Splicing Regulatory Factors in Breast Cancer Hallmarks and Disease Progression
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

SRC Antibody - N-Terminal Region (ARP32476 P050) Data Sheet
SRC antibody - N-terminal region (ARP32476_P050) Data Sheet Product Number ARP32476_P050 Product Name SRC antibody - N-terminal region (ARP32476_P050) Size 50ug Gene Symbol SRC Alias Symbols ASV; SRC1; c-SRC; p60-Src Nucleotide Accession# NM_005417 Protein Size (# AA) 536 amino acids Molecular Weight 60kDa Product Format Lyophilized powder NCBI Gene Id 6714 Host Rabbit Clonality Polyclonal Official Gene Full Name V-src sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog (avian) Gene Family SH2D This is a rabbit polyclonal antibody against SRC. It was validated on Western Blot by Aviva Systems Biology. At Aviva Systems Biology we manufacture rabbit polyclonal antibodies on a large scale (200-1000 Description products/month) of high throughput manner. Our antibodies are peptide based and protein family oriented. We usually provide antibodies covering each member of a whole protein family of your interest. We also use our best efforts to provide you antibodies recognize various epitopes of a target protein. For availability of antibody needed for your experiment, please inquire (). Peptide Sequence Synthetic peptide located within the following region: QTPSKPASADGHRGPSAAFAPAAAEPKLFGGFNSSDTVTSPQRAGPLAGG This gene is highly similar to the v-src gene of Rous sarcoma virus. This proto-oncogene may play a role in the Description of Target regulation of embryonic development and cell growth. SRC protein is a tyrosine-protein kinase whose activity can be inhibited by phosphorylation by c-SRC kinase. Mutations in this gene could be involved in the -

University of Groningen BRCA2 Deficiency Instigates Cgas
University of Groningen BRCA2 deficiency instigates cGAS-mediated inflammatory signaling and confers sensitivity to tumor necrosis factor-alpha-mediated cytotoxicity Heijink, Anne Margriet; Talens, Francien; Jae, Lucas T; van Gijn, Stephanie E; Fehrmann, Rudolf S N; Brummelkamp, Thijn R; van Vugt, Marcel A T M Published in: Nature Communications DOI: 10.1038/s41467-018-07927-y IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version below. Document Version Publisher's PDF, also known as Version of record Publication date: 2019 Link to publication in University of Groningen/UMCG research database Citation for published version (APA): Heijink, A. M., Talens, F., Jae, L. T., van Gijn, S. E., Fehrmann, R. S. N., Brummelkamp, T. R., & van Vugt, M. A. T. M. (2019). BRCA2 deficiency instigates cGAS-mediated inflammatory signaling and confers sensitivity to tumor necrosis factor-alpha-mediated cytotoxicity. Nature Communications, 10(1), [100]. https://doi.org/10.1038/s41467-018-07927-y Copyright Other than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons). The publication may also be distributed here under the terms of Article 25fa of the Dutch Copyright Act, indicated by the “Taverne” license. More information can be found on the University of Groningen website: https://www.rug.nl/library/open-access/self-archiving-pure/taverne- amendment. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
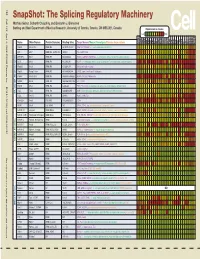
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

RET Gene Fusions in Malignancies of the Thyroid and Other Tissues
G C A T T A C G G C A T genes Review RET Gene Fusions in Malignancies of the Thyroid and Other Tissues Massimo Santoro 1,*, Marialuisa Moccia 1, Giorgia Federico 1 and Francesca Carlomagno 1,2 1 Department of Molecular Medicine and Medical Biotechnology, University of Naples “Federico II”, 80131 Naples, Italy; [email protected] (M.M.); [email protected] (G.F.); [email protected] (F.C.) 2 Institute of Endocrinology and Experimental Oncology of the CNR, 80131 Naples, Italy * Correspondence: [email protected] Received: 10 March 2020; Accepted: 12 April 2020; Published: 15 April 2020 Abstract: Following the identification of the BCR-ABL1 (Breakpoint Cluster Region-ABelson murine Leukemia) fusion in chronic myelogenous leukemia, gene fusions generating chimeric oncoproteins have been recognized as common genomic structural variations in human malignancies. This is, in particular, a frequent mechanism in the oncogenic conversion of protein kinases. Gene fusion was the first mechanism identified for the oncogenic activation of the receptor tyrosine kinase RET (REarranged during Transfection), initially discovered in papillary thyroid carcinoma (PTC). More recently, the advent of highly sensitive massive parallel (next generation sequencing, NGS) sequencing of tumor DNA or cell-free (cfDNA) circulating tumor DNA, allowed for the detection of RET fusions in many other solid and hematopoietic malignancies. This review summarizes the role of RET fusions in the pathogenesis of human cancer. Keywords: kinase; tyrosine kinase inhibitor; targeted therapy; thyroid cancer 1. The RET Receptor RET (REarranged during Transfection) was initially isolated as a rearranged oncoprotein upon the transfection of a human lymphoma DNA [1]. -

