Global Regulation of Differential Gene Expression by C-Abl/Arg Oncogenic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

PTPRA Phosphatase Regulates GDNF-Dependent RET Signaling and Inhibits the RET Mutant MEN2A Oncogenic Potential
Journal Pre-proof PTPRA phosphatase regulates GDNF-dependent RET signaling and inhibits the RET mutant MEN2A oncogenic potential Leena Yadav, Elina Pietilä, Tiina Öhman, Xiaonan Liu, Arun K. Mahato, Yulia Sidorova, Kaisa Lehti, Mart Saarma, Markku Varjosalo PII: S2589-0042(20)30055-9 DOI: https://doi.org/10.1016/j.isci.2020.100871 Reference: ISCI 100871 To appear in: ISCIENCE Received Date: 3 August 2019 Revised Date: 15 January 2020 Accepted Date: 26 January 2020 Please cite this article as: Yadav, L., Pietilä, E., Öhman, T., Liu, X., Mahato, A.K., Sidorova, Y., Lehti, K., Saarma, M., Varjosalo, M., PTPRA phosphatase regulates GDNF-dependent RET signaling and inhibits the RET mutant MEN2A oncogenic potential, ISCIENCE (2020), doi: https://doi.org/10.1016/ j.isci.2020.100871. This is a PDF file of an article that has undergone enhancements after acceptance, such as the addition of a cover page and metadata, and formatting for readability, but it is not yet the definitive version of record. This version will undergo additional copyediting, typesetting and review before it is published in its final form, but we are providing this version to give early visibility of the article. Please note that, during the production process, errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. © 2020 Growth factor RET PTPRA Cell surface Ras P P Complex formation RAF MEK ERK Growth Nucleus Proliferation Gene expression Migration 1 PTPRA phosphatase regulates GDNF-dependent RET signaling and inhibits the RET mutant MEN2A oncogenic potential Authors Leena Yadav 1, Elina Pietilä 3# , Tiina Öhman 1# , Xiaonan Liu 1, Arun K. -
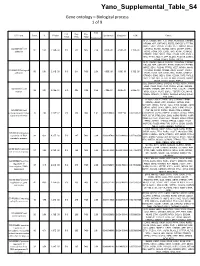
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Ptk7-Deficient Mice Have Decreased Hematopoietic Stem Cell Pools As a Result of Deregulated Proliferation and Migration
Ptk7-Deficient Mice Have Decreased Hematopoietic Stem Cell Pools as a Result of Deregulated Proliferation and Migration This information is current as Anne-Catherine Lhoumeau, Marie-Laure Arcangeli, Maria of September 24, 2021. De Grandis, Marilyn Giordano, Jean-Christophe Orsoni, Frédérique Lembo, Florence Bardin, Sylvie Marchetto, Michel Aurrand-Lions and Jean-Paul Borg J Immunol 2016; 196:4367-4377; Prepublished online 18 April 2016; Downloaded from doi: 10.4049/jimmunol.1500680 http://www.jimmunol.org/content/196/10/4367 Supplementary http://www.jimmunol.org/content/suppl/2016/04/16/jimmunol.150068 http://www.jimmunol.org/ Material 0.DCSupplemental References This article cites 55 articles, 24 of which you can access for free at: http://www.jimmunol.org/content/196/10/4367.full#ref-list-1 Why The JI? Submit online. by guest on September 24, 2021 • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright -

Functional Analysis of Somatic Mutations Affecting Receptor Tyrosine Kinase Family in Metastatic Colorectal Cancer
Author Manuscript Published OnlineFirst on March 29, 2019; DOI: 10.1158/1535-7163.MCT-18-0582 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Functional analysis of somatic mutations affecting receptor tyrosine kinase family in metastatic colorectal cancer Leslie Duplaquet1, Martin Figeac2, Frédéric Leprêtre2, Charline Frandemiche3,4, Céline Villenet2, Shéhérazade Sebda2, Nasrin Sarafan-Vasseur5, Mélanie Bénozène1, Audrey Vinchent1, Gautier Goormachtigh1, Laurence Wicquart6, Nathalie Rousseau3, Ludivine Beaussire5, Stéphanie Truant7, Pierre Michel8, Jean-Christophe Sabourin9, Françoise Galateau-Sallé10, Marie-Christine Copin1,6, Gérard Zalcman11, Yvan De Launoit1, Véronique Fafeur1 and David Tulasne1 1 Univ. Lille, CNRS, Institut Pasteur de Lille, UMR 8161 - M3T – Mechanisms of Tumorigenesis and Target Therapies, F-59000 Lille, France. 2 Univ. Lille, Plateau de génomique fonctionnelle et structurale, CHU Lille, F-59000 Lille, France 3 TCBN - Tumorothèque Caen Basse-Normandie, F-14000 Caen, France. 4 Réseau Régional de Cancérologie – OncoBasseNormandie – F14000 Caen – France. 5 Normandie Univ, UNIROUEN, Inserm U1245, IRON group, Rouen University Hospital, Normandy Centre for Genomic and Personalized Medicine, F-76000 Rouen, France. 6 Tumorothèque du C2RC de Lille, F-59037 Lille, France. 7 Department of Digestive Surgery and Transplantation, CHU Lille, Univ Lille, 2 Avenue Oscar Lambret, 59037, Lille Cedex, France. 8 Department of hepato-gastroenterology, Rouen University Hospital, Normandie Univ, UNIROUEN, Inserm U1245, IRON group, F-76000 Rouen, France. 9 Department of Pathology, Normandy University, INSERM 1245, Rouen University Hospital, F 76 000 Rouen, France. 10 Department of Pathology, MESOPATH-MESOBANK, Centre León Bérard, Lyon, France. 11 Thoracic Oncology Department, CIC1425/CLIP2 Paris-Nord, Hôpital Bichat-Claude Bernard, Paris, France. -
HCC and Cancer Mutated Genes Summarized in the Literature Gene Symbol Gene Name References*
HCC and cancer mutated genes summarized in the literature Gene symbol Gene name References* A2M Alpha-2-macroglobulin (4) ABL1 c-abl oncogene 1, receptor tyrosine kinase (4,5,22) ACBD7 Acyl-Coenzyme A binding domain containing 7 (23) ACTL6A Actin-like 6A (4,5) ACTL6B Actin-like 6B (4) ACVR1B Activin A receptor, type IB (21,22) ACVR2A Activin A receptor, type IIA (4,21) ADAM10 ADAM metallopeptidase domain 10 (5) ADAMTS9 ADAM metallopeptidase with thrombospondin type 1 motif, 9 (4) ADCY2 Adenylate cyclase 2 (brain) (26) AJUBA Ajuba LIM protein (21) AKAP9 A kinase (PRKA) anchor protein (yotiao) 9 (4) Akt AKT serine/threonine kinase (28) AKT1 v-akt murine thymoma viral oncogene homolog 1 (5,21,22) AKT2 v-akt murine thymoma viral oncogene homolog 2 (4) ALB Albumin (4) ALK Anaplastic lymphoma receptor tyrosine kinase (22) AMPH Amphiphysin (24) ANK3 Ankyrin 3, node of Ranvier (ankyrin G) (4) ANKRD12 Ankyrin repeat domain 12 (4) ANO1 Anoctamin 1, calcium activated chloride channel (4) APC Adenomatous polyposis coli (4,5,21,22,25,28) APOB Apolipoprotein B [including Ag(x) antigen] (4) AR Androgen receptor (5,21-23) ARAP1 ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 (4) ARHGAP35 Rho GTPase activating protein 35 (21) ARID1A AT rich interactive domain 1A (SWI-like) (4,5,21,22,24,25,27,28) ARID1B AT rich interactive domain 1B (SWI1-like) (4,5,22) ARID2 AT rich interactive domain 2 (ARID, RFX-like) (4,5,22,24,25,27,28) ARID4A AT rich interactive domain 4A (RBP1-like) (28) ARID5B AT rich interactive domain 5B (MRF1-like) (21) ASPM Asp (abnormal -

Enhancing the Thermostability and Activity of Uronate Dehydrogenase from Agrobacterium Tumefaciens LBA4404 by Semi-Rational Engi
Su et al. Bioresour. Bioprocess. (2019) 6:36 https://doi.org/10.1186/s40643-019-0267-3 RESEARCH Open Access Enhancing the thermostability and activity of uronate dehydrogenase from Agrobacterium tumefaciens LBA4404 by semi-rational engineering Hui‑Hui Su1†, Fei Peng1†, Pei Xu1, Xiao‑Ling Wu1, Min‑Hua Zong1, Ji‑Guo Yang2 and Wen‑Yong Lou1* Abstract Background: Glucaric acid, one of the aldaric acids, has been declared a “top value‑added chemical from biomass”, and is especially important in the food and pharmaceutical industries. Biocatalytic production of glucaric acid from glucuronic acid is more environmentally friendly, efcient and economical than chemical synthesis. Uronate dehydro‑ genases (UDHs) are the key enzymes for the preparation of glucaric acid in this way, but the poor thermostability and low activity of UDH limit its industrial application. Therefore, improving the thermostability and activity of UDH, for example by semi‑rational design, is a major research goal. Results: In the present work, three UDHs were obtained from diferent Agrobacterium tumefaciens strains. The three UDHs have an approximate molecular weight of 32 kDa and all contain typically conserved UDH motifs. All three UDHs showed optimal activity within a pH range of 6.0–8.5 and at a temperature of 30 °C, but the UDH from A. tume- 1 1 faciens (At) LBA4404 had a better catalytic efciency than the other two UDHs (800 vs 600 and 530 s− mM− ). To fur‑ ther boost the catalytic performance of the UDH from AtLBA4404, site‑directed mutagenesis based on semi‑rational design was carried out. An A39P/H99Y/H234K triple mutant showed a 400‑fold improvement in half‑life at 59 °C, a 10 5 °C improvement in T50 value and a 2.5‑fold improvement in specifc activity at 30 °C compared to wild‑type UDH. -

Wnt5a Modulates Integrin Expression in a Receptor-Dependent Manner In
www.nature.com/scientificreports OPEN Wnt5A modulates integrin expression in a receptor‑dependent manner in ovarian cancer cells Vajihe Azimian‑Zavareh 1,2, Zeinab Dehghani‑Ghobadi1, Marzieh Ebrahimi 3*, Kian Mirzazadeh1, Irina Nazarenko4 & Ghamartaj Hossein 1,4* Wnt5A signals through various receptors that confer versatile biological functions. Here, we used Wnt5A overexpressing human ovarian SKOV‑3 and OVCAR‑3 stable clones for assessing integrin expression, cell proliferation, migration, invasion, and the ability of multicellular aggregates (MCAs) formation. We found here, that Wnt5A regulates diferently the expression of its receptors in the stable Wnt5A overexpressing clones. The expression levels of Frizzled (FZD)‑2 and ‑5, were increased in diferent clones. However ROR‑1, ‑2 expression levels were diferently regulated in clones. Wnt5A overexpressing clones showed increased cell proliferation, migration, and clonogenicity. Moreover, Wnt5A overexpressing SKOV‑3 clone showed increased MCAs formation ability. Cell invasion had been increased in OVCAR‑3‑derived clones, while this was decreased in SKOV‑3‑derived clone. Importantly, αv integrin expression levels were increased in all assessed clones, accompanied by increased cell attachment to fbronectin and focal adhesion kinase activity. Moreover, the treatment of clones with Box5 as a Wnt5A/FZD5 antagonist abrogates ITGAV increase, cell proliferation, migration, and their attachment to fbronectin. Accordingly, we observed signifcantly higher expression levels of ITGAV and ITGB3 in human high‑grade serous ovarian cancer specimens and ITGAV correlated positively with Wnt5A in metastatic serous type ovarian cancer. In summary, we hypothesize here, that Wnt5A/FZD‑5 signaling modulate αv integrin expression levels that could be associated with ovarian cancer cell proliferation, migration, and fbronectin attachment. -
Loss of Receptor Tyrosine Kinase-Like Orphan Receptor 2 Impairs The
Lei et al. Stem Cell Research & Therapy (2020) 11:137 https://doi.org/10.1186/s13287-020-01646-2 RESEARCH Open Access Loss of receptor tyrosine kinase-like orphan receptor 2 impairs the osteogenesis of mBMSCs by inhibiting signal transducer and activator of transcription 3 Lizhen Lei1,2, Zhuwei Huang1,2, Jingyi Feng1,2, Zijing Huang1,2, Yiwei Tao1,2, Xiaoli Hu1,2* and Xiaolei Zhang1,2* Abstract Background: Receptor tyrosine kinase-like orphan receptor 2 (Ror2) plays a key role in bone formation, but its signaling pathway is not completely understood. Signal transducer and activator of transcription 3 (Stat3) takes part in maintaining bone homeostasis. The aim of this study is to reveal the role and mechanism of Ror2 in the osteogenic differentiation from mouse bone marrow mesenchymal stem cells (mBMSCs) and to explore the effect of Stat3 on Ror2-mediated osteogenesis. Methods: Ror2 CKO mice were generated via the Cre-loxp recombination system using Prrx1-Cre transgenic mice. Quantitative real-time PCR and western blot were performed to assess the expression of Stat3 and osteogenic markers in Ror2-knockdown mBMSCs (mBMSC-sh-Ror2). After being incubated in osteogenic induction medium for 3 weeks, Alizarin Red staining and western blot were used to examine the calcium deposit and osteogenic markers in Stat3 overexpression in mBMSC-sh-Ror2. Results: Loss of Ror2 in mesenchymal or osteoblast progenitor cells led to a dwarfism phenotype in vivo. The mRNA expression of osteogenic markers (osteocalcin, osteopontin (OPN), and collagen I) in the ulna proximal epiphysis of Ror2 CKO mice was significantly decreased (P < 0.05). -

Ror2 Functions As a Noncanonical Wnt Receptor That Regulates NMDAR-Mediated Synaptic Transmission
RoR2 functions as a noncanonical Wnt receptor that regulates NMDAR-mediated synaptic transmission Waldo Cerpaa,1, Elena Latorre-Estevesa,b, and Andres Barriaa,2 aDepartment of Physiology and Biophysics and bMolecular and Cellular Biology Program, University of Washington, Seattle, WA 98195 Edited* by Richard L. Huganir, The Johns Hopkins University School of Medicine, Baltimore, MD, and approved March 6, 2015 (received for review September 16, 2014) Wnt signaling has a well-established role as a regulator of nervous receptor for noncanonical Wnt ligands capable of regulating syn- system development, but its role in the maintenance and regula- aptic NMDARs. In hippocampal neurons, activation of RoR2 by tion of established synapses in the mature brain remains poorly noncanonical Wnt ligand Wnt5a activates PKC and JNK, two understood. At excitatory glutamatergic synapses, NMDA receptors kinases involved in the regulation of NMDAR currents. In ad- (NMDARs) have a fundamental role in synaptogenesis, synaptic dition, we show that signaling through RoR2 is necessary for plasticity, and learning and memory; however, it is not known what the maintenance of basal NMDAR-mediated synaptic trans- controls their number and subunit composition. Here we show that mission and the acute regulation of NMDAR synaptic responses the receptor tyrosine kinase-like orphan receptor 2 (RoR2) func- by Wnt5a. tions as a Wnt receptor required to maintain basal NMDAR- Identification of RoR2 as a Wnt receptor that regulates syn- aptic NMDARs provides a mechanism for Wnt signaling to mediated synaptic transmission. In addition, RoR2 activation by a control synaptic transmission and synaptic plasticity acutely, and noncanonical Wnt ligand activates PKC and JNK and acutely en- is a critical first step toward understanding the role played by hances NMDAR synaptic responses. -

Functional and Physiological Discovery in the Mannonate Dehydratase Subgroup of the Enolase Superfamily
FUNCTIONAL AND PHYSIOLOGICAL DISCOVERY IN THE MANNONATE DEHYDRATASE SUBGROUP OF THE ENOLASE SUPERFAMILY BY DANIEL JOSEPH WICHELECKI DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biochemistry in the Graduate College of the University of Illinois at Urbana-Champaign, 2014 Urbana, Illinois Doctoral Committee: Professor John Gerlt, Chair Professor John Cronan Professor Scott Silverman Professor Wilfred van der Donk ABSTRACT In the current post-genomic world, the exponential amassing of protein sequences is overwhelming the scientific community’s ability to experimentally assign each protein’s function. The use of automated, homology-based annotations has allowed a reprieve from this efflux of data, but has led to widespread misannotation and nonannotation in protein sequence databases. This dissertation details the functional and physiological characterization of the mannonate dehydratase subgroup (ManD) of the enolase superfamily (ENS). The outcome affirms the dangers of homology-based annotations while discovering novel metabolic pathways. Furthermore, the experimental verification of these pathways ( in vitro and in vivo ) has provided a platform to test the general strategies for improved functional and metabolic characterization being developed by the Enzyme Function Initiative (EFI). Prior to this study, one member of the ManD subgroup had been characterized and was shown to dehydrate D-mannonate to 2-keto-3-deoxy-D-gluconate. Forty-two additional members of the ManD, selected from across the sequence space of the subgroup, were screened for activity and kinetic constants were determined. The members of the once isofunctional subgroup were found to differ in both catalytic efficiency and substrate specificity: 1) high 3 4 -1 -1 efficiency (k cat /K M = 10 to 10 M s ) dehydration of D-mannonate, 2) low efficiency (k cat /K M = 10 1 to 10 2 M-1s-1) dehydration of D-mannonate and/or D-gluconate, and 3) no-activity with either D-mannonate or D-gluconate (or any other acid sugar tested). -

The Regulation of Human Carbonyl Reductase 3 (CBR3) in Epithelial Cell Lines
Charles university in Prague Faculty of pharmacy in Hradec Kralov´ e´ Department of biochemical sciences Diploma thesis The regulation of human carbonyl reductase 3 (CBR3) in epithelial cell lines Regulace lidske´ karbonylreduktasy 3 v buneˇcnˇ ych´ lini´ıch epithelu Petra Malatkov´ a´ 2009 Supervisors: Prof. Dr. Edmund Maser Dr. Bettina Ebert Christian Albrechts University in Kiel Faculty of Medicine Department of Toxicology and Pharmacology for Natural Scientists Prof. Ing. Vladim´ır Wsol,´ Ph.D. Charles University in Prague Faculty of Pharmacy in Hradec Kralov´ e´ Department of Biochemical Sciences Declaration I declare that this thesis is my original author’s work which I have worked out independently. All literature and other sources, from which I gather during elaboration, are listed in the bibli- ography and properly cited in the work. 4th May 2009 2 Acknowledgements First, I would like to express sincere thanks to Dr. Bettina Ebert for teaching me new techniques and performing cell culture experiments. I thank you for lending your time and providing de- tailed comments. Also, I would like to thank to Prof. Dr. Edmund Maser and Prof. Ing. Vladim´ır Wsol,´ Ph.D. for offering useful advice and, in particular, giving me chance to do the diploma thesis at Department of Toxicology and Pharmacology in Kiel. Next, I would like to thank to Yasser El-Hawari, who helps me with LaTex and contributes to improving my German skills. I would also like to thank to Michael Kisiela for his helpful assistance with any occurred problems. Finally, I would like to extend a big thank to my parents for their love and support. -

ROR2 Is Epigenetically Inactivated in the Early Stages of Colorectal Neoplasia and Is Associated with Proliferation and Migration Sean S
Ma et al. BMC Cancer (2016) 16:508 DOI 10.1186/s12885-016-2576-7 RESEARCH ARTICLE Open Access ROR2 is epigenetically inactivated in the early stages of colorectal neoplasia and is associated with proliferation and migration Sean S. Q. Ma1, Sameer Srivastava2,3, Estelle Llamosas1, Nicholas J. Hawkins4, Luke B. Hesson2, Robyn L. Ward2 and Caroline E. Ford1* Abstract Background: Colorectal cancer (CRC) is closely linked to Wnt signalling, with 94 % of cases exhibiting a Wnt related mutation. ROR2 is a receptor tyrosine kinase that is thought to repress β-catenin dependent Wnt signalling. Our study aims to determine if ROR2 is epigenetically silenced in CRC and determine if in vitro silencing of ROR2 potentiates Wnt signalling, and alters the proliferative, migratory or invasive potential of cells. Methods: ROR2 expression was examined in CRC cell lines and patient adenomas using qRT-PCR, while COBRA and bisulphite sequencing was used to analyse ROR2 promoter methylation. 258 patient primary tumour samples from publicly available databases were also examined for ROR2 expression and methylation. In addition, the functional effects of ROR2 modulation were investigated in HCT116 cells following ROR2 siRNA knockdown and in RKO and SW620 cells following ectopic ROR2 expression. Results: Reduced ROR2 expression was found to correlate with ROR2 promoter hypermethylation in colorectal cancer cell lines, carcinomas and adenomas. ROR2 expression was downregulated in 76.7 % (23/30) of CRC cell lines with increasing ROR2 promoter hypermethylation correlating with progressively lower expression. Analysis of 239 primary tumour samples from a publicly available cohort also found a significant correlation between reduced ROR2 expression and increased promoter methylation.