Cell–Cell Adhesion Molecules During Neocortical Development
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Adhesion Molecules in Non-Melanoma Skin Cancers: a Comprehensive Review JOANNA POGORZELSKA-DYRBUS 1 and JACEK C
in vivo 35 : 1327-1336 (2021) doi:10.21873/invivo.12385 Review Adhesion Molecules in Non-melanoma Skin Cancers: A Comprehensive Review JOANNA POGORZELSKA-DYRBUS 1 and JACEK C. SZEPIETOWSKI 2 1“Estevita” Specialist Medical Practice, Tychy, Poland; 2Department of Dermatology, Venereology and Allergology, Wroclaw Medical University, Wroclaw, Poland Abstract. Basal cell carcinoma (BCC) and squamous cell of NMSC develops from basal epithelial cells of hair carcinoma (SCC) are the most frequently diagnosed cancers, follicles or pluripotent epidermal basal cells and has a generating significant medical and financial problems. metastatic rate of only 0.0028-0.05% (1). Depending on the Cutaneous carcinogenesis is a very complex process different features of tumor cells, there are many characterized by genetic and molecular alterations, and histological types of BCC, with different, but still low mediated by various proteins and pathways. Cell adhesion metastatic potential (2). SCC develops from the molecules (CAMs) are transmembrane proteins responsible proliferating squamous layer of the epidermis, shows a for cell-to-cell and cell-to-extracellular matrix adhesion, metastatic rate of 0,1-9,9% and contributes to engaged in all steps of tumor progression. Based on their approximately 75% of deaths due to NMSC (3, 4). structures they are divided into five major groups: cadherins, Although both skin cancers generally have a good integrins, selectins, immunoglobulins and the CD44 family. prognosis, due to their high prevalence, they generate Cadherins, -

Propranolol-Mediated Attenuation of MMP-9 Excretion in Infants with Hemangiomas
Supplementary Online Content Thaivalappil S, Bauman N, Saieg A, Movius E, Brown KJ, Preciado D. Propranolol-mediated attenuation of MMP-9 excretion in infants with hemangiomas. JAMA Otolaryngol Head Neck Surg. doi:10.1001/jamaoto.2013.4773 eTable. List of All of the Proteins Identified by Proteomics This supplementary material has been provided by the authors to give readers additional information about their work. © 2013 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 10/01/2021 eTable. List of All of the Proteins Identified by Proteomics Protein Name Prop 12 mo/4 Pred 12 mo/4 Δ Prop to Pred mo mo Myeloperoxidase OS=Homo sapiens GN=MPO 26.00 143.00 ‐117.00 Lactotransferrin OS=Homo sapiens GN=LTF 114.00 205.50 ‐91.50 Matrix metalloproteinase‐9 OS=Homo sapiens GN=MMP9 5.00 36.00 ‐31.00 Neutrophil elastase OS=Homo sapiens GN=ELANE 24.00 48.00 ‐24.00 Bleomycin hydrolase OS=Homo sapiens GN=BLMH 3.00 25.00 ‐22.00 CAP7_HUMAN Azurocidin OS=Homo sapiens GN=AZU1 PE=1 SV=3 4.00 26.00 ‐22.00 S10A8_HUMAN Protein S100‐A8 OS=Homo sapiens GN=S100A8 PE=1 14.67 30.50 ‐15.83 SV=1 IL1F9_HUMAN Interleukin‐1 family member 9 OS=Homo sapiens 1.00 15.00 ‐14.00 GN=IL1F9 PE=1 SV=1 MUC5B_HUMAN Mucin‐5B OS=Homo sapiens GN=MUC5B PE=1 SV=3 2.00 14.00 ‐12.00 MUC4_HUMAN Mucin‐4 OS=Homo sapiens GN=MUC4 PE=1 SV=3 1.00 12.00 ‐11.00 HRG_HUMAN Histidine‐rich glycoprotein OS=Homo sapiens GN=HRG 1.00 12.00 ‐11.00 PE=1 SV=1 TKT_HUMAN Transketolase OS=Homo sapiens GN=TKT PE=1 SV=3 17.00 28.00 ‐11.00 CATG_HUMAN Cathepsin G OS=Homo -

