Rare and Low-Frequency Coding Variants Alter Human Adult Height
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

EGFR Phosphorylation of DCBLD2 Recruits TRAF6 and Stimulates AKT-Promoted Tumorigenesis
The Journal of Clinical Investigation RESEARCH ARTICLE EGFR phosphorylation of DCBLD2 recruits TRAF6 and stimulates AKT-promoted tumorigenesis Haizhong Feng,1,2 Giselle Y. Lopez,3 Chung Kwon Kim,2 Angel Alvarez,2 Christopher G. Duncan,3 Ryo Nishikawa,4 Motoo Nagane,5 An-Jey A. Su,6 Philip E. Auron,6 Matthew L. Hedberg,7 Lin Wang,7 Jeffery J. Raizer,2 John A. Kessler,2 Andrew T. Parsa,8 Wei-Qiang Gao,1 Sung-Hak Kim,9 Mutsuko Minata,9 Ichiro Nakano,9 Jennifer R. Grandis,7 Roger E. McLendon,3 Darell D. Bigner,3 Hui-Kuan Lin,10 Frank B. Furnari,11 Webster K. Cavenee,11 Bo Hu,2 Hai Yan,3 and Shi-Yuan Cheng1,2 1State Key Laboratory of Oncogenes and Related Genes, Renji-Med X Clinical Stem Cell Research Center, Ren Ji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai, China. 2Department of Neurology and Northwestern Brain Tumor Institute, Center for Genetic Medicine, Robert H. Lurie Comprehensive Cancer Center, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA. 3Pediatric Brain Tumor Foundation Institute at Duke, The Preston Robert Tisch Brain Tumor Center, and Department of Pathology, Duke University Medical Center, Durham, North Carolina, USA. 4Department of Neuro-Oncology/Neurosurgery, International Medical Center, Saitama Medical University, Saitama, Japan. 5Department of Neurosurgery, Kyorin University, Tokyo, Japan. 6Department of Biological Sciences, Duquesne University, Pittsburgh, Pennsylvania, USA. 7Departments of Otolaryngology and Pharmacology and Chemical Biology, University of Pittsburgh School of Medicine, Pittsburgh, Pennsylvania, USA. 8Department of Neurological Surgery and Northwestern Brain Tumor Institute, Robert H. Lurie Comprehensive Cancer Center, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA. -

Downloaded the “Top Edge” Version
bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Drosophila models of pathogenic copy-number variant genes show global and 2 non-neuronal defects during development 3 Short title: Non-neuronal defects of fly homologs of CNV genes 4 Tanzeen Yusuff1,4, Matthew Jensen1,4, Sneha Yennawar1,4, Lucilla Pizzo1, Siddharth 5 Karthikeyan1, Dagny J. Gould1, Avik Sarker1, Yurika Matsui1,2, Janani Iyer1, Zhi-Chun Lai1,2, 6 and Santhosh Girirajan1,3* 7 8 1. Department of Biochemistry and Molecular Biology, Pennsylvania State University, 9 University Park, PA 16802 10 2. Department of Biology, Pennsylvania State University, University Park, PA 16802 11 3. Department of Anthropology, Pennsylvania State University, University Park, PA 16802 12 4 contributed equally to work 13 14 *Correspondence: 15 Santhosh Girirajan, MBBS, PhD 16 205A Life Sciences Building 17 Pennsylvania State University 18 University Park, PA 16802 19 E-mail: [email protected] 20 Phone: 814-865-0674 21 1 bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 22 ABSTRACT 23 While rare pathogenic copy-number variants (CNVs) are associated with both neuronal and non- 24 neuronal phenotypes, functional studies evaluating these regions have focused on the molecular 25 basis of neuronal defects. -

