A Novel Mode of Capping Protein- Regulation by Twinfilin
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

PARSANA-DISSERTATION-2020.Pdf
DECIPHERING TRANSCRIPTIONAL PATTERNS OF GENE REGULATION: A COMPUTATIONAL APPROACH by Princy Parsana A dissertation submitted to The Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland July, 2020 © 2020 Princy Parsana All rights reserved Abstract With rapid advancements in sequencing technology, we now have the ability to sequence the entire human genome, and to quantify expression of tens of thousands of genes from hundreds of individuals. This provides an extraordinary opportunity to learn phenotype relevant genomic patterns that can improve our understanding of molecular and cellular processes underlying a trait. The high dimensional nature of genomic data presents a range of computational and statistical challenges. This dissertation presents a compilation of projects that were driven by the motivation to efficiently capture gene regulatory patterns in the human transcriptome, while addressing statistical and computational challenges that accompany this data. We attempt to address two major difficulties in this domain: a) artifacts and noise in transcriptomic data, andb) limited statistical power. First, we present our work on investigating the effect of artifactual variation in gene expression data and its impact on trans-eQTL discovery. Here we performed an in-depth analysis of diverse pre-recorded covariates and latent confounders to understand their contribution to heterogeneity in gene expression measurements. Next, we discovered 673 trans-eQTLs across 16 human tissues using v6 data from the Genotype Tissue Expression (GTEx) project. Finally, we characterized two trait-associated trans-eQTLs; one in Skeletal Muscle and another in Thyroid. Second, we present a principal component based residualization method to correct gene expression measurements prior to reconstruction of co-expression networks. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
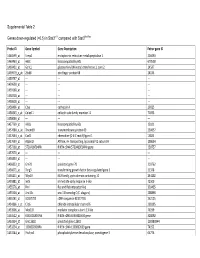
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Supplementary Table S1. Correlation Between the Mutant P53-Interacting Partners and PTTG3P, PTTG1 and PTTG2, Based on Data from Starbase V3.0 Database
Supplementary Table S1. Correlation between the mutant p53-interacting partners and PTTG3P, PTTG1 and PTTG2, based on data from StarBase v3.0 database. PTTG3P PTTG1 PTTG2 Gene ID Coefficient-R p-value Coefficient-R p-value Coefficient-R p-value NF-YA ENSG00000001167 −0.077 8.59e-2 −0.210 2.09e-6 −0.122 6.23e-3 NF-YB ENSG00000120837 0.176 7.12e-5 0.227 2.82e-7 0.094 3.59e-2 NF-YC ENSG00000066136 0.124 5.45e-3 0.124 5.40e-3 0.051 2.51e-1 Sp1 ENSG00000185591 −0.014 7.50e-1 −0.201 5.82e-6 −0.072 1.07e-1 Ets-1 ENSG00000134954 −0.096 3.14e-2 −0.257 4.83e-9 0.034 4.46e-1 VDR ENSG00000111424 −0.091 4.10e-2 −0.216 1.03e-6 0.014 7.48e-1 SREBP-2 ENSG00000198911 −0.064 1.53e-1 −0.147 9.27e-4 −0.073 1.01e-1 TopBP1 ENSG00000163781 0.067 1.36e-1 0.051 2.57e-1 −0.020 6.57e-1 Pin1 ENSG00000127445 0.250 1.40e-8 0.571 9.56e-45 0.187 2.52e-5 MRE11 ENSG00000020922 0.063 1.56e-1 −0.007 8.81e-1 −0.024 5.93e-1 PML ENSG00000140464 0.072 1.05e-1 0.217 9.36e-7 0.166 1.85e-4 p63 ENSG00000073282 −0.120 7.04e-3 −0.283 1.08e-10 −0.198 7.71e-6 p73 ENSG00000078900 0.104 2.03e-2 0.258 4.67e-9 0.097 3.02e-2 Supplementary Table S2. -

