The Role of Sin1 in Cell Survival
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -

1 Supporting Information for a Microrna Network Regulates
Supporting Information for A microRNA Network Regulates Expression and Biosynthesis of CFTR and CFTR-ΔF508 Shyam Ramachandrana,b, Philip H. Karpc, Peng Jiangc, Lynda S. Ostedgaardc, Amy E. Walza, John T. Fishere, Shaf Keshavjeeh, Kim A. Lennoxi, Ashley M. Jacobii, Scott D. Rosei, Mark A. Behlkei, Michael J. Welshb,c,d,g, Yi Xingb,c,f, Paul B. McCray Jr.a,b,c Author Affiliations: Department of Pediatricsa, Interdisciplinary Program in Geneticsb, Departments of Internal Medicinec, Molecular Physiology and Biophysicsd, Anatomy and Cell Biologye, Biomedical Engineeringf, Howard Hughes Medical Instituteg, Carver College of Medicine, University of Iowa, Iowa City, IA-52242 Division of Thoracic Surgeryh, Toronto General Hospital, University Health Network, University of Toronto, Toronto, Canada-M5G 2C4 Integrated DNA Technologiesi, Coralville, IA-52241 To whom correspondence should be addressed: Email: [email protected] (M.J.W.); yi- [email protected] (Y.X.); Email: [email protected] (P.B.M.) This PDF file includes: Materials and Methods References Fig. S1. miR-138 regulates SIN3A in a dose-dependent and site-specific manner. Fig. S2. miR-138 regulates endogenous SIN3A protein expression. Fig. S3. miR-138 regulates endogenous CFTR protein expression in Calu-3 cells. Fig. S4. miR-138 regulates endogenous CFTR protein expression in primary human airway epithelia. Fig. S5. miR-138 regulates CFTR expression in HeLa cells. Fig. S6. miR-138 regulates CFTR expression in HEK293T cells. Fig. S7. HeLa cells exhibit CFTR channel activity. Fig. S8. miR-138 improves CFTR processing. Fig. S9. miR-138 improves CFTR-ΔF508 processing. Fig. S10. SIN3A inhibition yields partial rescue of Cl- transport in CF epithelia. -

Ten Commandments for a Good Scientist
Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Wageningen 2010 Thesis committee Thesis supervisors Prof. dr. ir. Ivonne M.C.M. Rietjens Professor of Toxicology Wageningen University Prof. dr. Albertinka J. Murk Personal chair at the sub-department of Toxicology Wageningen University Thesis co-supervisor Dr. ir. Jacques J.M. Vervoort Associate professor at the Laboratory of Biochemistry Wageningen University Other members Prof. dr. Michael R. Muller, Wageningen University Prof. dr. ir. Huub F.J. Savelkoul, Wageningen University Prof. dr. Everardus J. van Zoelen, Radboud University Nijmegen Dr. ir. Toine F.H. Bovee, RIKILT, Wageningen This research was conducted under the auspices of the Graduate School VLAG Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Thesis submitted in fulfillment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. dr. M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Tuesday 14 September 2010 at 4 p.m. in the Aula Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds. Ana María Sotoca Covaleda Thesis Wageningen University, Wageningen, The Netherlands, 2010, With references, and with summary in Dutch. ISBN: 978-90-8585-707-5 “Caminante no hay camino, se hace camino al andar. Al andar se hace camino, y al volver la vista atrás se ve la senda que nunca se ha de volver a pisar” - Antonio Machado – A mi madre. -

Comparison of Orthologs Across Multiple Species by Various Strategies
COMPARISON OF ORTHOLOGS ACROSS MULTIPLE SPECIES BY VARIOUS STRATEGIES BY HUI LIU DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biophysics and Computational Biology in the Graduate College of the University of Illinois at Urbana-Champaign, 2014 Urbana, Illinois Doctoral Committee: Professor Eric Jakobsson, Chair, Director of Research Professor Gene E. Robinson Associate Professor Saurabh Sinha Assistant Professor Jian Ma Abstract Thanks to the improvement of genome sequencing technology, abundant multi-species genomic data now became available and comparative genomics continues to be a fast prospering filed of biological research. Through the comparison of genomes of different organisms, we can understand what, at the molecular level, distinguishes different life forms from each other. It shed light on revealing the evolution of biology. And it also helps to refine the annotations and functions of individual genomes. For example, through comparisons across mammalian genomes, we can give an estimate of the conserved set of genes across mammals and correspondingly, find the species-specific sets of genes or functions. However, comparative genomics can be feasible only if a meaningful classification of genes exists. A natural way to do so is to delineate sets of orthologous genes. However, debates exist about the appropriate way to define orthologs. It is originally defined as genes in different species which derive from speciation events. But such definition is not sufficient to derive orthologous genes due to the complexity of evolutionary events such as gene duplication and gene loss. While it is possible to correctly figure out all the evolutionary events with the true phylogenetic tree, the true phylogenetic tree itself is impractical to be inferred. -

(12) Patent Application Publication (10) Pub. No.: US 2006/0088532 A1 Alitalo Et Al
US 20060O88532A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2006/0088532 A1 Alitalo et al. (43) Pub. Date: Apr. 27, 2006 (54) LYMPHATIC AND BLOOD ENDOTHELIAL Related U.S. Application Data CELL GENES (60) Provisional application No. 60/363,019, filed on Mar. (76) Inventors: Kari Alitalo, Helsinki (FI); Taija 7, 2002. Makinen, Helsinki (FI); Tatiana Petrova, Helsinki (FI); Pipsa Publication Classification Saharinen, Helsinki (FI); Juha Saharinen, Helsinki (FI) (51) Int. Cl. A6IR 48/00 (2006.01) Correspondence Address: A 6LX 39/395 (2006.01) MARSHALL, GERSTEIN & BORUN LLP A6II 38/18 (2006.01) 233 S. WACKER DRIVE, SUITE 6300 (52) U.S. Cl. .............................. 424/145.1: 514/2: 514/44 SEARS TOWER (57) ABSTRACT CHICAGO, IL 60606 (US) The invention provides polynucleotides and genes that are (21) Appl. No.: 10/505,928 differentially expressed in lymphatic versus blood vascular endothelial cells. These genes are useful for treating diseases (22) PCT Filed: Mar. 7, 2003 involving lymphatic vessels, such as lymphedema, various inflammatory diseases, and cancer metastasis via the lym (86). PCT No.: PCT/USO3FO6900 phatic system. Patent Application Publication Apr. 27, 2006 Sheet 1 of 2 US 2006/0088532 A1 integrin O9 integrin O1 KIAAO711 KAAO644 ApoD Fig. 1 Patent Application Publication Apr. 27, 2006 Sheet 2 of 2 US 2006/0088532 A1 CN g uueleo-gº US 2006/0O88532 A1 Apr. 27, 2006 LYMPHATIC AND BLOOD ENDOTHELLAL CELL lymphatic vessels, such as lymphangiomas or lymphang GENES iectasis. Witte, et al., Regulation of Angiogenesis (eds. Goldber, I. D. & Rosen, E. M.) 65-112 (Birkauser, Basel, BACKGROUND OF THE INVENTION Switzerland, 1997). -

Supplementary Table S2
1-high in cerebrotropic Gene P-value patients Definition BCHE 2.00E-04 1 Butyrylcholinesterase PLCB2 2.00E-04 -1 Phospholipase C, beta 2 SF3B1 2.00E-04 -1 Splicing factor 3b, subunit 1 BCHE 0.00022 1 Butyrylcholinesterase ZNF721 0.00028 -1 Zinc finger protein 721 GNAI1 0.00044 1 Guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 GNAI1 0.00049 1 Guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 PDE1B 0.00069 -1 Phosphodiesterase 1B, calmodulin-dependent MCOLN2 0.00085 -1 Mucolipin 2 PGCP 0.00116 1 Plasma glutamate carboxypeptidase TMX4 0.00116 1 Thioredoxin-related transmembrane protein 4 C10orf11 0.00142 1 Chromosome 10 open reading frame 11 TRIM14 0.00156 -1 Tripartite motif-containing 14 APOBEC3D 0.00173 -1 Apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3D ANXA6 0.00185 -1 Annexin A6 NOS3 0.00209 -1 Nitric oxide synthase 3 SELI 0.00209 -1 Selenoprotein I NYNRIN 0.0023 -1 NYN domain and retroviral integrase containing ANKFY1 0.00253 -1 Ankyrin repeat and FYVE domain containing 1 APOBEC3F 0.00278 -1 Apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3F EBI2 0.00278 -1 Epstein-Barr virus induced gene 2 ETHE1 0.00278 1 Ethylmalonic encephalopathy 1 PDE7A 0.00278 -1 Phosphodiesterase 7A HLA-DOA 0.00305 -1 Major histocompatibility complex, class II, DO alpha SOX13 0.00305 1 SRY (sex determining region Y)-box 13 ABHD2 3.34E-03 1 Abhydrolase domain containing 2 MOCS2 0.00334 1 Molybdenum cofactor synthesis 2 TTLL6 0.00365 -1 Tubulin tyrosine ligase-like family, member 6 SHANK3 0.00394 -1 SH3 and multiple ankyrin repeat domains 3 ADCY4 0.004 -1 Adenylate cyclase 4 CD3D 0.004 -1 CD3d molecule, delta (CD3-TCR complex) (CD3D), transcript variant 1, mRNA. -

Analysis of Pax6 Contiguous Gene Deletions in the Mouse, Mus
Genetics: Published Articles Ahead of Print, published on May 27, 2009 as 10.1534/genetics.109.104562 Analysis of Pax6 contiguous gene deletions in the mouse, Mus musculus, identifies regions distinct from Pax6 responsible for extreme small eye and belly spotting phenotypes Jack Favor*, Alan Bradley†, Nathalie Conte†, Dirk Janik‡, Walter Pretsch*, Peter Reitmeir§, Michael Rosemann**, Wolfgang Schmahl‡, Johannes Wienberg†† and Irmgard Zaus* Institute of Human Genetics*, Institute of Health Management§, Institute of Radiation Biology**, Helmholtz Zentrum München, German Research Center for Environmental Health, Neuherberg D- 85764, Germany † Wellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, UK ‡ Lehrstuhl für Allgemeine Pathologie und Neuropathologie, Tierärztliche Fakultät, Ludwig-Maximilians-Universität, München D-80539, Germany †† Chrombios GmbH, Raubling D-83064, Germany The mutant allele symbols Del(2)Pax611Neu/1Neu, Del(2)Pax612Neu/2Neu and Del(2)Pax613Neu/3Neu were submitted to and approved by the Mouse Genetic Nomenclature Committee, and assigned the MGI accession ID numbers 3698295, 3698296 and 3710946, respectively. 1 Running head: Mouse Pax6 contiguous gene deletions Key words: Mouse, Pax6, contiguous gene deletions, microphthalmia, belly spotting Corresponding author: Jack Favor Institute of Human Genetics Helmholtz Zentrum München German Research Center for Environmental Health Ingolstädter Lanstr. 1 D-85764 Neuherberg Germany Telephone No. +49-89-3187-2395 FAX No. +49-89-3187-3297 e-mail [email protected] -

Gene Expression Signatures and Biomarkers of Noninvasive And
Oncogene (2006) 25, 2328–2338 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc ORIGINAL ARTICLE Gene expression signatures and biomarkers of noninvasive and invasive breast cancer cells: comprehensive profiles by representational difference analysis, microarrays and proteomics GM Nagaraja1, M Othman2, BP Fox1, R Alsaber1, CM Pellegrino3, Y Zeng2, R Khanna2, P Tamburini3, A Swaroop2 and RP Kandpal1 1Department of Biological Sciences, Fordham University, Bronx, NY, USA; 2Department of Ophthalmology and Visual Sciences, University of Michigan, Ann Arbor, MI, USA and 3Bayer Corporation, West Haven, CT, USA We have characterized comprehensive transcript and Keywords: representational difference analysis; micro- proteomic profiles of cell lines corresponding to normal arrays; proteomics; breast carcinoma; biomarkers; breast (MCF10A), noninvasive breast cancer (MCF7) and copper homeostasis invasive breast cancer (MDA-MB-231). The transcript profiles were first analysed by a modified protocol for representational difference analysis (RDA) of cDNAs between MCF7 and MDA-MB-231 cells. The majority of genes identified by RDA showed nearly complete con- Introduction cordance withmicroarray results, and also led to the identification of some differentially expressed genes such The transformation of a normal cell into a cancer cell as lysyl oxidase, copper transporter ATP7A, EphB6, has been correlated to altered expression of a variety of RUNX2 and a variant of RUNX2. The altered transcripts genes (Perou et al., 2000; Becker et al., 2005). The identified by microarray analysis were involved in cell–cell expression of some of these genes is a direct result of or cell–matrix interaction, Rho signaling, calcium home- sequence mutation, whereas other changes occur due to ostasis and copper-binding/sensitive activities. -
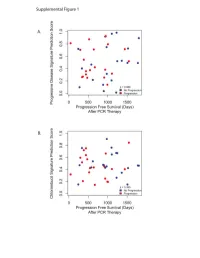
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Leveraging the Role of the Metastatic Associated Protein Anterior Gradient Homologue 2 in Unfolded Protein Degradation: a Novel Therapeutic Biomarker for Cancer
cancers Review Leveraging the Role of the Metastatic Associated Protein Anterior Gradient Homologue 2 in Unfolded Protein Degradation: A Novel Therapeutic Biomarker for Cancer Reem Alsereihi 1,2, Hans-Juergen Schulten 3,4 , Sherin Bakhashab 3,5 , Kulvinder Saini 6, Ahmed M. Al-Hejin 2,7 and Deema Hussein 1,4,* 1 Neurooncology Translational Group, King Fahd Medical Research Center, King Abdulaziz University, P.O. Box 80216, Jeddah 21589, Saudi Arabia 2 Department of Biological Sciences, Faculty of Science, King Abdulaziz University, P.O. Box 80203, Jeddah 21589, Saudi Arabia 3 Center of Excellence in Genomic Medicine Research, King Abdulaziz University, P.O. Box 80216, Jeddah 21589, Saudi Arabia 4 Department of Medical Laboratory Technology, Faculty of Applied Medical Sciences, King Abdulaziz University, Jeddah 21589, Saudi Arabia 5 Biochemistry Department, King Abdulaziz University, P.O. Box 80218, Jeddah 21589, Saudi Arabia 6 School of Biotechnology, Eternal University, Baru Sahib-173101, Himachal Pradesh, India 7 Microbiology Unit, King Fahad Medical Research Center, King Abdulaziz University, P.O. Box 80216, Jeddah 21589, Saudi Arabia * Correspondence: [email protected] or [email protected]; Tel.: +966-533-381-010 Received: 19 May 2019; Accepted: 21 June 2019; Published: 26 June 2019 Abstract: Effective diagnostic, prognostic and therapeutic biomarkers can help in tracking disease progress, predict patients’ survival, and considerably affect the drive for successful clinical management. The present review aims to determine how the metastatic-linked protein anterior gradient homologue 2 (AGR2) operates to affect cancer progression, and to identify associated potential diagnostic, prognostic and therapeutic biomarkers, particularly in central nervous system (CNS) tumors. -

