently uitin ligases belonging to three different families (RING, Ccaused by the expression of the BCR-ABL fusion HERCT, RING-between-RING or RBR type E3).7 protein. The expression of BCR-ABL fusion protein with The ubiquitin conjugating enzymes including UBE2N constitutive and elevated tyrosine kinase activity is suffi- (UBC13) and UBE2C are over-expressed in a myriad of cient to induce transformation of hematopoietic stem tumors such as breast, pancreas, colon, prostate, lym- cells (HSC) and the development of CML.1 Despite the phoma, and ovarian carcinomas.8 Higher expression of introduction of tyrosine kinase inhibitors (TKI), the dis- UBE2A is associated with poor prognosis of hepatocellu- ease may progress from a manageable chronic phase to a lar cancer.9 In leukemia, bone marrow (BM) cells from clinically challenging blast crisis phase with a poor prog- pediatric acute lymphoblastic patients show higher levels nosis,2 in which myeloid or lymphoid blasts fail to differ- of UBE2Q2 -

The Regulation of Carbamoyl Phosphate Synthetase-Aspartate Transcarbamoylase-Dihydroorotase (Cad) by Phosphorylation and Protein-Protein Interactions
THE REGULATION OF CARBAMOYL PHOSPHATE SYNTHETASE-ASPARTATE TRANSCARBAMOYLASE-DIHYDROOROTASE (CAD) BY PHOSPHORYLATION AND PROTEIN-PROTEIN INTERACTIONS Eric M. Wauson A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Pharmacology. Chapel Hill 2007 Approved by: Lee M. Graves, Ph.D. T. Kendall Harden, Ph.D. Gary L. Johnson, Ph.D. Aziz Sancar M.D., Ph.D. Beverly S. Mitchell, M.D. 2007 Eric M. Wauson ALL RIGHTS RESERVED ii ABSTRACT Eric M. Wauson: The Regulation of Carbamoyl Phosphate Synthetase-Aspartate Transcarbamoylase-Dihydroorotase (CAD) by Phosphorylation and Protein-Protein Interactions (Under the direction of Lee M. Graves, Ph.D.) Pyrimidines have many important roles in cellular physiology, as they are used in the formation of DNA, RNA, phospholipids, and pyrimidine sugars. The first rate- limiting step in the de novo pyrimidine synthesis pathway is catalyzed by the carbamoyl phosphate synthetase II (CPSase II) part of the multienzymatic complex Carbamoyl phosphate synthetase, Aspartate transcarbamoylase, Dihydroorotase (CAD). CAD gene induction is highly correlated to cell proliferation. Additionally, CAD is allosterically inhibited or activated by uridine triphosphate (UTP) or phosphoribosyl pyrophosphate (PRPP), respectively. The phosphorylation of CAD by PKA and ERK has been reported to modulate the response of CAD to allosteric modulators. While there has been much speculation on the identity of CAD phosphorylation sites, no definitive identification of in vivo CAD phosphorylation sites has been performed. Therefore, we sought to determine the specific CAD residues phosphorylated by ERK and PKA in intact cells. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -
![UBE2B (HR6B) [Untagged] E2 – Ubiquitin Conjugating Enzyme](https://docslib.b-cdn.net/cover/5816/ube2b-hr6b-untagged-e2-ubiquitin-conjugating-enzyme-305816.webp)
UBE2B (HR6B) [Untagged] E2 – Ubiquitin Conjugating Enzyme
UBE2B (HR6B) [untagged] E2 – Ubiquitin Conjugating Enzyme Alternate Names: HHR6B, HR6B, RAD6B, Ubiquitin carrier protein B, Ubiquitin protein ligase B Cat. No. 62-0004-100 Quantity: 100 µg Lot. No. 1456 Storage: -70˚C FOR RESEARCH USE ONLY NOT FOR USE IN HUMANS CERTIFICATE OF ANALYSIS Page 1 of 2 Background Physical Characteristics The enzymes of the ubiquitylation Species: human Protein Sequence: pathway play a pivotal role in a num- GPLGSSTPARRRLMRDFKRLQEDPPVGVS ber of cellular processes including Source: E. coli expression GAPSENNIMQWNAVIFGPEGTPFEDGT regulated and targeted proteasomal FKLVIEFSEEYPNKPPTVRFLSKMFHPNVY degradation of substrate proteins. Quantity: 100 μg ADGSICLDILQNRWSPTYDVSSILTSIQSLL DEPNPNSPANSQAAQLYQENKREYEKRV Three classes of enzymes are in- Concentration: 1 mg/ml SAIVEQSWNDS volved in the process of ubiquitylation; activating enzymes (E1s), conjugating Formulation: 50 mM HEPES pH 7.5, enzymes (E2s) and protein ligases 150 mM sodium chloride, 2 mM The residues underlined remain after cleavage and removal of the purification tag. (E3s). UBE2B is a member of the E2 dithiothreitol, 10% glycerol UBE2B (regular text): Start bold italics (amino acid residues ubiquitin-conjugating enzyme family 2-152) and cloning of the human gene was Molecular Weight: ~17 kDa Accession number: NP_003328 first described by Koken et al. (1991). UBE2B shares 70% identity with its Purity: >98% by InstantBlue™ SDS-PAGE yeast homologue but lacks the acidic Stability/Storage: 12 months at -70˚C; C-terminal domain. The ring finger aliquot as required proteins RAD5 and RAD18 interact with UBE2B and other members of the RAD6 pathway (Notenboom et al., Quality Assurance 2007; Ulrich and Jentsch, 2000). In complex UBE2B and RAD18 trigger Purity: Protein Identification: replication fork stalling at DNA dam- 4-12% gradient SDS-PAGE Confirmed by mass spectrometry InstantBlue™ staining age sites during the post replicative Lane 1: MW markers E2-Ubiquitin Thioester Loading Assay: repair process (Tsuji et al., 2008). -

DNA Replication Stress Response Involving PLK1, CDC6, POLQ
DNA replication stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients C. Allera-Moreau, I. Rouquette, B. Lepage, N. Oumouhou, M. Walschaerts, E. Leconte, V. Schilling, K. Gordien, L. Brouchet, Mb Delisle, et al. To cite this version: C. Allera-Moreau, I. Rouquette, B. Lepage, N. Oumouhou, M. Walschaerts, et al.. DNA replica- tion stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients. Oncogenesis, Nature Publishing Group: Open Access Journals - Option C, 2012, 1, pp.e30. 10.1038/oncsis.2012.29. hal-00817701 HAL Id: hal-00817701 https://hal.archives-ouvertes.fr/hal-00817701 Submitted on 9 Jun 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial - NoDerivatives| 4.0 International License Citation: Oncogenesis (2012) 1, e30; doi:10.1038/oncsis.2012.29 & 2012 Macmillan Publishers Limited All rights reserved 2157-9024/12 www.nature.com/oncsis ORIGINAL ARTICLE DNA replication stress response involving PLK1, CDC6, POLQ, RAD51 and CLASPIN upregulation prognoses the outcome of early/mid-stage non-small cell lung cancer patients C Allera-Moreau1,2,7, I Rouquette2,7, B Lepage3, N Oumouhou3, M Walschaerts4, E Leconte5, V Schilling1, K Gordien2, L Brouchet2, MB Delisle1,2, J Mazieres1,2, JS Hoffmann1, P Pasero6 and C Cazaux1 Lung cancer is the leading cause of cancer deaths worldwide. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Dysfunction of Human Rad18 Results in Defective Postreplication Repair and Hypersensitivity to Multiple Mutagens
Dysfunction of human Rad18 results in defective postreplication repair and hypersensitivity to multiple mutagens Satoshi Tateishi*, Yoshiyuki Sakuraba†, Sadaharu Masuyama*, Hirokazu Inoue†, and Masaru Yamaizumi*‡ *Institute of Molecular Embryology and Genetics, Kumamoto University School of Medicine, Kumamoto 862-0976, Japan; and †Department of Regulation Biology, Faculty of Science, Saitama University, Urawa 338-8570, Japan Communicated by Philip C. Hanawalt, Stanford University, Stanford, CA, April 7, 2000 (received for review January 25, 2000) Postreplication repair functions in gap-filling of a daughter strand CCATACTCAACTATCC-3Ј, which amplify a 622-bp fragment on replication of damaged DNA. The yeast Saccharomyces cerevi- in the 3Ј untranslated region of hRAD18. Human hHR6A and siae Rad18 protein plays a pivotal role in the process together with hHR6B cDNA were obtained by PCR using a cDNA library the Rad6 protein. Here, we have cloned a human homologue of prepared from normal human fibroblasts. RAD18, hRAD18. It maps on chromosome 3p24–25, where dele- tions are often found in lung, breast, ovary, and testis cancers. In Cell Culture and Transfection. Cells were cultured in DMEM vivo, hRad18 protein binds to hHR6 protein through a conserved supplemented with 10% FCS, 100 units͞ml penicillin G, and 100 ͞ ring-finger motif. Stable transformants with hRad18 mutated in g ml streptomycin in a humidified 5% CO2 incubator. Mori-SV this motif become sensitive to UV, methyl methanesulfonate, and is a normal human skin fibroblast immortalized with simian virus mitomycin C, and are defective in the replication of UV-damaged 40. Mutant hRAD18 cDNAs were produced with a Quick- DNA. Thus, hRAD18 is a functional homologue of RAD18. -
Supplementary Table 1
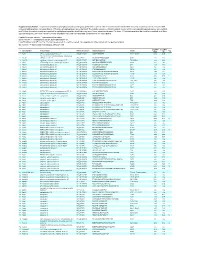
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Requirement of Rad18 Protein for Replication Through DNA Lesions in Mouse and Human Cells
Requirement of Rad18 protein for replication through DNA lesions in mouse and human cells Jung-Hoon Yoon, Satya Prakash, and Louise Prakash1 Department of Biochemistry and Molecular Biology, University of Texas Medical Branch at Galveston, Galveston, TX 77555 Edited* by Jerard Hurwitz, Memorial Sloan-Kettering Cancer Center, New York, NY, and approved March 21, 2012 (received for review March 8, 2012) In yeast, the Rad6-Rad18 ubiquitin conjugating enzyme plays a translocase activity that could function directly in lesion bypass critical role in promoting replication although DNA lesions by by promoting replication fork regression and template switching translesion synthesis (TLS). In striking contrast, a number of (11). Genetic studies in yeast have provided evidence for the studies have indicated that TLS can occur in the absence of requirement of Mms2, Ubc13, and Rad5 for a template-switch- Rad18 in human and other mammalian cells, and also in chicken ing mode of lesion bypass and they have shown that both the cells. In this study, we determine the role of Rad18 in TLS that ubiquitin ligase and DNA translocase activities are essential for occurs during replication in human and mouse cells, and show that Rad5 to carry out its role in lesion bypass (6, 12). in the absence of Rad18, replication of duplex plasmids containing Mammalian cells have two RAD6 homologs, RAD6A and a cis-syn TT dimer or a (6-4) TT photoproduct is severely inhibited RAD6B, and a single RAD18 gene. Importantly, in contrast to in human cells and that mutagenesis resulting from TLS opposite the indispensability of the Rad6-Rad18 enzyme and of PCNA cyclobutane pyrimidine dimers and (6-4) photoproducts formed at ubiquitylation for TLS in yeast cells, several studies have in- the TT, TC, and CC dipyrimidine sites in the chromosomal cII gene dicated that Rad6-Rad18 and PCNA ubiquitylation may not play in UV-irradiated mouse cells is abolished. -

2. Improvement of Itraq-SPROX Protocol
Development and Application of Mass Spectrometry-Based Approaches for Thermodynamic Analysis of Protein-Ligand Binding Interactions by Xiaopu (Lorrain) Jin Department of Chemistry Duke University Date:_______________________ Approved: ___________________________ Michael C. Fitzgerald, Supervisor ___________________________ Jie Liu ___________________________ Terrence G. Oas ___________________________ Katherine J. Franz Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Chemistry in the Graduate School of Duke University 2017 i v ABSTRACT Development and Application of Mass Spectrometry-Based Approaches for Thermodynamic Analysis of Protein-Ligand Binding Interactions by Xiaopu (Lorrain) Jin Department of Chemistry Duke University Date:_______________________ Approved: ___________________________ Michael C. Fitzgerald, Supervisor ___________________________ Jie Liu ___________________________ Terrence G. Oas ___________________________ Katherine J. Franz An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Chemistry in the Graduate School of Duke University 2017 Copyright by Xiaopu (Lorrain) Jin 2017 Abstract The characterization of protein stability changes and protein-ligand interactions on the proteomic scale is important for understanding the biology of cellular processes. The identification and quantification of protein-ligand binding affinities is critical for disease -

ABSTRACT Title of Dissertation: INTEGRATION OF
ABSTRACT Title of Dissertation: INTEGRATION OF 18O LABELING AND SOLUTION ISOELECTRIC FOCUSING IN A SHOTGUN ANALYSIS OF MITOCHONDRIAL PROTEINS Jinshan Wang, Doctor of Philosophy, 2007 Dissertation Directed by: Professor Catherine Fenselau Department of Chemistry and Biochemistry The coupling of efficient separations and mass spectrometry instrumentation is highly desirable to provide global proteomic analysis. When quantitative comparisons are part of the strategy, separation and analytical methods should be selected, which optimize the isotope labeling procedure. Enzyme-catalyzed 18O labeling is considered to be the labeling method most compatible with analysis of proteins from tissue and other limited samples. The introduction of label at the peptide stage mandates that protein manipulation be minimized in favor of peptide fractionation post-labeling. In the present study, forward and reverse 18O labeling are integrated with solution isoelectric focusing and capillary LC-tandem mass spectrometry to study changes in mitochondrial proteins associated with drug resistance in human cancer cells. A total of 637 peptides corresponding to 278 proteins were identified in this analysis. Of these, twelve proteins have been demonstrated from the forward and reverse labeling experiments to have abundances altered by greater than a factor of two between the drug susceptible MCF-7 cell line and the MCF-7 cell line selected for resistance to mitoxantrone. Galectin-3 binding protein precursor was detected in the resistant cell line, but was not detected in the drug susceptible line. Such proteins are challenging to 18O and other isotope strategies and a solution is offered, based on reverse labeling. These twelve proteins play a role in several pathways including apoptosis, oxidative phosphorylation, fatty acid metabolism and amino acid metabolism. -
![UBE2A (HR6A) [GST-Tagged] E2 – Ubiquitin Conjugating Enzyme](https://docslib.b-cdn.net/cover/1509/ube2a-hr6a-gst-tagged-e2-ubiquitin-conjugating-enzyme-1351509.webp)
UBE2A (HR6A) [GST-Tagged] E2 – Ubiquitin Conjugating Enzyme
UBE2A (HR6A) [GST-tagged] E2 – Ubiquitin Conjugating Enzyme Alternate Names: HHR6A, HR6A, RAD6A, UBC2, EC 6.3.2.19, Ubiquitin-conjugating enzyme E2A Cat. No. 62-0001-020 Quantity: 20 µg Lot. No. 1384 Storage: -70˚C FOR RESEARCH USE ONLY NOT FOR USE IN HUMANS CERTIFICATE OF ANALYSIS Background Physical Characteristics The enzymes of the ubiquitylation pathway Species: human Protein Sequence: play a pivotal role in a number of cellular MSPILGYWKIKGLVQPTRLLLEYLEEKYEEH processes including the regulated and tar- Source: E. coli expression LYERDEGDKWRNKKFELGLEFPNLPYYIDGD geted proteosomal degradation of sub- VKLTQSMAIIRYIADKHNMLGGCPKER strate proteins. Three classes of enzymes Quantity: 20 µg AEISMLEGAVLDIRYGVSRIAYSKDFETLKVD are involved in the process of ubiquityla- FLSKLPEMLKMFEDRLCHKTYLNGDHVTHP tion; activating enzymes (E1s), conjugating Concentration: 1 mg/ml DFMLYDALDVVLYMDPMCLDAFPKLVCFK enzymes (E2s) and protein ligases (E3s). KRIEAIPQIDKYLKSSKYIAWPLQGWQATFG UBE2A is a member of the E2 conjugating Formulation: 50 mM HEPES pH 7.5, GGDHPPKSDLEVLFQGPLGSPNSRVDSTPAR enzyme family and cloning of the human 150 mM sodium chloride, 2 mM RRLMRDFKRLQEDPPAGVSGAPSENNIMVW gene was first described by Koken et al. dithiothreitol, 10% glycerol NAVIFGPEGTPFEDGTFKLTIEFTEEYPNKPPT (1991). UBE2A shares 70% identity with VRFVSKMFHPNVYADGSICLDILQNRWSPTYD its yeast homologue but lacks the acidic c- Molecular Weight: ~45 kDa VSSILTSIQSLLDEPNPNSPANSQAAQLYQENK terminal domain. The ring finger proteins REYEKRVSAIVEQSWRDC RAD5 and RAD18 interact with UBE2A and Purity: >98% by InstantBlue™ SDS-PAGE Tag (bold text): N-terminal glutathione-S-transferase (GST) other members of the RAD6 pathway (Ul- Protease cleavage site: PreScission™ (LEVLFQtGP) rich and Jentsch, 2000). Phosphorylation of Stability/Storage: 12 months at -70˚C; UBE2A (regular text): Start bold italics (amino acid UBE2A by CDK1 and 2 increases its activ- aliquot as required residues 2-152) Accession number: NP_003327 ity during the G2/M phase of the cell cycle (Sarcevic et al., 2002).