EGFR Phosphorylation of DCBLD2 Recruits TRAF6 and Stimulates AKT-Promoted Tumorigenesis
EGFR phosphorylation of DCBLD2 recruits TRAF6 and stimulates AKT-promoted tumorigenesis Haizhong Feng, … , Hai Yan, Shi-Yuan Cheng J Clin Invest. 2014;124(9):3741-3756. https://doi.org/10.1172/JCI73093. Research Article Oncology Aberrant activation of EGFR in human cancers promotes tumorigenesis through stimulation of AKT signaling. Here, we determined that the discoidina neuropilin-like membrane protein DCBLD2 is upregulated in clinical specimens of glioblastomas and head and neck cancers (HNCs) and is required for EGFR-stimulated tumorigenesis. In multiple cancer cell lines, EGFR activated phosphorylation of tyrosine 750 (Y750) of DCBLD2, which is located within a recently identified binding motif for TNF receptor-associated factor 6 (TRAF6). Consequently, phosphorylation of DCBLD2 Y750 recruited TRAF6, leading to increased TRAF6 E3 ubiquitin ligase activity and subsequent activation of AKT, thereby enhancing EGFR-driven tumorigenesis. Moreover, evaluation of patient samples of gliomas and HNCs revealed an association among EGFR activation, DCBLD2 phosphorylation, and poor prognoses. Together, our findings uncover a pathway in which DCBLD2 functions as a signal relay for oncogenic EGFR signaling to promote tumorigenesis and suggest DCBLD2 and TRAF6 as potential therapeutic targets for human cancers that are associated with EGFR activation. Find the latest version: https://jci.me/73093/pdf The Journal of Clinical Investigation RESEARCH ARTICLE EGFR phosphorylation of DCBLD2 recruits TRAF6 and stimulates AKT-promoted tumorigenesis Haizhong Feng,1,2 Giselle Y. Lopez,3 Chung Kwon Kim,2 Angel Alvarez,2 Christopher G. Duncan,3 Ryo Nishikawa,4 Motoo Nagane,5 An-Jey A. Su,6 Philip E. Auron,6 Matthew L. Hedberg,7 Lin Wang,7 Jeffery J. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

A Homozygous Nonsense Mutation in DCBLD2 Is a Candidate Cause of Developmental Delay, Dysmorphic Features and Restrictive Cardiomyopathy Kheloud M
www.nature.com/scientificreports OPEN A homozygous nonsense mutation in DCBLD2 is a candidate cause of developmental delay, dysmorphic features and restrictive cardiomyopathy Kheloud M. Alhamoudi1, Tlili Barhoumi2, Hamad Al‑Eidi1, Abdulaziz Asiri3, Marwan Nashabat4, Manal Alaamery5, Masheal Alharbi1, Yazeid Alhaidan1, Brahim Tabarki6, Muhammad Umair1 & Majid Alfadhel1,4* DCBLD2 encodes discodin, CUB and LCCL domain‑containing protein 2, a type‑I transmembrane receptor that is involved in intracellular receptor signalling pathways and the regulation of cell growth. In this report, we describe a 5‑year‑old female who presented severe clinical features, including restrictive cardiomyopathy, developmental delay, spasticity and dysmorphic features. Trio‑ whole‑exome sequencing and segregation analysis were performed to identify the genetic cause of the disease within the family. A novel homozygous nonsense variant in the DCBLD2 gene (c.80G > A, p.W27*) was identifed as the most likely cause of the patient’s phenotype. This nonsense variant falls in the extracellular N‑terminus of DCBLD2 and thus might afect proper protein function of the transmembrane receptor. A number of in vitro investigations were performed on the proband’s skin fbroblasts compared to normal fbroblasts, which allowed a comprehensive assessment resulting in the functional characterization of the identifed DCBLD2 nonsense variant in diferent cellular processes. Our data propose a signifcant association between the identifed variant and the observed reduction in cell proliferation, -