Cdh2 Coordinates Myosin-II Dependent Internalisation of the Zebrafish Neural Plate
www.nature.com/scientificreports Corrected: Publisher Correction OPEN Cdh2 coordinates Myosin-II dependent internalisation of the zebrafsh neural plate Received: 4 June 2018 Claudio Araya1,2, Hanna-Maria Häkkinen 1, Luis Carcamo1, Mauricio Cerda3,4, Thierry Savy5,6, Accepted: 7 December 2018 Christopher Rookyard7, Nadine Peyriéras5,6 & Jonathan D. W. Clarke7 Published online: 12 February 2019 Tissue internalisation is a key morphogenetic mechanism by which embryonic tissues generate complex internal organs and a number of studies of epithelia have outlined a general view of tissue internalisation. Here we have used quantitative live imaging and mutant analysis to determine whether similar mechanisms are responsible for internalisation in a tissue that apparently does not have a typical epithelial organisation – the zebrafsh neural plate. We found that although zebrafsh embryos begin neurulation without a conventional epithelium, medially located neural plate cells adopt strategies typical of epithelia in order to constrict their dorsal surface membrane during cell internalisation. Furthermore, we show that Myosin-II activity is a signifcant driver of this transient cell remodeling which also depends on Cdh2 (N-cadherin). Abrogation of Cdh2 results in defective Myosin-II distribution, mislocalised internalisation events and defective neural plate morphogenesis. Our work suggests Cdh2 coordinates Myosin-II dependent internalisation of the zebrafsh neural plate. Te internalisation of superfcial sheets of cells is a widely used developmental strategy to generate complex three-dimensional structures with well-defned shape and size. Trough this mechanism, animal tissues form a number of internal organs including the vertebrate central nervous system1,2. In recent years, live imaging studies and mutant analysis has begun to defne the cellular, molecular, and biomechanical mechanisms responsible for tissue internalisation in a growing number of tractable model systems3,4. -

The N-Cadherin Interactome in Primary Cardiomyocytes As Defined Using Quantitative Proximity Proteomics Yang Li1,*, Chelsea D
© 2019. Published by The Company of Biologists Ltd | Journal of Cell Science (2019) 132, jcs221606. doi:10.1242/jcs.221606 TOOLS AND RESOURCES The N-cadherin interactome in primary cardiomyocytes as defined using quantitative proximity proteomics Yang Li1,*, Chelsea D. Merkel1,*, Xuemei Zeng2, Jonathon A. Heier1, Pamela S. Cantrell2, Mai Sun2, Donna B. Stolz1, Simon C. Watkins1, Nathan A. Yates1,2,3 and Adam V. Kwiatkowski1,‡ ABSTRACT requires multiple adhesion, cytoskeletal and signaling proteins, The junctional complexes that couple cardiomyocytes must transmit and mutations in these proteins can cause cardiomyopathies (Ehler, the mechanical forces of contraction while maintaining adhesive 2018). However, the molecular composition of ICD junctional homeostasis. The adherens junction (AJ) connects the actomyosin complexes remains poorly defined. – networks of neighboring cardiomyocytes and is required for proper The core of the AJ is the cadherin catenin complex (Halbleib and heart function. Yet little is known about the molecular composition of the Nelson, 2006; Ratheesh and Yap, 2012). Classical cadherins are cardiomyocyte AJ or how it is organized to function under mechanical single-pass transmembrane proteins with an extracellular domain that load. Here, we define the architecture, dynamics and proteome of mediates calcium-dependent homotypic interactions. The adhesive the cardiomyocyte AJ. Mouse neonatal cardiomyocytes assemble properties of classical cadherins are driven by the recruitment of stable AJs along intercellular contacts with organizational and cytosolic catenin proteins to the cadherin tail, with p120-catenin β structural hallmarks similar to mature contacts. We combine (CTNND1) binding to the juxta-membrane domain and -catenin β quantitative mass spectrometry with proximity labeling to identify the (CTNNB1) binding to the distal part of the tail. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Binding Mode of the Side-By-Side Two-Igv Molecule CD226/DNAM-1 to Its Ligand CD155/Necl-5