TITLE PAGE Oxidative Stress and Response to Thymidylate Synthase
Downloaded from molpharm.aspetjournals.org at ASPET Journals on October 2, 2021 -Targeted -Targeted 1 , University of of , University SC K.W.B., South Columbia, (U.O., Carolina, This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. -

The Kinesin-8 Member Kif19 Alters Microtubule Dynamics, Suppresses
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.04.282657; this version posted September 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 2 3 4 The kinesin-8 member Kif19 alters microtubule dynamics, suppresses cell adhesion, and 5 promotes cancer cell invasion 6 Short title: Kif19, microtubule dynamics, cell adhesion, and cancer invasion 7 8 Samuel C. Eisenberg1, Abhinav Dey2, Rayna Birnbaum1, David J. Sharp1* 9 10 11 12 1 Physiology and Biophysics, Albert Einstein College of Medicine, Bronx, New York, United 13 States of America 14 2 MicroCures, Albert Einstein College of Medicine, Bronx, New York, United States of America 15 16 17 * Corresponding Author 18 Email: [email protected] (DJS) 19 20 21 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.09.04.282657; this version posted September 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 22 Abstract 23 Metastasis is one of the deadliest aspects of cancer. Initial Metastatic spread is 24 dependent on the detachment and dissemination of cells from a parent tumor, and invasion into 25 the surrounding tissue. In this study, we characterize the kinesin-8 member Kif19 as a promoter 26 of cancer cell invasion that suppresses cell-cell adherens junctions and cell-matrix focal 27 adhesions. -

A Novel Mode of Capping Protein-Regulation by Twinfilin
Washington University School of Medicine Digital Commons@Becker Open Access Publications 10-23-2018 A novel mode of capping protein-regulation by Twinfilin Adam B. Johnston Denise M. Hilton Patrick McConnell Britney Johnson Meghan T. Harris See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Authors Adam B. Johnston, Denise M. Hilton, Patrick McConnell, Britney Johnson, Meghan T. Harris, Avital Simone, Gaya K. Amarasinghe, John A. Cooper, and Bruce L. Goode RESEARCH ARTICLE A novel mode of capping protein- regulation by twinfilin Adam B Johnston1†, Denise M Hilton1†, Patrick McConnell2, Britney Johnson3, Meghan T Harris1, Avital Simone1, Gaya K Amarasinghe3, John A Cooper2, Bruce L Goode1* 1Department of Biology, Rosenstiel Basic Medical Science Research Center, Brandeis University, Waltham, United States; 2Department of Biochemistry and Molecular Biophysics, Washington University, St Louis, United states; 3Department of Pathology and Immunology, Washington University, St Louis, United States Abstract Cellular actin assembly is controlled at the barbed ends of actin filaments, where capping protein (CP) limits polymerization. Twinfilin is a conserved in vivo binding partner of CP, yet the significance of this interaction has remained a mystery. Here, we discover that the C-terminal tail of Twinfilin harbors a CP-interacting (CPI) motif, identifying it as a novel CPI-motif protein. Twinfilin and the CPI-motif protein CARMIL have overlapping binding sites on CP. Further, Twinfilin binds competitively with CARMIL to CP, protecting CP from barbed-end displacement by CARMIL. Twinfilin also accelerates dissociation of the CP inhibitor V-1, restoring CP to an active capping state. -

