Downloaded from Bioscientifica.Com at 09/23/2021 08:17:44PM Via Free Access
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Methods in and Applications of the Sequencing of Short Non-Coding Rnas" (2013)
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2013 Methods in and Applications of the Sequencing of Short Non- Coding RNAs Paul Ryvkin University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Bioinformatics Commons, Genetics Commons, and the Molecular Biology Commons Recommended Citation Ryvkin, Paul, "Methods in and Applications of the Sequencing of Short Non-Coding RNAs" (2013). Publicly Accessible Penn Dissertations. 922. https://repository.upenn.edu/edissertations/922 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/922 For more information, please contact [email protected]. Methods in and Applications of the Sequencing of Short Non-Coding RNAs Abstract Short non-coding RNAs are important for all domains of life. With the advent of modern molecular biology their applicability to medicine has become apparent in settings ranging from diagonistic biomarkers to therapeutics and fields angingr from oncology to neurology. In addition, a critical, recent technological development is high-throughput sequencing of nucleic acids. The convergence of modern biotechnology with developments in RNA biology presents opportunities in both basic research and medical settings. Here I present two novel methods for leveraging high-throughput sequencing in the study of short non- coding RNAs, as well as a study in which they are applied to Alzheimer's Disease (AD). The computational methods presented here include High-throughput Annotation of Modified Ribonucleotides (HAMR), which enables researchers to detect post-transcriptional covalent modifications ot RNAs in a high-throughput manner. In addition, I describe Classification of RNAs by Analysis of Length (CoRAL), a computational method that allows researchers to characterize the pathways responsible for short non-coding RNA biogenesis. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

The Centriole Protein CEP76 Negatively Regulates PLK1 Activity in the Cytoplasm for Proper Mitotic Progression
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs241281. doi:10.1242/jcs.241281 RESEARCH ARTICLE The centriole protein CEP76 negatively regulates PLK1 activity in the cytoplasm for proper mitotic progression Yutaka Takeda, Kaho Yamazaki, Kaho Hashimoto, Koki Watanabe, Takumi Chinen*,‡ and Daiju Kitagawa*,‡ ABSTRACT are important for accurate mitotic progression (Golsteyn, 1995; Polo-like kinase 1 (PLK1) dynamically changes its localization and Schmucker and Sumara, 2014). Several factors regulate PLK1 plays important roles in proper mitotic progression. In particular, strict localization during mitosis. PLK1 is recruited to centrosomes in a control of cytoplasmic PLK1 is needed to prevent mitotic defects. CDK1-dependent manner (Lee et al., 2014). The initial recruitment However, the regulation of cytoplasmic PLK1 is not fully understood. In of PLK1 to kinetochores is mediated by its interaction with PBIP1 this study, we show that CEP76, a centriolar protein, physically (also known as CENPU; Kang et al., 2006). During anaphase, PLK1 interacts with PLK1 and tightly controls the activation of cytoplasmic binds to PRC1 and changes its localization to the midbody (Hu et al., PLK1 during mitosis in human cells. We found that removal of 2012). In addition, strict control of PLK1 in the cytoplasm is needed centrosomes induced ectopic aggregation of PLK1, which is highly to prevent its aggregation and mitotic defects (Mukhopadhyay and phosphorylated, in the cytoplasm during mitosis. Importantly, a Dasso, 2010; Schmit et al., 2012; Zhao et al., 2016). However, the targeted RNAi screen revealed that depletion of CEP76 resulted in a detailed mechanisms through which the localization, expression and similar phenotype. -

Is Cep70, a Centrosomal Protein with New Roles in Breast Cancer Dissemination and Metastasis, a Facilitator of Epithelial-Mesenchymal Transition?