Binding mode of the side-by-side two-IgV molecule CD226/DNAM-1 to its ligand CD155/Necl-5 Han Wanga, Jianxun Qib, Shuijun Zhangb,1, Yan Lib, Shuguang Tanb,2, and George F. Gaoa,b,2 aResearch Network of Immunity and Health (RNIH), Beijing Institutes of Life Science, Chinese Academy of Sciences (CAS), 100101 Beijing, China; and bCAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, 100101 Beijing, China Edited by K. Christopher Garcia, Stanford University School of Medicine, Stanford, CA, and approved December 3, 2018 (received for review September 11, 2018) Natural killer (NK) cells are important component of innate immu- CD226, also known as DNAM-1, belongs to the Ig superfamily nity and also contribute to activating and reshaping the adaptive and contains two extracellular Ig-like domains (CD226-D1 and immune responses. The functions of NK cells are modulated by CD226-D2), and is widely expressed in monocytes, platelets, multiple inhibitory and stimulatory receptors. Among these recep- T cells, and most of the resting NK cells (8, 13, 19, 20). The tors, the activating receptor CD226 (DNAM-1) mediates NK cell intracellular domain of CD226 does not contain a tyrosine-based activation via binding to its nectin-like (Necl) family ligand, CD155 activation motif, which is accepted as responsible for activating (Necl-5). Here, we present a unique side-by-side arrangement signal transduction of stimulatory molecules (13). Instead, it pattern of two tandem immunoglobulin V-set (IgV) domains transmits the downstream signaling by phosphorylation of in- deriving from the ectodomains of both human CD226 (hCD226- tracellular phosphorylation sites and subsequent association with ecto) and mouse CD226 (mCD226-ecto), which is substantially integrin lymphocyte function-associated antigen 1 (21). -
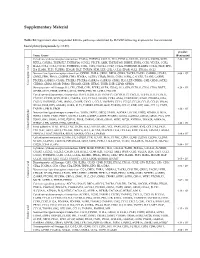
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Endocytosis Elicited by Nectins Transfers Cytoplasmic Cargo, Including Infectious Material, Between Cells Alex R
© 2019. Published by The Company of Biologists Ltd | Journal of Cell Science (2019) 132, jcs235507. doi:10.1242/jcs.235507 RESEARCH ARTICLE Trans-endocytosis elicited by nectins transfers cytoplasmic cargo, including infectious material, between cells Alex R. Generous1,2, Oliver J. Harrison3, Regina B. Troyanovsky4, Mathieu Mateo1,*, Chanakha K. Navaratnarajah1, Ryan C. Donohue1,2, Christian K. Pfaller1,2, Olga Alekhina5, Alina P. Sergeeva3,6, Indrajyoti Indra4, Theresa Thornburg7,‡, Irina Kochetkova7, Daniel D. Billadeau5, Matthew P. Taylor7, Sergey M. Troyanovsky4, Barry Honig3,6, Lawrence Shapiro3 and Roberto Cattaneo1,2,§ ABSTRACT development, where the Bride of sevenless protein is internalized by the Sevenless tyrosine kinase receptor (Cagan et al., 1992). Here, we show that cells expressing the adherens junction protein Transfer of specific transmembrane proteins also occurs during nectin-1 capture nectin-4-containing membranes from the surface tissue patterning in embryonic development of higher of adjacent cells in a trans-endocytosis process. We find that vertebrates, during epithelial cell movement and at the immune internalized nectin-1–nectin-4 complexes follow the endocytic synapse (Gaitanos et al., 2016; Hudrisier et al., 2001; Marston et al., pathway. The nectin-1 cytoplasmic tail controls transfer: its deletion 2003; Matsuda et al., 2004; Qureshi et al., 2011). At the immune prevents trans-endocytosis, while its exchange with the nectin-4 tail synapse, the CTLA-4 protein captures its ligands CD80 and CD86 reverses transfer direction. Nectin-1-expressing cells acquire dye- from donor cells by a process of trans-endocytosis; after removal, labeled cytoplasmic proteins synchronously with nectin-4, a process these ligands are degraded inside the acceptor cell, resulting in most active during cell adhesion. -