IDENTIFICATION and CHARACTERIZATION of ACTIN-REGULATORY PROTEINS in the HAIR CELL's CUTICULAR PLATE by LANA MARY POLLOCK Subm
IDENTIFICATION AND CHARACTERIZATION OF ACTIN-REGULATORY PROTEINS IN THE HAIR CELL’S CUTICULAR PLATE by LANA MARY POLLOCK Submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy Dissertation advisor: Brian M. McDermott Jr., Ph.D. Department of Genetics and Genome Sciences CASE WESTERN RESERVE UNIVERSITY January 2016 Case Western Reserve University School of Graduate Studies We, the thesis committee, hereby approve the thesis/dissertation of Lana Pollock, candidate for the degree of Doctor of Philosophy (PhD).* (signed)_________Zhenghe Wang, Ph.D._________________ (chair of committee) ___________Brian McDermott, Ph.D._______________ ___________ Hua Lou, Ph.D._____________________ ___________Stephen Maricich, Ph.D., M.D.___________ ___________Anthony Wynshaw-Boris, Ph.D., M.D._____ Date of defense_____September 8th, 2015_______________ *we also certify that written approval has been obtained for release of any proprietary material contained therein 2 This thesis is dedicated to Daniel Margevicius. Thank you for your unwavering love and support. Ačiū!! 3 Table of contents List of Tables ........................................................................................................ 7 List of Figures ....................................................................................................... 8 List of abbreviations ............................................................................................ 13 Abstract ............................................................................................................. -

Kif18a Regulates Sirt2-Mediated Tubulin Acetylation for Spindle
Tang et al. Cell Div (2018) 13:9 https://doi.org/10.1186/s13008-018-0042-4 Cell Division RESEARCH Open Access Kif18a regulates Sirt2‑mediated tubulin acetylation for spindle organization during mouse oocyte meiosis Feng Tang, Meng‑Hao Pan, Xiang Wan, Yujie Lu, Yu Zhang and Shao‑Chen Sun* Abstract Background: During oocyte meiosis, the cytoskeleton dynamics, especially spindle organization, are critical for chro‑ mosome congression and segregation. However, the roles of the kinesin superfamily in this process are still largely unknown. Results: In the present study, Kif18a, a member of the kinesin-8 family, regulated spindle organization through its efects on tubulin acetylation in mouse oocyte meiosis. Our results showed that Kif18a is expressed and mainly local‑ ized in the spindle region. Knock down of Kif18a caused the failure of frst polar body extrusion, dramatically afecting spindle organization and resulting in severe chromosome misalignment. Further analysis showed that the disrup‑ tion of Kif18a caused an increase in acetylated tubulin level, which might be the reason for the spindle organization defects after Kif18a knock down in oocyte meiosis, and the decreased expression of deacetylase Sirt2 was found after Kif18a knock down. Moreover, microinjections of tubulin K40R mRNA, which could induce tubulin deacetylation, pro‑ tected the oocytes from the efects of Kif18a downregulation, resulting in normal spindle morphology in Kif18a-knock down oocytes. Conclusions: Taken together, our results showed that Kif18a afected Sirt2-mediated tubulin acetylation level for spindle organization during mouse oocyte meiosis. Our results not only revealed the critical efect of Kif18a on micro‑ tubule stability, but also extended our understanding of kinesin activity in meiosis. -

Selective and ATP‐Competitive Kinesin KIF18A Inhibitor Suppresses
Received: 16 October 2019 | Revised: 13 February 2020 | Accepted: 6 March 2020 DOI: 10.1111/jcmm.15200 ORIGINAL ARTICLE Selective and ATP-competitive kinesin KIF18A inhibitor suppresses the replication of influenza A virus Yong-Bin Cho1 | Sungguan Hong2 | Kyung-Won Kang3 | Ji-Hun Kang1 | Sang- Myeong Lee3 | Young-Jin Seo1 1Department of Life Science, Chung-Ang University, Seoul, South Korea Abstract 2Department of Chemistry, Chung-Ang The influenza virus is one of the major public health threats. However, the devel- University, Seoul, South Korea opment of efficient vaccines and therapeutic drugs to combat this virus is greatly 3Division of Biotechnology, College of Environmental and Bioresources, Jeonbuk limited by its frequent genetic mutations. Because of this, targeting the host factors National University, Iksan, South Korea required for influenza virus replication may be a more effective strategy for inhibit- Correspondence ing a broader spectrum of variants. Here, we demonstrated that inhibition of a motor Sang-Myeong Lee, Division of protein kinesin family member 18A (KIF18A) suppresses the replication of the influ- Biotechnology, College of Environmental and Bioresources Sciences, Jeonbuk enza A virus (IAV). The expression of KIF18A in host cells was increased following National University, Gobong-ro 79, Iksan-si, IAV infection. Intriguingly, treatment with the selective and ATP-competitive mitotic Jeollabuk-do 570-752, Korea. Email: [email protected] kinesin KIF18A inhibitor BTB-1 substantially decreased the expression of viral RNAs and proteins, and the production of infectious viral particles, while overexpression Young-Jin Seo, Department of Life Science, College of Natural Sciences, Chung-Ang of KIF18A enhanced the replication of IAV. -

The Proximal End of Mouse Chromosome 17: New Molecular Markers Identify a Deletion Associatedwith Quaking"'"""
Copyright 0 1992 by the Genetics Societyof America The Proximal End of Mouse Chromosome 17: New Molecular Markers Identify a Deletion AssociatedWith quaking"'""" Thomas Ebersole, Okkyung Rho and KarenArtzt Department of Zoology, The University of Texas, Austin, Texas 78712-1064 Manuscript received November 4, 1991 Accepted for publication January10, 1992 ABSTRACT Five randomly identified cosmids have been mapped proximal to the Leh66D locus on mouse chromosome 17. Two of these cosmids, AulO and Aull9, map near the neurological mutationquaking. Au119 is deleted in qk-blt/qk~ab'cDNA, whereas AulO is not. Au76 maps to a gene-rich region near the Tme locus. The Au76 locus encodes a member of a low copy gene family expressed in embryos, the adult central nervous system and testis. A second member of this family has been mapped to chromosome 15 near c-sis (PDGF-B). At the centromeric end of chromosome 17, Au116 maps near the Tu1 locus, and along with Au217rs identifies a region of unusually high recombinational activity between t-haplotypes and wild-type chromosomes. Au217I and II map to the large inverted repeats found at the proximal end of the wild-type chromosome. In addition, the Au2171 and/or II loci encode testis transcripts not expressed from t-haplotypes. HE attention given the proximal third of mouse and classical recombination analysis, many markers T chromosome 17 derives, in large part, from the have been mapped into the proximal end. The distri- analysis of t-haplotypes, a variant form found in wild bution of markers is, however, quite variable. The populations of mice. -

Post-Transcriptionally Impaired De Novo Mutations Contribute to The
bioRxiv preprint doi: https://doi.org/10.1101/175844; this version posted November 26, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Post-transcriptionally impaired de novo mutations 2 contribute to the genetic etiology of four neuropsychiatric 3 disorders 4 5 Fengbiao Mao1,2¶, Lu Wang3¶, Xiaolu Zhao2, Zhongshan Li4, Luoyuan Xiao5, 6 Rajesh C. Rao2, Jinchen Li4, Huajing Teng1*, Xin He6*, and Zhong Sheng Sun1,4* 7 8 1 Beijing Institutes of Life Science, Chinese Academy of Sciences, Beijing 100101, 9 China. 10 2 Department of Pathology, University of Michigan, Ann Arbor, MI 48109, USA. 11 3 Institute of Life Science, Southeast University, Nanjing 210096, China. 12 4 Institute of Genomic Medicine, Wenzhou Medical University, Wenzhou 325027, 13 China 14 5 Department of Computer Science and Technology, Tsinghua University, Beijing 15 100084, China. 16 6 Department of Human Genetics, University of Chicago, Chicago, IL, USA. 17 18 ¶These authors contributed equally to this work 19 * Corresponding authors 20 E-mail: 21 [email protected] (Z.S.S.) 22 [email protected] (X.H.) 23 [email protected] (H.T.) 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/175844; this version posted November 26, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity.