Association Mapping Based on a Common-Garden Migration Experiment Reveals
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Tepzz¥ 6Z54za T
(19) TZZ¥ ZZ_T (11) EP 3 260 540 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 27.12.2017 Bulletin 2017/52 C12N 15/113 (2010.01) A61K 9/127 (2006.01) A61K 31/713 (2006.01) C12Q 1/68 (2006.01) (21) Application number: 17000579.7 (22) Date of filing: 12.11.2011 (84) Designated Contracting States: • Sarma, Kavitha AL AT BE BG CH CY CZ DE DK EE ES FI FR GB Philadelphia, PA 19146 (US) GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO • Borowsky, Mark PL PT RO RS SE SI SK SM TR Needham, MA 02494 (US) • Ohsumi, Toshiro Kendrick (30) Priority: 12.11.2010 US 412862 P Cambridge, MA 02141 (US) 20.12.2010 US 201061425174 P 28.07.2011 US 201161512754 P (74) Representative: Clegg, Richard Ian et al Mewburn Ellis LLP (62) Document number(s) of the earlier application(s) in City Tower accordance with Art. 76 EPC: 40 Basinghall Street 11840099.3 / 2 638 163 London EC2V 5DE (GB) (71) Applicant: The General Hospital Corporation Remarks: Boston, MA 02114 (US) •Thecomplete document including Reference Tables and the Sequence Listing can be downloaded from (72) Inventors: the EPO website • Lee, Jeannie T •This application was filed on 05-04-2017 as a Boston, MA 02114 (US) divisional application to the application mentioned • Zhao, Jing under INID code 62. San Diego, CA 92122 (US) •Claims filed after the date of receipt of the divisional application (Rule 68(4) EPC). (54) POLYCOMB-ASSOCIATED NON-CODING RNAS (57) This invention relates to long non-coding RNAs (IncRNAs), libraries of those ncRNAs that bind chromatin modifiers, such as Polycomb Repressive Complex 2, inhibitory nucleic acids and methods and compositions for targeting IncRNAs. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Advances in Prognostic Methylation Biomarkers for Prostate Cancer
cancers Review Advances in Prognostic Methylation Biomarkers for Prostate Cancer 1 1,2 1,2, 1,2, , Dilys Lam , Susan Clark , Clare Stirzaker y and Ruth Pidsley * y 1 Epigenetics Research Laboratory, Genomics and Epigenetics Division, Garvan Institute of Medical Research, Sydney, New South Wales 2010, Australia; [email protected] (D.L.); [email protected] (S.C.); [email protected] (C.S.) 2 St. Vincent’s Clinical School, University of New South Wales, Sydney, New South Wales 2010, Australia * Correspondence: [email protected]; Tel.: +61-2-92958315 These authors have contributed equally. y Received: 22 September 2020; Accepted: 13 October 2020; Published: 15 October 2020 Simple Summary: Prostate cancer is a major cause of cancer-related death in men worldwide. There is an urgent clinical need for improved prognostic biomarkers to better predict the likely outcome and course of the disease and thus inform the clinical management of these patients. Currently, clinically recognised prognostic markers lack sensitivity and specificity in distinguishing aggressive from indolent disease, particularly in patients with localised, intermediate grade prostate cancer. Thus, there is major interest in identifying new molecular biomarkers to complement existing standard clinicopathological markers. DNA methylation is a frequent alteration in the cancer genome and offers potential as a reliable and robust biomarker. In this review, we provide a comprehensive overview of the current state of DNA methylation biomarker studies in prostate cancer prognosis. We highlight advances in this field that have enabled the discovery of novel prognostic genes and discuss the potential of methylation biomarkers for noninvasive liquid-biopsy testing. -
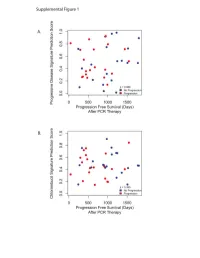
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Abstracts from the 51St European Society of Human Genetics Conference: Electronic Posters
