UNIVERSITY of CALIFORNIA, IRVINE Molecular and Metabolic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Assembly Factors for the Membrane Arm of Human Complex I
Assembly factors for the membrane arm of human complex I Byron Andrews, Joe Carroll, Shujing Ding, Ian M. Fearnley, and John E. Walker1 Medical Research Council Mitochondrial Biology Unit, Cambridge CB2 0XY, United Kingdom Contributed by John E. Walker, October 14, 2013 (sent for review September 12, 2013) Mitochondrial respiratory complex I is a product of both the nuclear subunits in a fungal enzyme from Yarrowia lipolytica seem to be and mitochondrial genomes. The integration of seven subunits distributed similarly (12, 13). encoded in mitochondrial DNA into the inner membrane, their asso- The assembly of mitochondrial complex I involves building the ciation with 14 nuclear-encoded membrane subunits, the construc- 44 subunits emanating from two genomes into the two domains of tion of the extrinsic arm from 23 additional nuclear-encoded the complex. The enzyme is put together from preassembled sub- proteins, iron–sulfur clusters, and flavin mononucleotide cofactor complexes, and their subunit compositions have been characterized require the participation of assembly factors. Some are intrinsic to partially (14, 15). Extrinsic assembly factors of unknown function the complex, whereas others participate transiently. The suppres- become associated with subcomplexes that accumulate when as- sion of the expression of the NDUFA11 subunit of complex I dis- sembly and the activity of complex I are impaired by pathogenic rupted the assembly of the complex, and subcomplexes with mutations. Some assembly factor mutations also impair its activ- masses of 550 and 815 kDa accumulated. Eight of the known ex- ity (16). Other pathogenic mutations are found in all of the core trinsic assembly factors plus a hydrophobic protein, C3orf1, were subunits, and in 10 supernumerary subunits (NDUFA1, NDUFA2, associated with the subcomplexes. -

Biogenesis of NDUFS3-Less Complex I Indicates TMEM126A/OPA7 As an Assembly Factor of the ND4-Module
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.22.350587; this version posted October 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Biogenesis of NDUFS3-less complex I indicates TMEM126A/OPA7 as an assembly factor of the ND4-module Running title: NDUFS3-dependent CI disassembly pathway Luigi D’Angelo,1* Elisa Astro,1* Monica De Luise,2 Ivana Kurelac,2 Nikkitha Umesh-Ganesh,2 Shujing Ding,3 Ian M. Fearnley,3 Massimo Zeviani,3,4 Giuseppe Gasparre,2,5 Anna Maria Porcelli,1,6# Erika Fernandez-Vizarra,3,7# and Luisa Iommarini1# 1Department of Pharmacy and Biotechnology (FABIT), University of Bologna, Bologna, Italy 2Department of Medical and Surgical Sciences (DIMEC), University of Bologna, Bologna, Italy 3Medical Research Council-Mitochondrial Biology Unit, University of Cambridge, Cambridge, UK 4Venetian Institute of Molecular Medicine, Via Orus 2, 35128 Padova, Italy; Department of Neurosciences, University of Padova, via Giustiniani 2, 35128 Padova, Italy 5Center for Applied Biomedical Research (CRBA), University of Bologna, Bologna, Italy 6Interdepartmental Center of Industrial Research (CIRI) Life Science and Health Technologies, University of Bologna, Ozzano dell'Emilia, Italy 7Institute of Molecular, Cell and Systems Biology, University of Glasgow, Glasgow, UK. *Co-first authors #Co-last authors To whom correspondence should be addressed: Luisa Iommarini Department of Pharmacy and Biotechnology (FABIT) University of Bologna Via Francesco Selmi 3, 40126 Bologna, Italy Tel. +39 051 2091282 e-mail [email protected] Erika Fernandez-Vizarra Institute of Molecular, Cell and Systems Biology University of Glasgow University Avenue Glasgow G12 8QQ, UK Tel. -

Electron Transport Chain Activity Is a Predictor and Target for Venetoclax Sensitivity in Multiple Myeloma
ARTICLE https://doi.org/10.1038/s41467-020-15051-z OPEN Electron transport chain activity is a predictor and target for venetoclax sensitivity in multiple myeloma Richa Bajpai1,7, Aditi Sharma 1,7, Abhinav Achreja2,3, Claudia L. Edgar1, Changyong Wei1, Arusha A. Siddiqa1, Vikas A. Gupta1, Shannon M. Matulis1, Samuel K. McBrayer 4, Anjali Mittal3,5, Manali Rupji 6, Benjamin G. Barwick 1, Sagar Lonial1, Ajay K. Nooka 1, Lawrence H. Boise 1, Deepak Nagrath2,3,5 & ✉ Mala Shanmugam 1 1234567890():,; The BCL-2 antagonist venetoclax is highly effective in multiple myeloma (MM) patients exhibiting the 11;14 translocation, the mechanistic basis of which is unknown. In evaluating cellular energetics and metabolism of t(11;14) and non-t(11;14) MM, we determine that venetoclax-sensitive myeloma has reduced mitochondrial respiration. Consistent with this, low electron transport chain (ETC) Complex I and Complex II activities correlate with venetoclax sensitivity. Inhibition of Complex I, using IACS-010759, an orally bioavailable Complex I inhibitor in clinical trials, as well as succinate ubiquinone reductase (SQR) activity of Complex II, using thenoyltrifluoroacetone (TTFA) or introduction of SDHC R72C mutant, independently sensitize resistant MM to venetoclax. We demonstrate that ETC inhibition increases BCL-2 dependence and the ‘primed’ state via the ATF4-BIM/NOXA axis. Further, SQR activity correlates with venetoclax sensitivity in patient samples irrespective of t(11;14) status. Use of SQR activity in a functional-biomarker informed manner may better select for MM patients responsive to venetoclax therapy. 1 Department of Hematology and Medical Oncology, Winship Cancer Institute, School of Medicine, Emory University, Atlanta, GA, USA. -

SARS-Cov-2 Nucleocapsid Protein Attenuates Stress Granule Formation and Alters Gene Expression Via Direct Interaction with Host Mrnas
SARS-CoV-2 Nucleocapsid protein attenuates stress granule formation and alters gene expression via direct interaction with host mRNAs Syed Nabeel-Shah1,2,5, Hyunmin Lee1,3,5, Nujhat Ahmed1,2,5, Edyta Marcon1,5, Shaghayegh Farhangmehr1,2, Shuye Pu1, Giovanni L. Burke1,2, Kanwal Ashraf1, Hong Wei4, Guoqing Zhong1, Hua Tang1, Jianyi Yang4, Benjamin J. Blencowe1,2, Zhaolei Zhang1,2,3, Jack F. Greenblatt1,2, * 1. Donnelly Centre, University of Toronto, Toronto, M5S 3E1, Canada 2. Department of Molecular Genetics, University of Toronto, Toronto, M5S 1A8, Canada. 3. Department of Computer Sciences, University of Toronto, Toronto, M5S 1A8, Canada. 4. School of Mathematical Sciences, Nankai University, Tianjin 300071, China 5 These authors contributed equally to this work * Corresponding author: Jack F. Greenblatt; Email: [email protected] Keywords: SARS-CoV-2, COVID-19, Nucleocapsid N protein, Stress granules, G3BP1 and G3BP2, Gene expression regulation, host mRNA-binding Supplemental Figures Extended Figure 1: Dot plot overview of the interaction partners identified with 21 SARS-CoV- 2 proteins expressed in HEK293 cells. Inner circle color represents the average spectral count, the circle size maps to the relative prey abundance across all samples shown, and the circle outer edge represents the SAINT FDR. Legend is provided. Extended Figure 2: KEGG pathway (A) and Biological processes (B) enrichment analysis using cellular proteins identified as high confidence (FDR<1%) interaction partners for the SARS- CoV-2 proteins. Only top 10 most significant terms are shown (Q0.05). Darker nodes are more significantly enriched gene sets. Bigger nodes represent larger gene sets. Thicker edges represent more overlapped genes. -

Mitochondrial Disease Associated with Complex I (NADH-Coq Oxidoreductase) Deficiency
Mitochondrial disease associated with complex I (NADH-CoQ oxidoreductase) deficiency Immo E. Scheffler Division of Biology (Molecular Biology Section), University of California San Diego, 9500 Gilman Dr., La Jolla, CA 92093-0322, USA e-mail: [email protected] Abstract: Mitochondrial diseases due to a reduced capacity for oxidative phosphorylation were first identified more than 20 years ago, and their incidence is now recognized to be quite significant. In a large proportion of cases the problem can be traced to a complex I (NADH-CoQ oxidoreductase) deficiency (Phenotype MIM #252010). Because the complex consists of 44 subunits, there are many potential targets for pathogenic mutations, both on the nuclear and mitochondrial genomes. Surprisingly, however, almost half of the complex I deficiencies are due to defects in as yet unidentified genes that encode proteins other than the structural proteins of the complex. This review attempts to summarize what we know about the molecular basis of complex I deficiencies: mutations in the known structural genes, and mutations in an increasing num- ber of genes encoding “assembly factors”, that is, proteins required for the biogenesis of a functional complex I that are not found in the final complex I. More such genes must be identified before definitive genetic counselling can be applied in all cases of affected families. Introduction: Until about 25 years ago the words ‘mitochondria’ and ‘mitochondrial diseases’ probably did not exist in the vocabulary of the majority of physicians. In 1988 several pioneering papers opened a new era of biomedical research resulting in an avalanche of publications with no end in sight (Holt et al 1988a, b; Wallace et al 1988a, b). -

The Host Cellular Protein Ndufaf4 Interacts with the Vesicular Stomatitis Virus M Protein and Affects Viral Propagation
The host cellular protein Ndufaf4 interacts with the vesicular stomatitis virus M protein and affects viral propagation Wei Pan Shandong Normal University Hongmei Wang Shandong Normal University Hongbin He ( [email protected] ) Shandong Normal University https://orcid.org/0000-0002-7438-0638 Research article Keywords: VSV, Matrix Protein, Ndufaf4, Interaction Posted Date: March 25th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-18819/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/13 Abstract Background: Vesicular stomatitis virus (VSV) is an archetypal member of Mononegavirales which causes important diseases in cattle, horses and pigs. The matrix protein (M) of VSV plays critical roles in the replication, assembly/budding and pathogenesis of VSV. To further investigate the role of M during viral growth, we used a two-hybrid system to screen for host factors that interact with the M protein. Results: Here, NADH: ubiquinone oxidoreductase complex assembly factor 4 (Ndufaf4) was identied as an M-binding partner, and this interaction was conrmed by yeast cotransformation and GST pulldown assays. The globular domain of M was mapped and shown to be critical for the M-Ndufaf4 interaction. Two double mutations (E156A/H157A, D180A/E181A) in M impaired the M-Ndufaf4 interaction. Overexpression of Ndufaf4 inhibited VSV propagation, and knockdown of Ndufaf4 by short hairpin RNA (shRNA) markedly promoted VSV replication. Finally, we also demonstrate that the anti-VSV effect of Ndufaf4 is independent of activation of the type I IFN response. These results indicated that Ndufaf4 might exploit other mechanisms to affect VSV replication. -

Mitochondrial Structure and Bioenergetics in Normal and Disease Conditions
International Journal of Molecular Sciences Review Mitochondrial Structure and Bioenergetics in Normal and Disease Conditions Margherita Protasoni 1 and Massimo Zeviani 1,2,* 1 Mitochondrial Biology Unit, The MRC and University of Cambridge, Cambridge CB2 0XY, UK; [email protected] 2 Department of Neurosciences, University of Padova, 35128 Padova, Italy * Correspondence: [email protected] Abstract: Mitochondria are ubiquitous intracellular organelles found in almost all eukaryotes and involved in various aspects of cellular life, with a primary role in energy production. The interest in this organelle has grown stronger with the discovery of their link to various pathologies, including cancer, aging and neurodegenerative diseases. Indeed, dysfunctional mitochondria cannot provide the required energy to tissues with a high-energy demand, such as heart, brain and muscles, leading to a large spectrum of clinical phenotypes. Mitochondrial defects are at the origin of a group of clinically heterogeneous pathologies, called mitochondrial diseases, with an incidence of 1 in 5000 live births. Primary mitochondrial diseases are associated with genetic mutations both in nuclear and mitochondrial DNA (mtDNA), affecting genes involved in every aspect of the organelle function. As a consequence, it is difficult to find a common cause for mitochondrial diseases and, subsequently, to offer a precise clinical definition of the pathology. Moreover, the complexity of this condition makes it challenging to identify possible therapies or drug targets. Keywords: ATP production; biogenesis of the respiratory chain; mitochondrial disease; mi-tochondrial electrochemical gradient; mitochondrial potential; mitochondrial proton pumping; mitochondrial respiratory chain; oxidative phosphorylation; respiratory complex; respiratory supercomplex Citation: Protasoni, M.; Zeviani, M. -
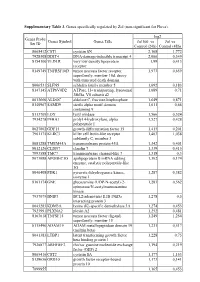
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

Accessory Subunits Are Integral for Assembly and Function of Human Mitochondrial Complex I
Accessory subunits are integral for assembly and function of human mitochondrial complex I David A. Stroud1*, Elliot E. Surgenor1, Luke E. Formosa1,2, Boris Reljic2†, Ann E. Frazier3,4, Marris G. Dibley1, Laura D. Osellame1, Tegan Stait3, Traude H. Beilharz1, David R. Thorburn3-5, Agus Salim6, Michael T. Ryan1* 1Department of Biochemistry and Molecular Biology, Monash Biomedicine Discovery Institute, Monash University, 3800, Melbourne, Australia. 2Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University 3086, Melbourne, Australia. 3Murdoch Childrens Research Institute, Royal Children’s Hospital, Melbourne 3052, Australia 4Department of Pediatrics, University of Melbourne, Melbourne 3052, Australia. 5Victorian Clinical Genetics Services, Royal Children’s Hospital 3052, Melbourne, Australia. 6Department of Mathematics and Statistics, La Trobe University 3086, Melbourne Australia. *Correspondence to: [email protected] and [email protected] †Current address: Walter and Eliza Hall Institute of Medical Research, Parkville, Melbourne, Victoria 3052, Australia Complex I (NADH:ubiquinone oxidoreductase) is the first enzyme of the mitochondrial respiratory chain (RC) and is composed of 44 different subunits in humans, making it one of the largest known multi-subunit membrane protein complexes1. Complex I exists in supercomplex forms with RC complexes III and IV, which are together required for the generation of a transmembrane proton gradient used for the synthesis of ATP2. Complex I is also a major source of damaging reactive oxygen species and its dysfunction is associated with mitochondrial disease, Parkinson’s disease and aging3-5. Bacterial and human complex I share 14 core subunits essential for enzymatic function, however the role and requirement of the remaining 31 human accessory subunits is unclear1,6. -

A High-Density Human Mitochondrial Proximity Interaction Network
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.01.020479; this version posted April 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A high-density human mitochondrial proximity interaction network Hana Antonicka1,2, Zhen-Yuan Lin3, Alexandre Janer1,2, Woranontee Weraarpachai1,5, Anne- Claude Gingras3,4,*, Eric A. Shoubridge1,2* 1 Montreal Neurological Institute, McGill University, Montreal, QC, Canada 2 Department of Human Genetics, McGill University, Montreal, QC, Canada 3 Lunenfeld-Tanenbaum Research Institute, Toronto, ON, Canada 4 Department of Molecular Genetics, University of Toronto, Toronto, ON, Canada 5 Department of Biochemistry, Faculty of Medicine, Chiang Mai University, Chiang Mai, Thailand Data deposition: Mass spectrometry data have been deposited in the Mass spectrometry Interactive Virtual Environment (MassIVE, http://massive.ucsd.edu). * corresponding authors Correspondence: e-mail: [email protected] Phone: (514) 398-1997 Fax: (514) 398-1509 e-mail: [email protected] Phone: (416) 586-5027 Fax: (416) 586-8869 Summary We used BioID, a proximity-dependent biotinylation assay, to interrogate 100 mitochondrial baits from all mitochondrial sub-compartments to create a high resolution human mitochondrial proximity interaction network. We identified 1465 proteins, producing 15626 unique high confidence proximity interactions. Of these, 528 proteins were previously annotated as mitochondrial, nearly half of the mitochondrial proteome defined by Mitocarta 2.0. Bait-bait analysis showed a clear separation of mitochondrial compartments, and correlation analysis among preys across all baits allowed us to identify functional clusters involved in diverse mitochondrial functions, and to assign uncharacterized proteins to specific modules. -