New Approach for Untangling the Role of Uncommon Calcium-Binding Proteins in the Central Nervous System
brain sciences Review New Approach for Untangling the Role of Uncommon Calcium-Binding Proteins in the Central Nervous System Krisztina Kelemen * and Tibor Szilágyi Department of Physiology, Doctoral School, Faculty of Medicine, George Emil Palade University of Medicine, Pharmacy, Science, and Technology of Targu Mures, 540142 Târgu Mures, , Romania; [email protected] * Correspondence: [email protected]; Tel.: +40-746-248064 Abstract: Although Ca2+ ion plays an essential role in cellular physiology, calcium-binding proteins (CaBPs) were long used for mainly as immunohistochemical markers of specific cell types in different regions of the central nervous system. They are a heterogeneous and wide-ranging group of proteins. Their function was studied intensively in the last two decades and a tremendous amount of informa- tion was gathered about them. Girard et al. compiled a comprehensive list of the gene-expression profiles of the entire EF-hand gene superfamily in the murine brain. We selected from this database those CaBPs which are related to information processing and/or neuronal signalling, have a Ca2+- buffer activity, Ca2+-sensor activity, modulator of Ca2+-channel activity, or a yet unknown function. In this way we created a gene function-based selection of the CaBPs. We cross-referenced these findings with publicly available, high-quality RNA-sequencing and in situ hybridization databases (Human Protein Atlas (HPA), Brain RNA-seq database and Allen Brain Atlas integrated into the HPA) and created gene expression heat maps of the regional and cell type-specific expression levels of the selected CaBPs. This represents a useful tool to predict and investigate different expression patterns and functions of the less-known CaBPs of the central nervous system. -

Egfrviii Promotes Cell Survival During Endoplasmic Reticulum Stress Through a Reticulocalbin 1-Dependent Mechanism
cancers Article EGFRvIII Promotes Cell Survival during Endoplasmic Reticulum Stress through a Reticulocalbin 1-Dependent Mechanism Juliana Gomez 1, Zammam Areeb 1, Sarah F. Stuart 1, Hong P. T. Nguyen 1, Lucia Paradiso 1, Ahmad Zulkifli 1 , Sonakshi Madan 1, Vijay Rajagopal 2 , Magdalene K. Montgomery 3, Hui K. Gan 4 , Andrew M. Scott 4, Jordan Jones 1,5, Andrew H. Kaye 1,6, Andrew P. Morokoff 1,5 and Rodney B. Luwor 1,* 1 Department of Surgery, The University of Melbourne, The Royal Melbourne Hospital, Parkville, VIC 3050, Australia; [email protected] (J.G.); [email protected] (Z.A.); [email protected] (S.F.S.); [email protected] (H.P.T.N.); [email protected] (L.P.); [email protected] (A.Z.); [email protected] (S.M.); [email protected] (J.J.); [email protected] (A.H.K.); [email protected] (A.P.M.) 2 Cell Structure and Mechanobiology Group, Department of Biomedical Engineering, Melbourne School of Engineering, The University of Melbourne, Parkville, VIC 3010, Australia; [email protected] 3 Department of Physiology, The University of Melbourne, Parkville, VIC 3010, Australia; [email protected] 4 Olivia Newton-John Cancer Research Institute, La Trobe University, Heidelberg, VIC 3084, Australia; [email protected] (H.K.G.); [email protected] (A.M.S.) 5 Department of Neurosurgery, The Royal Melbourne Hospital, Parkville, VIC 3050, Australia 6 Department of Neurosurgery, Hadassah Hebrew University Medical Centre, Jerusalem 91120, Israel * Correspondence: [email protected]; Tel.: +61-3-8344-3027; Fax: +61-393-476-488 Citation: Gomez, J.; Areeb, Z.; Stuart, S.F.; Nguyen, H.P.T.; Paradiso, L.; Simple Summary: A key molecule, EGFRvIII has been shown to provide several growth advantages Zulkifli, A.; Madan, S.; Rajagopal, V.; for brain tumors.