European Journal of Human Genetics (2019) 27:870–1041 https://doi.org/10.1038/s41431-019-0408-3 MEETING ABSTRACTS Abstracts from the 51st European Society of Human Genetics Conference: Electronic Posters © European Society of Human Genetics 2019 June 16–19, 2018, Fiera Milano Congressi, Milan Italy Sponsorship: Publication of this supplement was sponsored by the European Society of Human Genetics. All content was reviewed and approved by the ESHG Scientific Programme Committee, which held full responsibility for the abstract selections. Disclosure Information: In order to help readers form their own judgments of potential bias in published abstracts, authors are asked to declare any competing financial interests. Contributions of up to EUR 10 000.- (Ten thousand Euros, or equivalent value in kind) per year per company are considered "Modest". Contributions above EUR 10 000.- per year are considered "Significant". 1234567890();,: 1234567890();,: E-P01 Reproductive Genetics/Prenatal Genetics then compared this data to de novo cases where research based PO studies were completed (N=57) in NY. E-P01.01 Results: MFSIQ (66.4) for familial deletions was Parent of origin in familial 22q11.2 deletions impacts full statistically lower (p = .01) than for de novo deletions scale intelligence quotient scores (N=399, MFSIQ=76.2). MFSIQ for children with mater- nally inherited deletions (63.7) was statistically lower D. E. McGinn1,2, M. Unolt3,4, T. B. Crowley1, B. S. Emanuel1,5, (p = .03) than for paternally inherited deletions (72.0). As E. H. Zackai1,5, E. Moss1, B. Morrow6, B. Nowakowska7,J. compared with the NY cohort where the MFSIQ for Vermeesch8, A. -

Identification of DNA Methylation Associated Gene Signatures in Endometrial Cancer Via Integrated Analysis of DNA Methylation and Gene Expression Systematically
J Gynecol Oncol. 2017 Nov;28(6):e83 https://doi.org/10.3802/jgo.2017.28.e83 pISSN 2005-0380·eISSN 2005-0399 Original Article Identification of DNA methylation associated gene signatures in endometrial cancer via integrated analysis of DNA methylation and gene expression systematically Chuandi Men ,1,2 Hongjuan Chai ,1 Xumin Song ,1 Yue Li ,1 Huawen Du ,1 Qing Ren 1 1Department of Gynecology and Obstetrics, Shanghai Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China Received: May 11, 2017 2Graduate School, Bengbu Medical College, Bengbu, China Revised: Aug 2, 2017 Accepted: Aug 10, 2017 Correspondence to ABSTRACT Qing Ren Department of Gynecology and Obstetrics, Objective: Endometrial cancer (EC) is a common gynecologic cancer worldwide. However, Shanghai Ninth People's Hospital, Shanghai the pathogenesis of EC has not been epigenetically elucidated. Here, this study aims Jiao Tong University School of Medicine, No. to describe the DNA methylation profile and identify favorable gene signatures highly 280, Mohe Road, Baoshan District, Shanghai 201900, China. associated with aberrant DNA methylation changes in EC. E-mail: [email protected] Methods: The data regarding DNA methylation and gene expression were downloaded from The Cancer Genome Atlas (TCGA) database. Differentially methylated CpG sites (DMCs), Copyright © 2017. Asian Society of differentially methylated regions (DMRs), and differentially expressed genes (DEGs) were Gynecologic Oncology, Korean Society of Gynecologic Oncology identified, and the relationship between the 2 omics was further analyzed. In addition, This is an Open Access article distributed weighted CpG site co-methylation network (WCCN) was constructed followed by an under the terms of the Creative Commons integrated analysis of DNA methylation and gene expression data. -

Genetics of Distal Hereditary Motor Neuropathies
GENETICSOFDISTALHEREDITARY MOTOR NEUROPATHIES By alexander peter drew A thesis submitted for the Degree of Doctor of Philosophy Supervised by Professor Garth A. Nicholson Dr. Ian P. Blair Faculty of Medicine University of Sydney 2012 statement No part of the work described in this thesis has been submitted in fulfilment of the requirements for any other academic degree or qualification. Except where due acknowledgement has been made, all experimental work was performed by the author. Alexander Peter Drew CONTENTS acknowledgements ............................. i summary .................................... ii list of figures ................................ v list of tables ................................ viii acronyms and abbreviations ..................... xi publications ................................. xiv 1 literature review ........................... 1 1.1 Molecular genetics and mechanisms of disease in Distal Hereditary Motor Neuropathies . .1 1.1.1 Small heat shock protein family . .2 1.1.2 Dynactin 1 (DCTN1).....................9 1.1.3 Immunoglobulin mu binding protein 2 gene (IGHMBP2) 11 1.1.4 Senataxin (SETX)....................... 14 1.1.5 Glycyl-tRNA synthase (GARS)............... 16 1.1.6 Berardinelli-Seip congenital lipodystrophy 2 (SEIPIN) gene (BSCL2)......................... 18 1.1.7 ATPase, Cu2+-transporting, alpha polypeptide gene (ATP7A) 20 1.1.8 Pleckstrin homology domain-containing protein, G5 gene (PLEKHG5)........................... 21 1.1.9 Transient receptor potential cation channel, V4 gene (TRPV4) 22 1.1.10 DYNC1H1 ........................... 23 1.1.11 Clinical variability in dHMN . 24 1.1.12 Common disease mechanisms in dHMN . 29 2 general materials and methods ................. 32 2.1 General materials and reagents . 32 2.1.1 Reagents and Enzymes . 32 2.1.2 Equipment . 33 2.1.3 Plasticware . 33 2.2 Study participants . 34 2.3 DNA methods . -

Ep 3039174 B1
(19) TZZ¥Z¥__T (11) EP 3 039 174 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C12Q 1/6886 (2018.01) C12Q 1/6883 (2018.01) 16.10.2019 Bulletin 2019/42 (86) International application number: (21) Application number: 14840036.9 PCT/US2014/053306 (22) Date of filing: 28.08.2014 (87) International publication number: WO 2015/031694 (05.03.2015 Gazette 2015/09) (54) OLIGONUCLEOTIDE PROBES AND USES THEREOF OLIGONUKLEOTIDSONDEN UND VERWENDUNGEN DAVON SONDES OLIGONUCLÉOTIDIQUES ET LEURS UTILISATIONS (84) Designated Contracting States: (74) Representative: Patent Boutique LLP AL AT BE BG CH CY CZ DE DK EE ES FI FR GB 10A Printing House Yard GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO Hackney Road PL PT RO RS SE SI SK SM TR London E2 7PR (GB) (30) Priority: 28.08.2013 US 201361871107 P (56) References cited: 06.09.2013 US 201361874621 P WO-A1-2011/066589 WO-A2-2010/056337 06.11.2013 US 201361900975 P WO-A2-2011/088226 WO-A2-2013/022995 05.12.2013 US 201361912471 P 06.01.2014 US 201461924192 P • TERESA JANAS ET AL: "The selection of 06.03.2014 US 201461949216 P aptamers specific for membrane molecular 03.04.2014 US 201461974949 P targets", CELLULAR & MOLECULAR BIOLOGY 07.05.2014 US 201461990085 P LETTERS, vol. 16, no. 1, 1 March 2011 16.05.2014 US 201461994704 P (2011-03-01), pages 25-39, XP055001923, ISSN: 14.07.2014 US 201462024436 P 1425-8153, DOI: 10.2478/s11658-010-0023-3 • HENNING ULRICH ET AL: "Disease-specific (43) Date of publication of application: biomarkerdiscovery by aptamers", CYTOMETRY 06.07.2016 Bulletin 2016/27 PART A, vol. -

(B6;129.Cg-Gt(ROSA)26Sor Tm20(CAG-Ctgf-GFP)Jsd) Were Crossed with Female Foxd1cre/+ Heterozygote Mice 1, and Experimental Mice Were Selected As Foxd1cre/+; Rs26cig/+
Supplemental Information SI Methods Animal studies Heterozygote mice (B6;129.Cg-Gt(ROSA)26Sor tm20(CAG-Ctgf-GFP)Jsd) were crossed with female Foxd1Cre/+ heterozygote mice 1, and experimental mice were selected as Foxd1Cre/+; Rs26CIG/+. In some studies Coll-GFPTg or TCF/Lef:H2B-GFPTg mice or Foxd1Cre/+; Rs26tdTomatoR/+ mice were used as described 2; 3. Left kidneys were subjected to ureteral obstruction using a posterior surgical approach as described 2. In some experiments recombinant mouse DKK1 (0.5mg/kg) or an equal volume of vehicle was administered by daily IP injection. In the in vivo ASO experiment, either specific Lrp6 (TACCTCAATGCGATTT) or scrambled negative control ASO (AACACGTCTATACGC) (30mg/kg) (Exiqon, LNA gapmers) was administered by IP injection on d-1, d1, d4, and d7. In other experiments anti-CTGF domain-IV antibodies (5mg/kg) or control IgG were administered d-1, d1 and d6. All animal experiments were performed under approved IACUC protocols held at the University of Washington and Biogen. Recombinant protein and antibody generation and characterization Human CTGF domain I (sequence Met1 CPDEPAPRCPAGVSLVLDGCGCCRVCAKQLGELCTERDPCDPHKGLFC), domain I+II (sequence Met1CPDEPAPRCPAGVSLVLDGCGCCRVCAKQLGELCTERDPCDPHKGLFCCIFGGT VYRSGESFQSSCKYQCTCLDGAVGCMPLCSMDVRLPSPDCPFPRRVKLPGKCCEE) were cloned and expressed in 293 cells, and purified by Chelating SFF(Ni) Column, tested for single band by SEC and PAGE, and tested for absence of contamination. Domain-IV (sequence GKKCIRTPKISKPIKFELSGCTSMKTYRAKFCGVCTDGRCCTPHRTTTLPVEFKCPDGE VMKKNMMFIKTCACHYNCPGDNDIFESLYYRKMY) was purchased from Peprotech. Mouse or human DKK1 was generated from the coding sequence with some modifications and a tag. Secreted protein was harvested from 293 cells, and purified by nickel column, and tested for activity in a supertopflash (STF) assay 4. DKK1 showed EC50 of 0.69nM for WNT3a-induced WNT signaling in STF cells. -

Development of Network-Based Analysis Methods with Application to the Genetic Component of Asthma Yuanlong Liu
Development of network-based analysis methods with application to the genetic component of asthma Yuanlong Liu To cite this version: Yuanlong Liu. Development of network-based analysis methods with application to the genetic com- ponent of asthma. Human genetics. Université Sorbonne Paris Cité, 2017. English. NNT : 2017US- PCC329. tel-02466418 HAL Id: tel-02466418 https://tel.archives-ouvertes.fr/tel-02466418 Submitted on 4 Feb 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparé à l’Université Paris Diderot ÉCOLE DOCTORALE PIERRE LOUIS DE SANTÉ PUBLIQUE À PARIS ÉPIDÉMIOLOGIE ET SCIENCES DE L’INFORMATION BIOMÉDICALE (ED 393) Unité de recherche: UMR 946 - Variabilité Génétique et Maladies Humaines DOCTORAT Spécialité: Epidémiologie Génétique Yuanlong LIU Development of network-based analysis methods with application to the genetic component of asthma Thèse dirigée par Florence DEMENAIS Présentée et soutenue publiquement à Paris le 13 Novembre 2017 JURY M. Bertram MÜLLER-MYHSOK Professeur, Technische Universität München Rapporteur Mme Kristel VAN STEEN Professeur, Université de Liège Rapporteur M. Benno SCHWIKOWSKI Directeur de Recherche, Institut Pasteur Examinateur M. Mohamed NADIF Professeur, Université Paris-Descartes Examinateur M. -

Analysis of Molecular Events Involved in Chondrogenesis and Somitogenesis by Global Gene Expression Profiling
Technische Universit¨at Munc¨ hen GSF-Forschungszentrum Neuherberg Analysis of Molecular Events Involved in Chondrogenesis and Somitogenesis by Global Gene Expression Profiling Matthias Wahl Vollst¨andiger Abdruck der von der Fakult¨at Wissenschaftszentrum Weihen- stephan fur¨ Ern¨ahrung, Landnutzung und Umwelt der Technischen Univer- sit¨at Munc¨ hen zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ.-Prof. Dr. A. Gierl Prufer¨ der Dissertation: 1. Hon.-Prof. Dr. R. Balling, Technische Uni- versit¨at Carlo-Wilhelmina zu Braunschweig 2. Univ.-Prof. Dr. W. Wurst Die Dissertation wurde am 29. Oktober 2003 bei der Technischen Univer- sit¨at Munc¨ hen eingereicht und durch die Fakult¨at Wissenschaftszentrum Wei- henstephan fur¨ Ern¨ahrung, Landnutzung und Umwelt am 14. Januar 2004 angenommen. ii Zusammenfassung Um die molekularen Mechanismen, welche Knorpelentwicklung und Somito- genese steuern, aufzukl¨aren, wurde eine globale quantitative Genexpressions- analyse unter Verwendung von SAGE (Serial Analysis of Gene Expression) an der knorpelbildenden Zelllinie ATDC5 und an somitischem Gewebe, pr¨apariert von E10.5 M¨ausen, durchgefuhrt.¨ Unter insgesamt 43,656 von der murinen knorpelbildenden Zelllinie ATDC5 gewonnenen SAGE Tags (21,875 aus uninduzierten Zellen und 21,781 aus Zellen, die fur¨ 6h mit BMP4 induziert wurden) waren 139 Transkripte un- terschiedlich in den beiden Bibliotheken repr¨asentiert (P 0.05). 95 Tags ≤ konnten einzelnen UniGene Eintr¨agen zugeordnet werden -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons.