Truncated Cables1 Causes Agenesis of the Corpus Callosum in Mice
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Multidrug Transporter MRP4/ABCC4 As a Key Determinant of Pancreatic
www.nature.com/scientificreports OPEN Multidrug transporter MRP4/ ABCC4 as a key determinant of pancreatic cancer aggressiveness A. Sahores1, A. Carozzo1, M. May1, N. Gómez1, N. Di Siervi1, M. De Sousa Serro1, A. Yanef1, A. Rodríguez‑González2, M. Abba3, C. Shayo2 & C. Davio1* Recent fndings show that MRP4 is critical for pancreatic ductal adenocarcinoma (PDAC) cell proliferation. Nevertheless, the signifcance of MRP4 protein levels and function in PDAC progression is still unclear. The aim of this study was to determine the role of MRP4 in PDAC tumor aggressiveness. Bioinformatic studies revealed that PDAC samples show higher MRP4 transcript levels compared to normal adjacent pancreatic tissue and circulating tumor cells express higher levels of MRP4 than primary tumors. Also, high levels of MRP4 are typical of high-grade PDAC cell lines and associate with an epithelial-mesenchymal phenotype. Moreover, PDAC patients with high levels of MRP4 depict dysregulation of pathways associated with migration, chemotaxis and cell adhesion. Silencing MRP4 in PANC1 cells reduced tumorigenicity and tumor growth and impaired cell migration. Transcriptomic analysis revealed that MRP4 silencing alters PANC1 gene expression, mainly dysregulating pathways related to cell-to-cell interactions and focal adhesion. Contrarily, MRP4 overexpression signifcantly increased BxPC-3 growth rate, produced a switch in the expression of EMT markers, and enhanced experimental metastatic incidence. Altogether, our results indicate that MRP4 is associated with a more aggressive phenotype in PDAC, boosting pancreatic tumorigenesis and metastatic capacity, which could fnally determine a fast tumor progression in PDAC patients. Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal human malignancies, due to its late diag- nosis, inherent resistance to treatment and early dissemination 1. -

Estimation of Non-Null SNP Effect Size Distributions Enables the Detection
bioRxiv preprint doi: https://doi.org/10.1101/597484; this version posted May 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 1 Estimation of Non-null SNP Effect Size Distributions Enables 2 the Detection of Enriched Genes Underlying Complex Traits 3 1,2 1,2 2-4 4 Wei Cheng , Sohini Ramachandran y, and Lorin Crawford y 5 1 Department of Ecology and Evolutionary Biology, Brown University, Providence, RI, 6 USA 7 2 Center for Computational Molecular Biology, Brown University, Providence, RI, USA 8 3 Department of Biostatistics, Brown University, Providence, RI, USA 9 4 Center for Statistical Sciences, Brown University, Providence, RI, USA 10 Corresponding E-mail: [email protected]; lorin [email protected] y 11 Abstract 12 Traditional univariate genome-wide association studies generate false positives and negatives due to 13 difficulties distinguishing associated variants from variants with spurious nonzero effects that do not 14 directly influence the trait. Recent efforts have been directed at identifying genes or signaling pathways 15 enriched for mutations in quantitative traits or case-control studies, but these can be computationally 16 costly and hampered by strict model assumptions. Here, we present gene-", a new approach for identifying 17 statistical associations between sets of variants and quantitative traits. Our key insight is that enrichment 18 studies on the gene-level are improved when we reformulate the genome-wide SNP-level null hypothesis 19 to identify spurious small-to-intermediate SNP effects and classify them as non-causal. -

Noncoding Rnas As Novel Pancreatic Cancer Targets
NONCODING RNAS AS NOVEL PANCREATIC CANCER TARGETS by Amy Makler A Thesis Submitted to the Faculty of The Charles E. Schmidt College of Science In Partial Fulfillment of the Requirements for the Degree of Master of Science Florida Atlantic University Boca Raton, FL August 2018 Copyright 2018 by Amy Makler ii ACKNOWLEDGEMENTS I would first like to thank Dr. Narayanan for his continuous support, constant encouragement, and his gentle, but sometimes critical, guidance throughout the past two years of my master’s education. His faith in my abilities and his belief in my future success ensured I continue down this path of research. Working in Dr. Narayanan’s lab has truly been an unforgettable experience as well as a critical step in my future endeavors. I would also like to extend my gratitude to my committee members, Dr. Binninger and Dr. Jia, for their support and suggestions regarding my thesis. Their recommendations added a fresh perspective that enriched our initial hypothesis. They have been indispensable as members of my committee, and I thank them for their contributions. My parents have been integral to my successes in life and their support throughout my education has been crucial. They taught me to push through difficulties and encouraged me to pursue my interests. Thank you, mom and dad! I would like to thank my boyfriend, Joshua Disatham, for his assistance in ensuring my writing maintained a logical progression and flow as well as his unwavering support. He was my rock when the stress grew unbearable and his encouraging words kept me pushing along. -

Recalibrating the Genome-Wide Association Null Hypothesis
bioRxiv preprint doi: https://doi.org/10.1101/597484; this version posted October 4, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 1 Recalibrating the Genome-wide Association Null Hypothesis 2 Enables the Detection of Core Genes Underlying Complex 3 Traits 4 1,2 1,2 2-4 5 Wei Cheng , Sohini Ramachandran y, and Lorin Crawford y 6 1 Department of Ecology and Evolutionary Biology, Brown University, Providence, RI, 7 USA 8 2 Center for Computational Molecular Biology, Brown University, Providence, RI, USA 9 3 Department of Biostatistics, Brown University, Providence, RI, USA 10 4 Center for Statistical Sciences, Brown University, Providence, RI, USA 11 E-mail: [email protected]; lorin [email protected] y 12 Abstract 13 Traditional univariate genome-wide association studies generate false positives and negatives due to 14 difficulties distinguishing causal variants from \interactive" variants (i.e., variants correlated with causal 15 variants without directly influencing the trait). Recent efforts have been directed at identifying gene or 16 pathway associations, but these can be computationally costly and hampered by strict model assumptions. 17 Here, we present gene-", a new approach for identifying statistical associations between sets of variants 18 and quantitative traits. Our key innovation is a recalibration of the genome-wide null hypothesis to treat 19 small-yet-nonzero associations emitted by interactive variants as non-causal. gene-" efficiently identifies 20 core genes under a variety of simulated genetic architectures, achieving greater than a 90% true positive 21 rate at 1% false positive rate for polygenic traits. -

Loss-Of-Function Mutations in the CABLES1 Gene Are a Novel Cause
248 L C Hernández- CABLES1 mutations in 24:8 379–392 Research Ramírez et al. Cushing’s disease Open Access Loss-of-function mutations in the CABLES1 gene are a novel cause of Cushing’s disease Laura C Hernández-Ramírez1, Ryhem Gam2, Nuria Valdés1,3, Maya B Lodish1, Nathan Pankratz4, Aurelio Balsalobre2, Yves Gauthier2, Fabio R Faucz1, Giampaolo Trivellin1, Prashant Chittiboina5, John Lane4, Denise M Kay6 Aggeliki Dimopoulos7, Stephan Gaillard8,9, Mario Neou8, Jérôme Bertherat8,10, Guillaume Assié8,10, Chiara Villa8,11,12, James L Mills7,*, Jacques Drouin2,*, and Constantine A Stratakis1,* 1Section on Endocrinology and Genetics, Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), National Institutes of Health (NIH), Bethesda, Maryland, USA 2Laboratoire de Génétique Moléculaire, Institut de Recherches Cliniques de Montréal (IRCM), Montréal, Québec, Canada 3Service of Endocrinology and Nutrition, Hospital Universitario Central de Asturias, Instituto Universita- rio de Oncología del Principado de Asturias, Universidad de Oviedo, Oviedo, Spain 4Department of Laboratory Medicine and Pathology, University of Minnesota Medical School, Minneapolis, Minnesota, USA 5Surgical Neurology Branch, National Institute of Neurological Disorders and Stroke (NINDS), National Institutes of Health (NIH), Bethesda, Maryland, USA 6Newborn Screening Program, Wadsworth Center, New York State Department of Health, Albany, New York, USA 7Division of Intramural Population Health Research, Epidemiology Branch, Eunice Kennedy Shriver -

Molecular Characterization of Selectively Vulnerable Neurons in Alzheimer’S Disease
RESOURCE https://doi.org/10.1038/s41593-020-00764-7 Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease Kun Leng1,2,3,4,13, Emmy Li1,2,3,13, Rana Eser 5, Antonia Piergies5, Rene Sit2, Michelle Tan2, Norma Neff 2, Song Hua Li5, Roberta Diehl Rodriguez6, Claudia Kimie Suemoto7,8, Renata Elaine Paraizo Leite7, Alexander J. Ehrenberg 5, Carlos A. Pasqualucci7, William W. Seeley5,9, Salvatore Spina5, Helmut Heinsen7,10, Lea T. Grinberg 5,7,11 ✉ and Martin Kampmann 1,2,12 ✉ Alzheimer’s disease (AD) is characterized by the selective vulnerability of specific neuronal populations, the molecular sig- natures of which are largely unknown. To identify and characterize selectively vulnerable neuronal populations, we used single-nucleus RNA sequencing to profile the caudal entorhinal cortex and the superior frontal gyrus—brain regions where neurofibrillary inclusions and neuronal loss occur early and late in AD, respectively—from postmortem brains spanning the progression of AD-type tau neurofibrillary pathology. We identified RORB as a marker of selectively vulnerable excitatory neu- rons in the entorhinal cortex and subsequently validated their depletion and selective susceptibility to neurofibrillary inclusions during disease progression using quantitative neuropathological methods. We also discovered an astrocyte subpopulation, likely representing reactive astrocytes, characterized by decreased expression of genes involved in homeostatic functions. Our characterization of selectively vulnerable neurons in AD paves the way for future mechanistic studies of selective vulnerability and potential therapeutic strategies for enhancing neuronal resilience. elective vulnerability is a fundamental feature of neurodegen- Here, we performed snRNA-seq on postmortem brain tissue erative diseases, in which different neuronal populations show from a cohort of individuals spanning the progression of AD-type a gradient of susceptibility to degeneration. -

Peri-Implantation Lethality in Mice Carrying Megabase-Scale Deletion
www.nature.com/scientificreports OPEN Peri-implantation lethality in mice carrying megabase-scale deletion on 5qc3.3 is caused by Exoc1 null Received: 12 June 2015 Accepted: 03 August 2015 mutation Published: 08 September 2015 Seiya Mizuno*, Kohei Takami*, Yoko Daitoku, Yoko Tanimoto, Tra Thi Huong Dinh, Saori Mizuno-Iijima, Yoshikazu Hasegawa, Satoru Takahashi, Fumihiro Sugiyama & Ken-ichi Yagami We found a novel spontaneous mouse mutant with depigmentation in the ventral body, which we called White Spotting (WS) mouse. Genetic investigation revealed deletion of a > 1.2-Mb genomic region containing nine genes (Kit, Kdr, Srd5a3, Tmeme165, Clock, Pdcl2, Nmu, Exoc1, and Cep135). We designated this mutant allele KitWS. Interestingly, homozygous mutants (KitWS/WS) showed a peri- implantation lethal phenotype. Expression analyses of these nine genes in blastocysts suggested that Exoc1 was a prime candidate for this phenotype. We produced Exoc1 knockout mice, and the same peri-implantation lethal phenotype was seen in Exoc1−/− embryos. In addition, the polygenic effect without Exoc1 was investigated in genome-edited KitWE mice carrying the Mb-scale deletion induced by the CRISPR/Cas9 system. As KitWE/WE embryos did not exhibit the abnormal phenotype, which was seen in KitWS/WS. We concluded that peri-implantation lethality in KitWS/WS was caused by a monogenic defect of Exoc1. There have been many reports of spontaneous mouse mutants with depigmentation phenotypes1, among which Kit gene mutant mouse strains have been investigated in detail. More than 150 Kit mutant strains, including 72 spontaneous mutants, are listed in the Mouse Genome Informatics (MGI) database (http:// www.informatics.jax.org). -
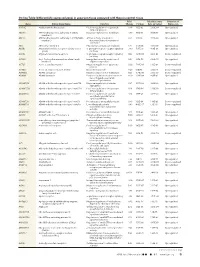
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

The Cancer Genome Atlas Dataset-Based Analysis of Aberrantly Expressed Genes by Geneanalytics in Thymoma Associated Myasthenia Gravis: Focusing on T Cells
2323 Original Article The Cancer Genome Atlas dataset-based analysis of aberrantly expressed genes by GeneAnalytics in thymoma associated myasthenia gravis: focusing on T cells Jianying Xi1#, Liang Wang1#, Chong Yan1, Jie Song1, Yang Song2, Ji Chen2, Yongjun Zhu2, Zhiming Chen2, Chun Jin3, Jianyong Ding3, Chongbo Zhao1,4 1Department of Neurology, 2Department of Thoracic Surgery, Huashan Hospital, Fudan University, Shanghai 200040, China; 3Department of Thoracic Surgery, Zhongshan Hospital, Fudan University, Shanghai 200030, China; 4Department of Neurology, Jing’an District Centre Hospital of Shanghai, Fudan University, Shanghai 200040, China Contributions: (I) Conception and design: J Xi, L Wang, C Zhao; (II) Administrative support: J Xi, L Wang, C Zhao; (III) Provision of study materials or patients: Y Song, Y Zhu, J Chen, Z Chen, C Jin, J Ding; (IV) Collection and assembly of data: C Jin, J Ding; (V) Data analysis and interpretation: C Yan, J Song; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Chongbo Zhao. Department of Neurology, Huashan Hospital, Fudan University, Shanghai 200040, China. Email: [email protected]. Background: Myasthenia gravis (MG) is a group of autoimmune disease which could be accompanied by thymoma. Many differences have been observed between thymoma-associated MG (TAMG) and non-MG thymoma (NMG). However, the molecular difference between them remained unknown. This study aimed to explore the differentially expressed genes (DEGs) between the two categories and to elucidate the possible pathogenesis of TAMG further. Methods: DEGs were calculated using the RNA-Sequencing data from 11 TAMG and 10 NMG in The Cancer Genome Atlas (TCGA) database. -

Oleuropein Activates Neonatal Neocortical Proteasomes
cells Article Oleuropein Activates Neonatal Neocortical Proteasomes, but Proteasome Gene Targeting by AAV9 Is Variable in a Clinically Relevant Piglet Model of Brain Hypoxia-Ischemia and Hypothermia Nagat El Demerdash 1, May W. Chen 2, Caitlin E. O’Brien 1, Shawn Adams 1, Ewa Kulikowicz 1, Lee J. Martin 3,† and Jennifer K. Lee 1,* 1 Department of Anesthesiology and Critical Care Medicine, Johns Hopkins University, Baltimore, MD 21287, USA; [email protected] (N.E.D.); [email protected] (C.E.O.); [email protected] (S.A.); [email protected] (E.K.) 2 Department of Pediatrics, School of Medicine, Johns Hopkins University, Baltimore, MD 21287, USA; [email protected] 3 Department of Pathology, Johns Hopkins University, Baltimore, MD 21287, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-410-955-6412; Fax: +1-410-502-5312 † Co-senior author (equal contribution). Abstract: Cerebral hypoxia-ischemia (HI) compromises the proteasome in a clinically relevant neona- tal piglet model. Protecting and activating proteasomes could be an adjunct therapy to hypothermia. Citation: El Demerdash, N.; Chen, We investigated whether chymotrypsin-like proteasome activity differs regionally and developmen- M.W.; O’Brien, C.E.; Adams, S.; tally in the neonatal brain. We also tested whether neonatal brain proteasomes can be modulated by Kulikowicz, E.; Martin, L.J.; Lee, J.K. oleuropein, an experimental pleiotropic neuroprotective drug, or by targeting a proteasome subunit Oleuropein Activates Neonatal gene using recombinant adeno-associated virus-9 (AAV). During post-HI hypothermia, we treated Neocortical Proteasomes, but piglets with oleuropein, used AAV-short hairpin RNA (shRNA) to knock down proteasome activator Proteasome Gene Targeting by AAV9 28γ (PA28γ), or enforced PA28γ using AAV-PA28γ with green fluorescent protein (GFP). -

Disruption of Entire Cables2 Locus Leads to Embryonic Lethality By
RESEARCH ARTICLE Disruption of entire Cables2 locus leads to embryonic lethality by diminished Rps21 gene expression and enhanced p53 pathway Tra Thi Huong Dinh1,2,3,4†, Hiroyoshi Iseki1,5†, Seiya Mizuno1,4†, Saori Iijima-Mizuno1,6, Yoko Tanimoto1, Yoko Daitoku1, Kanako Kato1, Yuko Hamada1, Ammar Shaker Hamed Hasan1,7, Hayate Suzuki1,7, Kazuya Murata1,4, Masafumi Muratani4,8, Masatsugu Ema9,10, Jun-Dal Kim11,12, Junji Ishida11, Akiyoshi Fukamizu11, Mitsuyasu Kato4,13, Satoru Takahashi1,4, Ken-ichi Yagami1, Valerie Wilson14, Ruth M Arkell15, Fumihiro Sugiyama1,4* 1Laboratory Animal Resource Center, Faculty of Medicine, University of Tsukuba, Tsukuba, Japan; 2Ph.D. Program in Human Biology, School of Integrative and Global Majors (SIGMA), University of Tsukuba, Tsukuba, Japan; 3Department of Traditional Medicine, University of Medicine and Pharmacy, Ho Chi Minh City, Viet Nam; 4Transborder Medical Research Center, Faculty of Medicine, University of Tsukuba, Tsukuba, Japan; 5International Institute for Integrative Sleep Medicine (WPI-IIIS), University of Tsukuba, Tsukuba, Japan; 6Experimental Animal Division, RIKEN BioResource Research Center, Tsukuba, Japan; 7Doctor’s Program in Biomedical Sciences, Graduate School of Comprehensive Human Science, University of Tsukuba, Tsukuba, Japan; 8Department of Genome Biology, Faculty of Medicine, University of Tsukuba, Tsukuba, Japan; 9Department of Stem Cells and Human Disease Models, Research Center for Animal Life Science, Shiga University of 10 *For correspondence: Medical Science, Otsu, Japan; -

DNA Methylation As a Mechanism for Caste System Determination in Solenopsis Invicta: Do the Genes Determine the Queen?
DNA Methylation as a Mechanism for Caste System Determination in Solenopsis invicta: Do the Genes Determine the Queen? By Kristen E. Walker A thesis submitted in partial fulfillment of the requirements of the University Honors Program University of South Florida St. Petersburg May 3, 2017 Thesis Director: David John, Ph.D. Lecturer, College of Arts and Sciences University Honors Program University of South Florida St. Petersburg CERTIFICATE OF APPROVAL ___________________________ Honors Thesis ___________________________ This is to certify that the Honors Thesis of Kristen E. Walker has been approved by the Examining Committee on May 3, 2017 as satisfying the thesis requirement of the University Honors Program Examining Committee: ___________________________ Thesis Director: David John, Ph.D. Lecturer, College of Arts and Sciences ____________________________ Thesis Committee Member: Thomas Smith, Ph.D. Honors Program Director and Associate Professor of Political Science, College of Arts and Sciences ___________________________ Thesis Committee Member: Debby Cassill, Ph.D. Associate Chairperson and Associate Professor, College of Arts and Sciences Abstract The emerging field of epigenetics seeks to study the interactions of the environment and other factors on gene expression in organisms. Epigenetic mechanisms involve modification to the DNA and/or surrounding structures, resulting in the activation or repression of gene expression. One epigenetic mechanism, DNA methylation, involves the addition of a methyl group to carbon 5 of cytosines in CpG dinucleotide regions in vertebrates and possibly other phyla. Vertebrate genomes tend to be globally methylated, whereas genes tend to be the target of methylation in insects. Genomic analyses of the order Hymenoptera revealed the evolutionary persistence of DNA methylation in social insects.