Sun, B. B., Maranville, J. C., Peters, J. E., Stacey, D., Staley, J. R., Blackshaw, J., Burgess, S., Jiang, T., Paige, E., Suren
Sun, B. B., Maranville, J. C., Peters, J. E., Stacey, D., Staley, J. R., Blackshaw, J., Burgess, S., Jiang, T., Paige, E., Surendran, P., Oliver- Williams, C., Kamat, M. A., Prins, B. P., Wilcox, S. K., Zimmerman, E. S., Chi, A., Bansal, N., Spain, S. L., Wood, A. M., ... Butterworth, A. S. (2018). Genomic atlas of the human plasma proteome. Nature, 558(7708), 73-79. https://doi.org/10.1038/s41586-018-0175-2 Peer reviewed version Link to published version (if available): 10.1038/s41586-018-0175-2 Link to publication record in Explore Bristol Research PDF-document This is the author accepted manuscript (AAM). The final published version (version of record) is available online via Nature Publishing Group at https://www.nature.com/articles/s41586-018-0175-2 . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/red/research-policy/pure/user-guides/ebr-terms/ Genomic atlas of the human plasma proteome Benjamin B. Sun1*, Joseph C. Maranville2*, James E. Peters1,3*, David Stacey1, James R. Staley1, James Blackshaw1, Stephen Burgess1,4, Tao Jiang1, Ellie Paige1,5, Praveen Surendran1, Clare Oliver-Williams1,6, Mihir A. Kamat1, Bram P. Prins1, Sheri K. Wilcox7, Erik S. Zimmerman7, An Chi2, Narinder Bansal1,8, Sarah L. Spain9, Angela M. Wood1, Nicholas W. Morrell10, John R. Bradley11, Nebojsa Janjic7, David J. Roberts12,13, Willem H. -

Mouse Dcbld2 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Dcbld2 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Dcbld2 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Dcbld2 gene (NCBI Reference Sequence: NM_028523 ; Ensembl: ENSMUSG00000035107 ) is located on Mouse chromosome 16. 16 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 16 (Transcript: ENSMUST00000046663). Exon 5~7 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Dcbld2 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-86O4 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a knock-out allele exhibit reduced postnatal angiogenesis and impaired recovery from femoral artery ligation with impaired blood flow and decreased capillary density. Exon 5 starts from about 26.66% of the coding region. The knockout of Exon 5~7 will result in frameshift of the gene. The size of intron 4 for 5'-loxP site insertion: 3059 bp, and the size of intron 7 for 3'-loxP site insertion: 884 bp. The size of effective cKO region: ~1888 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 5 6 7 8 9 16 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Dcbld2 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -
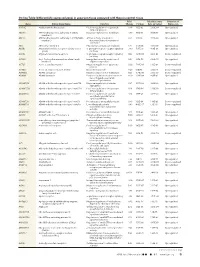
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

DCBLD2 (NM 080927) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC224483L4 DCBLD2 (NM_080927) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: DCBLD2 (NM_080927) Human Tagged ORF Clone Tag: mGFP Symbol: DCBLD2 Synonyms: CLCP1; ESDN Vector: pLenti-C-mGFP-P2A-Puro (PS100093) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(RC224483). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_080927 ORF Size: 2325 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 DCBLD2 (NM_080927) Human Tagged ORF Clone – RC224483L4 OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_080927.3 RefSeq Size: 6093 bp RefSeq ORF: 2328 bp Locus ID: 131566 UniProt ID: Q96PD2 Domains: F5_F8_type_C, CUB, LCCL Protein Families: Druggable Genome, Transmembrane MW: 84.9 kDa This product is to be used for laboratory only. -