Gorilla Results
Results http://cbl-gorilla.cs.technion.ac.il/GOrilla/2l4v7a1a/GOResultsPROCESS... P-value color scale > 10-3 10-3 to 10-5 10-5 to 10-7 10-7 to 10-9 < 10-9 null Description P-value Enrichment (N, B, n, b) Genes [-] Hide genes ATF3 - activating transcription factor 3 JUN - jun proto-oncogene ADM - adrenomedullin NAMPT - nicotinamide phosphoribosyltransferase GO:0008284 positive regulation of cell proliferation 6.46E-6 4.37 (297,17,40,10) C7orf68 - chromosome 7 open reading frame 68 FOSL2 - fos-like antigen 2 TNFAIP3 - tumor necrosis factor, alpha-induced protein 3 VEGFA - vascular endothelial growth factor a SOX9 - sry (sex determining region y)-box 9 EDN2 - endothelin 2 [-] Hide genes JUN - jun proto-oncogene SPRR1B - small proline-rich protein 1b GO:0030855 epithelial cell differentiation 4.33E-5 5.20 (297,10,40,7) SPRR1A - small proline-rich protein 1a PLAUR - plasminogen activator, urokinase receptor VEGFA - vascular endothelial growth factor a TXNIP - thioredoxin interacting protein ELF3 - e74-like factor 3 (ets domain transcription factor, epithelial-specific ) [-] Hide genes JUN - jun proto-oncogene KHDRBS1 - kh domain containing, rna binding, signal transduction associated 1 MAFF - v-maf musculoaponeurotic fibrosarcoma oncogene homolog f (avian) VEGFA - vascular endothelial growth factor a RRAS - related ras viral (r-ras) oncogene homolog SOX9 - sry (sex determining region y)-box 9 SAT1 - spermidine/spermine n1-acetyltransferase 1 GEM - gtp binding protein overexpressed in skeletal muscle GO:0006807 nitrogen compound metabolic process -

Flexible, Unbiased Analysis of Biological Characteristics Associated with Genomic Regions
bioRxiv preprint doi: https://doi.org/10.1101/279612; this version posted March 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. BioFeatureFinder: Flexible, unbiased analysis of biological characteristics associated with genomic regions Felipe E. Ciamponi 1,2,3; Michael T. Lovci 2; Pedro R. S. Cruz 1,2; Katlin B. Massirer *,1,2 1. Structural Genomics Consortium - SGC, University of Campinas, SP, Brazil. 2. Center for Molecular Biology and Genetic Engineering - CBMEG, University of Campinas, Campinas, SP, Brazil. 3. Graduate program in Genetics and Molecular Biology, PGGBM, University of Campinas, Campinas, SP, Brazil. *Corresponding author: [email protected] Mailing address: Center for Molecular Biology and Genetic Engineering - CBMEG, University of Campinas, Campinas, SP, Brazil. Av Candido Rondo, 400 Cidade Universitária CEP 13083-875, Campinas, SP Phone: 55-19-98121-937 bioRxiv preprint doi: https://doi.org/10.1101/279612; this version posted March 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Abstract BioFeatureFinder is a novel algorithm which allows analyses of many biological genomic landmarks (including alternatively spliced exons, DNA/RNA- binding protein binding sites, and gene/transcript functional elements, nucleotide content, conservation, k-mers, secondary structure) to identify distinguishing features. -
Supplementary Data Vigneswaran Et Al Supplementary Data
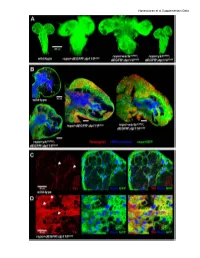
Vigneswaran et al Supplementary Data Vigneswaran et al Supplementary Data Figure S1: Yki is required for EGFR-PI3K-driven glial neoplasia in Drosophila (A) Optical projections of whole brain-nerve cord complexes from 3rd instar larvae approximately 130 hrs old. Dorsal view; anterior up. CD8-GFP (green) labels glial cell bodies. Compared to repo>dEGFRλ;dp110CAAX, warts knockdown (repo>wartsdsRNA; dEGFRλ;dp110CAAX) increased neoplastic brain overgrowth and yki knockdown (repo>ykidsRNA;dEGFRλ;dp110CAAX) decreased neoplastic brain overgrowth. (B) 3 µm optical projections of brain hemispheres, age-matched 3rd instar larvae. Frontal sections; anterior up; midline to left. Repo (red) labels glial cell nuclei; CD8-GFP (green) labels glial cell bodies; anti-HRP (blue) counter-stains for neurons and neuropil. (middle) repo>dEGFRλ;dp110CAAX showed increased glial cell numbers (red nuclei) compared to (upper left) wild-type. Compared to repo>dEGFRλ;dp110CAAX, (right) warts knockdown increased neoplastic glial cell numbers (red nuclei), whereas (lower left) yki knockdown reduced neoplastic glial cell numbers (red nuclei). (C, D) Low levels of Yki protein (red) was observed in wild-type central brain glia (white arrows, left panel in C) compared to high levels of cytoplasmic and nuclear Yki protein in dEGFRλ;dp110CAAX neoplastic glia (white arrows, left panel in D); Repo (blue) labels glial cell nuclei; CD8-GFP (green) labels glial cell bodies. Vigneswaran et al Supplementary Data Figure S2: YAP/TAZ expression confined to RTK-amplified tuMor cells and Maintained in patient-derived xenografts (A) On the left, immunohistochemical (IHC) staining in representative normal brain parenchyma in the cortex where YAP expression and TAZ expression was limited to vascular cells and was not detectable in normal neuronal and glial cells. -