Assembly Factors for the Membrane Arm of Human Complex I
Assembly factors for the membrane arm of human complex I Byron Andrews, Joe Carroll, Shujing Ding, Ian M. Fearnley, and John E. Walker1 Medical Research Council Mitochondrial Biology Unit, Cambridge CB2 0XY, United Kingdom Contributed by John E. Walker, October 14, 2013 (sent for review September 12, 2013) Mitochondrial respiratory complex I is a product of both the nuclear subunits in a fungal enzyme from Yarrowia lipolytica seem to be and mitochondrial genomes. The integration of seven subunits distributed similarly (12, 13). encoded in mitochondrial DNA into the inner membrane, their asso- The assembly of mitochondrial complex I involves building the ciation with 14 nuclear-encoded membrane subunits, the construc- 44 subunits emanating from two genomes into the two domains of tion of the extrinsic arm from 23 additional nuclear-encoded the complex. The enzyme is put together from preassembled sub- proteins, iron–sulfur clusters, and flavin mononucleotide cofactor complexes, and their subunit compositions have been characterized require the participation of assembly factors. Some are intrinsic to partially (14, 15). Extrinsic assembly factors of unknown function the complex, whereas others participate transiently. The suppres- become associated with subcomplexes that accumulate when as- sion of the expression of the NDUFA11 subunit of complex I dis- sembly and the activity of complex I are impaired by pathogenic rupted the assembly of the complex, and subcomplexes with mutations. Some assembly factor mutations also impair its activ- masses of 550 and 815 kDa accumulated. Eight of the known ex- ity (16). Other pathogenic mutations are found in all of the core trinsic assembly factors plus a hydrophobic protein, C3orf1, were subunits, and in 10 supernumerary subunits (NDUFA1, NDUFA2, associated with the subcomplexes. -

Mitochondrial DNA (Mtdna) Test Requisition
SHIP TO: Medical Genetics Laboratories BCM-MEDICAL GENETICS LABORATORIES Baylor College of Medicine PHONE: 800-411-GENE | FAX: 713-798-2787 | www.bcmgeneticlabs.org 2450 Holcombe, Grand Blvd. -Receiving Dock Houston, TX 77021-2024 MITOCHONDRIAL DNA (mtDNA) TEST REQUISITION Phone: 713-798-6555 PATIENT INFORMATION INDICATION FOR STUDY NAME: DATE OF COLLECTION: / / LAST NAME FIRST NAME MI MM DD YY HOSPITAL#: ACCESSION#: DATE OF BIRTH: / / GENDER (Please select one): FEMALE MALE MM DD YY SAMPLE TYPE (Please select one): ETHNIC BACKGROUND (Select all that apply): UNKNOWN BLOOD AFRICAN AMERICAN SKELETAL MUSCLE ASIAN DNA (Specify Source): ASHKENAZIC JEWISH EUROPEAN CAUCASIAN -OR- HISPANIC OTHER (Specify): NATIVE AMERICAN INDIAN PLACE PATIENT STICKER HERE OTHER JEWISH OTHER (Please specify): REPORTING INFORMATION ADDITIONAL PROFESSIONAL REPORT RECIPIENTS PHYSICIAN: NAME: INSTITUTION: PHONE: *FAX: PHONE: *FAX: NAME: EMAIL (INTERNATIONAL CLIENT REQUIREMENT): PHONE: *FAX: *BCM-MEDICAL GENETIC LABORATORIES HAS A FAX ONLY POLICY FOR REPORTING INDICATION FOR STUDY SYMPTOMATIC (Summarize below.): *FAMILIAL MUTATION/VARIANT ANALYSIS: Complete all fields below and attach the proband's report. GENE NAME: ASYMPTOMATIC/POSITIVE FAMILY HISTORY: (ATTACH FAMILY HISTORY) MUTATION/UNCLASSIFIED VARIANT: RELATIONSHIP TO PROBAND: THIS INDIVIDUAL IS CURRENTLY: SYMPTOMATIC ASYMPTOMATIC *If family mutation is known, complete the FAMILIAL MUTATION/ VARIANT ANALYSIS section. NAME OF PROBAND: ASYMPTOMATIC/POPULATION SCREENING RELATIONSHIP TO PROBAND: OTHER (Specify clinical findings below.): BCM LAB#: A COPY OF ORIGINAL RESULTS ATTACHED IF PROBAND TESTING WAS PERFORMED AT ANOTHER LAB, CALL TO DISCUSS PRIOR TO SENDING SAMPLE. A POSITIVE CONTROL MAY BE REQUIRED IN SOME CASES. REQUIRED: NEW YORK STATE PHYSICIAN SIGNATURE OF CONSENT I certify that the patient specified above and/or their legal guardian has been informed of the benefits, risks, and limitations of the laboratory test(s) requested.