Dcbld2/Esdn Is Essential for Proper Optic Tract Formation and Retinal Lamination Ryan Mears Joy University of Vermont
University of Vermont ScholarWorks @ UVM Graduate College Dissertations and Theses Dissertations and Theses 2016 Dcbld2/esdn Is Essential For Proper Optic Tract Formation And Retinal Lamination Ryan Mears Joy University of Vermont Follow this and additional works at: https://scholarworks.uvm.edu/graddis Part of the Cell and Developmental Biology Commons Recommended Citation Joy, Ryan Mears, "Dcbld2/esdn Is Essential For Proper Optic Tract Formation And Retinal Lamination" (2016). Graduate College Dissertations and Theses. 563. https://scholarworks.uvm.edu/graddis/563 This Thesis is brought to you for free and open access by the Dissertations and Theses at ScholarWorks @ UVM. It has been accepted for inclusion in Graduate College Dissertations and Theses by an authorized administrator of ScholarWorks @ UVM. For more information, please contact [email protected]. DCBLD2/ESDN IS ESSENTIAL FOR PROPER OPTIC TRACT FORMATION AND RETINAL LAMINATION A Thesis Presented by Ryan Joy to The Faculty of the Graduate College of The University of Vermont In Partial Fulfillment of the Requirements for the Degree of Master of Science Specializing in Biology May, 2016 Defense Date: January 7, 2016 Thesis Examination Committee: Bryan A. Ballif, Ph.D., Advisor Jason Stumpff, Ph.D., Chairperson Alicia M. Ebert, Ph.D. Paula Deming, Ph.D., MT Cynthia J. Forehand, Ph.D., Dean of the Graduate College ABSTRACT The Discoidin, CUB and LCCL domain-containing protein 2 (DCBLD2/ESDN/CLCP1) is a type-I, transmembrane receptor that mediates diverse cellular processes such as angiogenesis, vascular remodeling, cellular migration and proliferation. Identification of DCBLD2 in a proteomics screen to identify substrates of Src family tyrosine kinases that bind the Src homology 2 domain of CT10 regulator of kinase-Like (CrkL), a critical scaffolding protein for neuronal development, led to a need for further characterization of the protein. -

Systems Genetics Identifies Sestrin 3 As a Regulator of a Proconvulsant Gene Network in Human Epileptic Hippocampus
ARTICLE Received 1 Sep 2014 | Accepted 4 Dec 2014 | Published 23 Jan 2015 DOI: 10.1038/ncomms7031 Systems genetics identifies Sestrin 3 as a regulator of a proconvulsant gene network in human epileptic hippocampus Michael R. Johnson1,y, Jacques Behmoaras2,*, Leonardo Bottolo3,*, Michelle L. Krishnan4,*, Katharina Pernhorst5,*, Paola L. Meza Santoscoy6,*, Tiziana Rossetti7, Doug Speed8, Prashant K. Srivastava1,7, Marc Chadeau-Hyam9, Nabil Hajji10, Aleksandra Dabrowska10, Maxime Rotival7, Banafsheh Razzaghi7, Stjepana Kovac11, Klaus Wanisch11, Federico W. Grillo7, Anna Slaviero7, Sarah R. Langley1,7, Kirill Shkura1,7, Paolo Roncon12,13, Tisham De7, Manuel Mattheisen14,15,16, Pitt Niehusmann5, Terence J. O’Brien17, Slave Petrovski18, Marec von Lehe19, Per Hoffmann20,21, Johan Eriksson22,23,24, Alison J. Coffey25, Sven Cichon20,21, Matthew Walker11, Michele Simonato12,13,26,Be´ne´dicte Danis27, Manuela Mazzuferi27, Patrik Foerch27, Susanne Schoch5,28, Vincenzo De Paola7, Rafal M. Kaminski27, Vincent T. Cunliffe6, Albert J. Becker5,y & Enrico Petretto7,29,y Gene-regulatory network analysis is a powerful approach to elucidate the molecular processes and pathways underlying complex disease. Here we employ systems genetics approaches to characterize the genetic regulation of pathophysiological pathways in human temporal lobe epilepsy (TLE). Using surgically acquired hippocampi from 129 TLE patients, we identify a gene-regulatory network genetically associated with epilepsy that contains a specialized, highly expressed transcriptional module encoding proconvulsive cytokines and Toll-like receptor signalling genes. RNA sequencing analysis in a mouse model of TLE using 100 epileptic and 100 control hippocampi shows the proconvulsive module is preserved across-species, specific to the epileptic hippocampus and upregulated in chronic epilepsy.