SPINAL MUSCULAR ATROPHY: PATHOLOGY, DIAGNOSIS, CLINICAL PRESENTATION, THERAPEUTIC STRATEGIES & TREATMENTS Content
SPINAL MUSCULAR ATROPHY: PATHOLOGY, DIAGNOSIS, CLINICAL PRESENTATION, THERAPEUTIC STRATEGIES & TREATMENTS Content 1. DISCLAIMER 2. INTRODUCTION 3. SPINAL MUSCULAR ATROPHY: PATHOLOGY, DIAGNOSIS, CLINICAL PRESENTATION, THERAPEUTIC STRATEGIES & TREATMENTS 4. BIBLIOGRAPHY 5. GLOSSARY OF MEDICAL TERMS 1 SPINAL MUSCULAR ATROPHY: PATHOLOGY, DIAGNOSIS, CLINICAL PRESENTATION, THERAPEUTIC STRATEGIES & TREATMENTS Disclaimer The information in this document is provided for information purposes only. It does not constitute advice on any medical, legal, or regulatory matters and should not be used in place of consultation with appropriate medical, legal, or regulatory personnel. Receipt or use of this document does not create a relationship between the recipient or user and SMA Europe, or any other third party. The information included in this document is presented as a synopsis, may not be exhaustive and is dated November 2020. As such, it may no longer be current. Guidance from regulatory authorities, study sponsors, and institutional review boards should be obtained before taking action based on the information provided in this document. This document was prepared by SMA Europe. SMA Europe cannot guarantee that it will meet requirements or be error-free. The users and recipients of this document take on any risk when using the information contained herein. SMA Europe is an umbrella organisation, founded in 2006, which includes spinal muscular atrophy (SMA) patient and research organisations from across Europe. SMA Europe campaigns to improve the quality of life of people who live with SMA, to bring effective therapies to patients in a timely and sustainable way, and to encourage optimal patient care. SMA Europe is a non-profit umbrella organisation that consists of 23 SMA patients and research organisations from 22 countries across Europe. -

Mrna Editing, Processing and Quality Control in Caenorhabditis Elegans
| WORMBOOK mRNA Editing, Processing and Quality Control in Caenorhabditis elegans Joshua A. Arribere,*,1 Hidehito Kuroyanagi,†,1 and Heather A. Hundley‡,1 *Department of MCD Biology, UC Santa Cruz, California 95064, †Laboratory of Gene Expression, Medical Research Institute, Tokyo Medical and Dental University, Tokyo 113-8510, Japan, and ‡Medical Sciences Program, Indiana University School of Medicine-Bloomington, Indiana 47405 ABSTRACT While DNA serves as the blueprint of life, the distinct functions of each cell are determined by the dynamic expression of genes from the static genome. The amount and specific sequences of RNAs expressed in a given cell involves a number of regulated processes including RNA synthesis (transcription), processing, splicing, modification, polyadenylation, stability, translation, and degradation. As errors during mRNA production can create gene products that are deleterious to the organism, quality control mechanisms exist to survey and remove errors in mRNA expression and processing. Here, we will provide an overview of mRNA processing and quality control mechanisms that occur in Caenorhabditis elegans, with a focus on those that occur on protein-coding genes after transcription initiation. In addition, we will describe the genetic and technical approaches that have allowed studies in C. elegans to reveal important mechanistic insight into these processes. KEYWORDS Caenorhabditis elegans; splicing; RNA editing; RNA modification; polyadenylation; quality control; WormBook TABLE OF CONTENTS Abstract 531 RNA Editing and Modification 533 Adenosine-to-inosine RNA editing 533 The C. elegans A-to-I editing machinery 534 RNA editing in space and time 535 ADARs regulate the levels and fates of endogenous dsRNA 537 Are other modifications present in C. -

The Human Gene Connectome As a Map of Short Cuts for Morbid Allele Discovery
The human gene connectome as a map of short cuts for morbid allele discovery Yuval Itana,1, Shen-Ying Zhanga,b, Guillaume Vogta,b, Avinash Abhyankara, Melina Hermana, Patrick Nitschkec, Dror Friedd, Lluis Quintana-Murcie, Laurent Abela,b, and Jean-Laurent Casanovaa,b,f aSt. Giles Laboratory of Human Genetics of Infectious Diseases, Rockefeller Branch, The Rockefeller University, New York, NY 10065; bLaboratory of Human Genetics of Infectious Diseases, Necker Branch, Paris Descartes University, Institut National de la Santé et de la Recherche Médicale U980, Necker Medical School, 75015 Paris, France; cPlateforme Bioinformatique, Université Paris Descartes, 75116 Paris, France; dDepartment of Computer Science, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel; eUnit of Human Evolutionary Genetics, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3012, Institut Pasteur, F-75015 Paris, France; and fPediatric Immunology-Hematology Unit, Necker Hospital for Sick Children, 75015 Paris, France Edited* by Bruce Beutler, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 15, 2013 (received for review October 19, 2012) High-throughput genomic data reveal thousands of gene variants to detect a single mutated gene, with the other polymorphic genes per patient, and it is often difficult to determine which of these being of less interest. This goes some way to explaining why, variants underlies disease in a given individual. However, at the despite the abundance of NGS data, the discovery of disease- population level, there may be some degree of phenotypic homo- causing alleles from such data remains somewhat limited. geneity, with alterations of specific physiological pathways under- We developed the human gene connectome (HGC) to over- come this problem. -

Quantitative Interactomics in Primary T Cells Unveils TCR Signal
Quantitative interactomics in primary T cells unveils TCR signal diversification extent and dynamics Guillaume Voisinne, Kristof Kersse, Karima Chaoui, Liaoxun Lu, Julie Chaix, Lichen Zhang, Marisa Goncalves Menoita, Laura Girard, Youcef Ounoughene, Hui Wang, et al. To cite this version: Guillaume Voisinne, Kristof Kersse, Karima Chaoui, Liaoxun Lu, Julie Chaix, et al.. Quantitative interactomics in primary T cells unveils TCR signal diversification extent and dynamics. Nature Immunology, Nature Publishing Group, 2019, 20 (11), pp.1530 - 1541. 10.1038/s41590-019-0489-8. hal-03013469 HAL Id: hal-03013469 https://hal.archives-ouvertes.fr/hal-03013469 Submitted on 23 Nov 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. RESOURCE https://doi.org/10.1038/s41590-019-0489-8 Quantitative interactomics in primary T cells unveils TCR signal diversification extent and dynamics Guillaume Voisinne 1, Kristof Kersse1, Karima Chaoui2, Liaoxun Lu3,4, Julie Chaix1, Lichen Zhang3, Marisa Goncalves Menoita1, Laura Girard1,5, Youcef Ounoughene1, Hui Wang3, Odile Burlet-Schiltz2, Hervé Luche5,6, Frédéric Fiore5, Marie Malissen1,5,6, Anne Gonzalez de Peredo 2, Yinming Liang 3,6*, Romain Roncagalli 1* and Bernard Malissen 1,5,6* The activation of T cells by the T cell antigen receptor (TCR) results in the formation of signaling protein complexes (signalo somes), the composition of which has not been analyzed at a systems level.