Is Cep70, a centrosomal protein with new roles in breast cancer dissemination and metastasis, a facilitator of epithelial-mesenchymal transition? Pedro A. Lazo 1,2 1 Experimental Therapeutics and Translational Oncology Program, Instituto de Biología Molecular y Celular del Cáncer, Consejo Superior de Investigaciones Científicas (CSIC), Universidad de Salamanca, Salamanca, Spain 2 Instituto de Investigación Biomédica de Salamanca (IBSAL), Hospital Universitario de Salamanca, Salamanca, Spain Running title: Cep70 and EMT Disclosures: None declared. Contact address: [email protected] 1 Introduction Microtubules are driving mechanisms of chromosomes, intracellular organelle movement, and cell shape and motility. Microtubules are organized on centrosomes, which are assembled in microtubule-organizing centers (MTOC). In mitosis there is no nuclear envelope and centromeres associated to microtubules are mainly involved in chromosome redistribution into daughter cells. In differentiated cells the microtubule- organizing centers are dispersed in the cytoplasm (non centrosomal (ncMTOC)) and interact with the minus end of microtubules through γ-tubulin. 1 However, it is not known if the microtubule contribution to tumor biology is only by facilitating tumor aneuploidy. During tumor dissemination, a process not linked to cell division, important changes take place in cell shape and motility. Microtubules are very dynamic because of their inherent structural instability, and this plasticity facilitates their reorganization during the epithelial-mesenchymal -
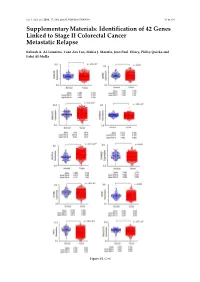
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

The Transformation of the Centrosome Into the Basal Body: Similarities and Dissimilarities Between Somatic and Male Germ Cells and Their Relevance for Male Fertility
cells Review The Transformation of the Centrosome into the Basal Body: Similarities and Dissimilarities between Somatic and Male Germ Cells and Their Relevance for Male Fertility Constanza Tapia Contreras and Sigrid Hoyer-Fender * Göttingen Center of Molecular Biosciences, Johann-Friedrich-Blumenbach Institute for Zoology and Anthropology-Developmental Biology, Faculty of Biology and Psychology, Georg-August University of Göttingen, 37077 Göttingen, Germany; [email protected] * Correspondence: [email protected] Abstract: The sperm flagellum is essential for the transport of the genetic material toward the oocyte and thus the transmission of the genetic information to the next generation. During the haploid phase of spermatogenesis, i.e., spermiogenesis, a morphological and molecular restructuring of the male germ cell, the round spermatid, takes place that includes the silencing and compaction of the nucleus, the formation of the acrosomal vesicle from the Golgi apparatus, the formation of the sperm tail, and, finally, the shedding of excessive cytoplasm. Sperm tail formation starts in the round spermatid stage when the pair of centrioles moves toward the posterior pole of the nucleus. The sperm tail, eventually, becomes located opposed to the acrosomal vesicle, which develops at the anterior pole of the nucleus. The centriole pair tightly attaches to the nucleus, forming a nuclear membrane indentation. An Citation: Tapia Contreras, C.; articular structure is formed around the centriole pair known as the connecting piece, situated in the Hoyer-Fender, S. The Transformation neck region and linking the sperm head to the tail, also named the head-to-tail coupling apparatus or, of the Centrosome into the Basal in short, HTCA. -

E-Mutpath: Computational Modelling Reveals the Functional Landscape of Genetic Mutations Rewiring Interactome Networks