The Poliovirus Receptor (CD155)
Cutting Edge: CD96 (Tactile) Promotes NK Cell-Target Cell Adhesion by Interacting with the Poliovirus Receptor (CD155) This information is current as Anja Fuchs, Marina Cella, Emanuele Giurisato, Andrey S. of September 27, 2021. Shaw and Marco Colonna J Immunol 2004; 172:3994-3998; ; doi: 10.4049/jimmunol.172.7.3994 http://www.jimmunol.org/content/172/7/3994 Downloaded from References This article cites 19 articles, 8 of which you can access for free at: http://www.jimmunol.org/content/172/7/3994.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 27, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2004 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. THE JOURNAL OF IMMUNOLOGY CUTTING EDGE Cutting Edge: CD96 (Tactile) Promotes NK Cell-Target Cell Adhesion by Interacting with the Poliovirus Receptor (CD155) Anja Fuchs, Marina Cella, Emanuele Giurisato, Andrey S. Shaw, and Marco Colonna1 The poliovirus receptor (PVR) belongs to a large family of activating receptor DNAM-1, also called CD226 (6, 7). -

Minimal 16Q Genomic Loss Implicates Cadherin-11 in Retinoblastoma
Minimal 16q Genomic Loss Implicates Cadherin-11 in Retinoblastoma Mellone N. Marchong,1,2 Danian Chen,1,6 Timothy W. Corson,1,3 Cheong Lee,1 Maria Harmandayan,1 Ella Bowles,1 Ning Chen,5 and Brenda L. Gallie1,2,3,4,5 1Divisions of Cancer Informatics and Cellular and Molecular Biology, Ontario Cancer Institute/Princess Margaret Hospital, University Health Network, Toronto, Ontario, Canada; Departments of 2Medical Biophysics, 3Molecular and Medical Genetics, and 4Ophthalmology, University of Toronto, Toronto, Ontario, Canada; 5Retinoblastoma Solutions, Toronto, Ontario, Canada; and 6Department of Ophthalmology, West China Hospital, Faculty of Medicine, Sichuan University, Chengdu, People’s Republic of China Abstract compared with normal adult human retina. Our analyses Retinoblastoma is initiated by loss of both RB1 alleles. implicate CDH11, but not CDH13, as a potential tumor Previous studies have shown that retinoblastoma suppressor gene in retinoblastoma. (Mol Cancer Res tumors also show further genomic gains and losses. We 2004;2(9):495–503) now define a 2.62 Mbp minimal region of genomic loss of chromosome 16q22, which is likely to contain tumor suppressor gene(s), in 76 retinoblastoma tumors, using Introduction loss of heterozygosity (30 of 76 tumors) and quantitative Retinoblastoma is the most common intraocular tumor in multiplex PCR (71 of 76 tumors). The sequence-tagged children. Mutations of both alleles (M1 and M2) of the RB1 site WI-5835 within intron 2 of the cadherin-11 (CDH11) gene at chromosome 13q14 are necessary (1) for retinoblastoma gene showed the highest frequency of loss (54%, 22 of tumor initiation but not sufficient for malignant transformation 41 samples tested). -

Human R-Cadherin / CDH4 Protein (His Tag)
Human R-Cadherin / CDH4 Protein (His Tag) Catalog Number: 10230-H08H General Information SDS-PAGE: Gene Name Synonym: CAD4; R-CAD; RCAD Protein Construction: A DNA sequence encoding the extracellular domain of human CAD4 (NP_001785.2) (Met 1-Ala 734) was expressed with a fused polyhistidine tag at the C-terminus. Source: Human Expression Host: HEK293 Cells QC Testing Purity: > 85 % as determined by SDS-PAGE Endotoxin: Protein Description < 1.0 EU per μg of the protein as determined by the LAL method The cadherin superfamily is a large family that engage in both homo- and heterotypic, calcium-dependent, cell-cell adhesion events, and can be Stability: divided into at least four subfamilies based on the extracellular (EC) regions and cytoplasmic domains, that is: classical cadherins, desmosomal Samples are stable for up to twelve months from date of receipt at -70 ℃ cadherins, protocadherins, and cadherin-like molecules. Human cadherin 4, type 1, R-cadherin (retinal), also known as CDH4, CAD4 and RCAD, is a Predicted N terminal: His 21 classical cadherin from the cadherin superfamily. It is a calcium-dependent Molecular Mass: adhesion molecule and a type I transmembrane glycoprotein composed of five extracellular cadherin repeats, a transmembrane region and a highly The recombinant pro form of human CAD4 comprises 725 amino acids and conserved cytoplasmic tail. CDH4 is thought to play an important role predicts a molecular mass of 80 kDa. As a result of glycosylation, rhCAD4 during brain segmentation and neuronal outgrowth, and also exerts critical migrates as an approximately 90-100 kDa protein in SDS-PAGE under actions in kidney and muscle development. -

Study on Expression of CDH4 in Lung Cancer Zhupeng Li1, Dan Su2, Lisha Ying2, Guangmao Yu1 and Weimin Mao3*
Li et al. World Journal of Surgical Oncology (2017) 15:26 DOI 10.1186/s12957-016-1083-2 RESEARCH Open Access Study on expression of CDH4 in lung cancer Zhupeng Li1, Dan Su2, Lisha Ying2, Guangmao Yu1 and Weimin Mao3* Abstract Background: The human CDH4 gene, which encodes the R-cadherin protein, has an important role in cell migration and cell adhesion, sorting, tissue morphogenesis, and tumor genesis. This study analyzed the relationship of CDH4 mRNA expression with lung cancer. Methods: Real time PCR was applied to detect CDH4 mRNA transcription in 142 paired cases of lung cancer and noncancerous regions. Results: No correlation was identified between CDH4 mRNA expression and gender, age, lymphnode metastasis, TNM stage, family history, smoking state, drinking state (P > 0.05), but grade and histotype (P < 0.05). The relative CDH4 mRNA value was remarkably decreased in lung cancer tissues compared with noncancerous tissues (P =0.001). Conclusions: We found that CDH4 mRNA expression was associated with grade and histotype. What is more, therelativeCDH4mRNAvaluewasdecreasedinthelung cancer tissues. Our results suggested that CDH4 might be a putative tumor suppressor gene (TSG) in lung cancer. Keywords: CDH4, Lung cancer, Tumor suppressor gene Background R-cadherin is a classic cadherin, which is important Lung cancer has the highest morbidity and mortality in for the differentiation of kidney, striated muscle, brain, malignant tumors and with a total of 1.5 million deaths and so on [6, 7]. R-cadherin plays a critical role in main- annually worldwide [1]. In the patients with lung cancer, taining cell polarity and tissue architecture in normal though diagnosis and treatment have been greatly gastrointestinal epithelial tissue [8].