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. e-MutPath: Computational modelling reveals the functional landscape of genetic mutations rewiring interactome networks Yongsheng Li1, Daniel J. McGrail1, Brandon Burgman2,3, S. Stephen Yi2,3,4,5 and Nidhi Sahni1,6,7,8,* 1Department oF Systems Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 2Department oF Oncology, Livestrong Cancer Institutes, Dell Medical School, The University oF Texas at Austin, Austin, TX 78712, USA 3Institute For Cellular and Molecular Biology (ICMB), The University oF Texas at Austin, Austin, TX 78712, USA 4Institute For Computational Engineering and Sciences (ICES), The University oF Texas at Austin, Austin, TX 78712, USA 5Department oF Biomedical Engineering, Cockrell School of Engineering, The University oF Texas at Austin, Austin, TX 78712, USA 6Department oF Epigenetics and Molecular Carcinogenesis, The University oF Texas MD Anderson Science Park, Smithville, TX 78957, USA 7Department oF BioinFormatics and Computational Biology, The University oF Texas MD Anderson Cancer Center, Houston, TX 77030, USA 8Program in Quantitative and Computational Biosciences (QCB), Baylor College oF Medicine, Houston, TX 77030, USA *To whom correspondence should be addressed. Nidhi Sahni. Tel: +1 512 2379506; Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.22.262386; this version posted August 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Suppl. Table 1
Suppl. Table 1. SiRNA library used for centriole overduplication screen. Entrez Gene Id NCBI gene symbol siRNA Target Sequence 1070 CETN3 TTGCGACGTGTTGCTAGAGAA 1070 CETN3 AAGCAATAGATTATCATGAAT 55722 CEP72 AGAGCTATGTATGATAATTAA 55722 CEP72 CTGGATGATTTGAGACAACAT 80071 CCDC15 ACCGAGTAAATCAACAAATTA 80071 CCDC15 CAGCAGAGTTCAGAAAGTAAA 9702 CEP57 TAGACTTATCTTTGAAGATAA 9702 CEP57 TAGAGAAACAATTGAATATAA 282809 WDR51B AAGGACTAATTTAAATTACTA 282809 WDR51B AAGATCCTGGATACAAATTAA 55142 CEP27 CAGCAGATCACAAATATTCAA 55142 CEP27 AAGCTGTTTATCACAGATATA 85378 TUBGCP6 ACGAGACTACTTCCTTAACAA 85378 TUBGCP6 CACCCACGGACACGTATCCAA 54930 C14orf94 CAGCGGCTGCTTGTAACTGAA 54930 C14orf94 AAGGGAGTGTGGAAATGCTTA 5048 PAFAH1B1 CCCGGTAATATCACTCGTTAA 5048 PAFAH1B1 CTCATAGATATTGAACAATAA 2802 GOLGA3 CTGGCCGATTACAGAACTGAA 2802 GOLGA3 CAGAGTTACTTCAGTGCATAA 9662 CEP135 AAGAATTTCATTCTCACTTAA 9662 CEP135 CAGCAGAAAGAGATAAACTAA 153241 CCDC100 ATGCAAGAAGATATATTTGAA 153241 CCDC100 CTGCGGTAATTTCCAGTTCTA 80184 CEP290 CCGGAAGAAATGAAGAATTAA 80184 CEP290 AAGGAAATCAATAAACTTGAA 22852 ANKRD26 CAGAAGTATGTTGATCCTTTA 22852 ANKRD26 ATGGATGATGTTGATGACTTA 10540 DCTN2 CACCAGCTATATGAAACTATA 10540 DCTN2 AACGAGATTGCCAAGCATAAA 25886 WDR51A AAGTGATGGTTTGGAAGAGTA 25886 WDR51A CCAGTGATGACAAGACTGTTA 55835 CENPJ CTCAAGTTAAACATAAGTCAA 55835 CENPJ CACAGTCAGATAAATCTGAAA 84902 CCDC123 AAGGATGGAGTGCTTAATAAA 84902 CCDC123 ACCCTGGTTGTTGGATATAAA 79598 LRRIQ2 CACAAGAGAATTCTAAATTAA 79598 LRRIQ2 AAGGATAATATCGTTTAACAA 51143 DYNC1LI1 TTGGATTTGTCTATACATATA 51143 DYNC1LI1 TAGACTTAGTATATAAATACA 2302 FOXJ1 CAGGACAGACAGACTAATGTA -

Regulation of the Centriole Life Cycle by Mdm1 a Dissertation Submitted to the Department of Biology and the Committee on Gradua
REGULATION OF THE CENTRIOLE LIFE CYCLE BY MDM1 A DISSERTATION SUBMITTED TO THE DEPARTMENT OF BIOLOGY AND THE COMMITTEE ON GRADUATE STUDIES OF STANFORD UNIVERSITY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY Daniel Patrick Van de Mark August 2014 © 2014 by Daniel Patrick Van de Mark. All Rights Reserved. Re-distributed by Stanford University under license with the author. This work is licensed under a Creative Commons Attribution- Noncommercial 3.0 United States License. http://creativecommons.org/licenses/by-nc/3.0/us/ This dissertation is online at: http://purl.stanford.edu/gf463wx1210 ii I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Tim Stearns, Primary Adviser I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Martha Cyert I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. William Nelson I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Rajat Rohatgi Approved for the Stanford University Committee on Graduate Studies. Patricia J. Gumport, Vice Provost for Graduate Education This signature page was generated electronically upon submission of this dissertation in electronic format. -

In Vivo ERK1/2 Reporter Predictively Models Response and Resistance to Combined
Author Manuscript Published OnlineFirst on July 3, 2019; DOI: 10.1158/1535-7163.MCT-18-1056 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. In vivo ERK1/2 reporter predictively models response and resistance to combined BRAF and MEK inhibitors in melanoma Ileine M. Sanchez1, Timothy J. Purwin1, Inna Chervoneva2, Dan A. Erkes1, Mai Q. Nguyen1, Michael A. Davies4, Katherine L. Nathanson5, Kristel Kemper6, Daniel S. Peeper6, and Andrew E. Aplin1,3 1 Department of Cancer Biology, Thomas Jefferson University, Philadelphia, PA 19107, USA 2 Division of Biostatistics, 3 Department of Pharmacology and Experimental Therapeutics Sidney Kimmel Cancer Center at Jefferson, Philadelphia, PA 19107, USA 4 Department of Melanoma Medical Oncology, Division of Cancer Medicine, University of Texas MD Anderson Cancer Center, Houston TX 77030, USA 5 Translational Medicine and Human Genetics, Department of Medicine, Perelman School of Medicine at the University of Pennsylvania, Abramson Cancer Center, Perelman School of Medicine at the University of Pennsylvania, Philadelphia PA 19104, USA 6 Division of Molecular Oncology & Immunology, Netherlands Cancer Institute, Plesmanlaan 121, 1066 CX Amsterdam, Netherlands Corresponding author: Andrew E. Aplin, Department of Cancer Biology, Thomas Jefferson University, 233 South 10th Street, Philadelphia, PA 19107. Tel: (215) 503- 7296. Fax: (215) 923-9248; E-mail: [email protected] Running title: In vivo pathway reporting on combination targeted therapy Keywords: melanoma, MEK, vemurafenib, resistance, BRAF 1 Downloaded from mct.aacrjournals.org on September 29, 2021. © 2019 American Association for Cancer Research. Author Manuscript Published OnlineFirst on July 3, 2019; DOI: 10.1158/1535-7163.MCT-18-1056 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2018 A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family Linda Molla Follow this and additional works at: https://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY A Thesis Presented to the Faculty of The Rockefeller University in Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Linda Molla June 2018 © Copyright by Linda Molla 2018 A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY Linda Molla, Ph.D. The Rockefeller University 2018 APOBEC2 is a member of the AID/APOBEC cytidine deaminase family of proteins. Unlike most of AID/APOBEC, however, APOBEC2’s function remains elusive. Previous research has implicated APOBEC2 in diverse organisms and cellular processes such as muscle biology (in Mus musculus), regeneration (in Danio rerio), and development (in Xenopus laevis). APOBEC2 has also been implicated in cancer. However the enzymatic activity, substrate or physiological target(s) of APOBEC2 are unknown. For this thesis, I have combined Next Generation Sequencing (NGS) techniques with state-of-the-art molecular biology to determine the physiological targets of APOBEC2. Using a cell culture muscle differentiation system, and RNA sequencing (RNA-Seq) by polyA capture, I demonstrated that unlike the AID/APOBEC family member APOBEC1, APOBEC2 is not an RNA editor. Using the same system combined with enhanced Reduced Representation Bisulfite Sequencing (eRRBS) analyses I showed that, unlike the AID/APOBEC family member AID, APOBEC2 does not act as a 5-methyl-C deaminase.