Dinoflagellate Nucleus Contains an Extensive Endomembrane Network, the Nuclear Net
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

This Article Appeared in a Journal Published by Elsevier. the Attached
This article appeared in a journal published by Elsevier. The attached copy is furnished to the author for internal non-commercial research and education use, including for instruction at the authors institution and sharing with colleagues. Other uses, including reproduction and distribution, or selling or licensing copies, or posting to personal, institutional or third party websites are prohibited. In most cases authors are permitted to post their version of the article (e.g. in Word or Tex form) to their personal website or institutional repository. Authors requiring further information regarding Elsevier’s archiving and manuscript policies are encouraged to visit: http://www.elsevier.com/copyright Author's personal copy ARTICLE IN PRESS European Journal of PROTISTOLOGY European Journal of Protistology 44 (2008) 299–307 www.elsevier.de/ejop Morphology and molecular phylogeny of Haplozoon praxillellae n. sp. (Dinoflagellata): A novel intestinal parasite of the maldanid polychaete Praxillella pacifica Berkeley Sonja RueckertÃ, Brian S. Leander Canadian Institute for Advanced Research, Program in Integrated Microbial Biodiversity, Departments of Botany and Zoology, University of British Columbia, Vancouver, BC, Canada V6T 1Z4 Received 11 December 2007; received in revised form 3 April 2008; accepted 5 April 2008 Abstract The genus Haplozoon comprises a group of endoparasites infecting the intestines of polychaete worms. Comparative studies using light microscopy, scanning and transmission electron microscopy, and small subunit rDNA have shown that these organisms are very unusual dinoflagellates. To date, there is only one species known from the Pacific Ocean, namely Haplozoon axiothellae Siebert. In this study, we describe Haplozoon praxillellae n. sp. from the intestine of the Pacific maldanid polychaete Praxillella pacifica Berkeley. -
Molecular Data and the Evolutionary History of Dinoflagellates by Juan Fernando Saldarriaga Echavarria Diplom, Ruprecht-Karls-Un
Molecular data and the evolutionary history of dinoflagellates by Juan Fernando Saldarriaga Echavarria Diplom, Ruprecht-Karls-Universitat Heidelberg, 1993 A THESIS SUBMITTED IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY in THE FACULTY OF GRADUATE STUDIES Department of Botany We accept this thesis as conforming to the required standard THE UNIVERSITY OF BRITISH COLUMBIA November 2003 © Juan Fernando Saldarriaga Echavarria, 2003 ABSTRACT New sequences of ribosomal and protein genes were combined with available morphological and paleontological data to produce a phylogenetic framework for dinoflagellates. The evolutionary history of some of the major morphological features of the group was then investigated in the light of that framework. Phylogenetic trees of dinoflagellates based on the small subunit ribosomal RNA gene (SSU) are generally poorly resolved but include many well- supported clades, and while combined analyses of SSU and LSU (large subunit ribosomal RNA) improve the support for several nodes, they are still generally unsatisfactory. Protein-gene based trees lack the degree of species representation necessary for meaningful in-group phylogenetic analyses, but do provide important insights to the phylogenetic position of dinoflagellates as a whole and on the identity of their close relatives. Molecular data agree with paleontology in suggesting an early evolutionary radiation of the group, but whereas paleontological data include only taxa with fossilizable cysts, the new data examined here establish that this radiation event included all dinokaryotic lineages, including athecate forms. Plastids were lost and replaced many times in dinoflagellates, a situation entirely unique for this group. Histones could well have been lost earlier in the lineage than previously assumed. -

Patrons De Biodiversité À L'échelle Globale Chez Les Dinoflagellés
! ! ! ! ! !"#$%&'%&'()!(*+!&'%&,-./01%*$0!2&30%**%&%!&4+*0%&).*0%& ! 0$'1&2(&3'!4!5&6(67&)!#2%&8)!9!:16()!;6136%2()!;&<)%=&3'!>?!@&<283! ! A%'=)83')!$2%! 45&/678&,9&:9;<6=! ! A6?% 6B3)8&% ()!7%2>) >) '()!%.*&>9&?-./01%*$0!2&30%**%&%!&4+*0%&).*0%! ! ! 0?C)3!>)!(2!3DE=)!4! ! @!!"#$%&'()*(+,%),-*$',#.(/(01.23*00*(40%+"0*(23*5(0*'( >A86B?7C9??D;&E?78<=68AFG9;&H7IA8;! ! ! ! 06?3)8?)!()!4!.+!FGH0!*+./! ! ;)<283!?8!C?%I!16#$6='!>)!4! ! 'I5&*6J987&$=9I8J!0&%!G(&=3)%!K2%>I!L6?8>23&68!M6%!N1)28!01&)81)!O0GKLN0PJ!A(I#6?3D!Q!H6I2?#)RS8&!! !!H2$$6%3)?%! 3I6B5&K78&37J?6J;LAJ!S8&<)%=&3'!>)!T)8E<)!Q!0?&==)! !!H2$$6%3)?%! 'I5&47IA87&468=I9;6IJ!032U&68)!V66(67&12!G8368!;6D%8!6M!W2$()=!Q!"32(&)! XY2#&823)?%! 3I6B5&,7I;&$=9HH788J!SAFZ,ZWH0!0323&68!V66(67&[?)!>)!@&(()M%281D)R=?%RF)%!Q!L%281)! XY2#&823)?%! 'I5&*7BB79?9&$A786J!;\WXZN,A)(276=J!"LHXFXH!!"#$%"&'"&(%")$*&+,-./0#1&Q!L%281)!!! !!!Z6R>&%)13)?%!>)!3DE=)! 'I5&)6?6HM78&>9&17IC7;J&SAFZ,ZWH0!0323&68!5&6(67&[?)!>)!H6=16MM!Q!L%281)! ! !!!!!!!!!;&%)13)?%!>)!3DE=)! ! ! ! "#$%&#'!()!*+,+-,*+./! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! ! Remerciements* ! Remerciements* A!l'issue!de!ce!travail!de!recherche!et!de!sa!rédaction,!j’ai!la!preuve!que!la!thèse!est!loin!d'être!un!travail! solitaire.! En! effet,! je! n'aurais! jamais! pu! réaliser! ce! travail! doctoral! sans! le! soutien! d'un! grand! nombre! de! personnes!dont!l’amitié,!la!générosité,!la!bonne!humeur%et%l'intérêt%manifestés%à%l'égard%de%ma%recherche%m'ont% permis!de!progresser!dans!cette!phase!délicate!de!«!l'apprentiGchercheur!».! -

In Situ Determination of Cellular DMSP and Pigment Quotas in a Prorocentrum Minimum Bloom Near the Falkland Islands Tyler Cyronaka, Erin O’Reilly B, Peter A
This article was downloaded by: [MUSC Library], [Peter Lee] On: 20 October 2014, At: 12:18 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 1072954 Registered office: Mortimer House, 37-41 Mortimer Street, London W1T 3JH, UK Advances in Oceanography and Limnology Publication details, including instructions for authors and subscription information: http://www.tandfonline.com/loi/taol20 In situ determination of cellular DMSP and pigment quotas in a Prorocentrum minimum bloom near the Falkland Islands Tyler Cyronaka, Erin O’Reilly b, Peter A. Leeb & Giacomo R. DiTulliob a Scripps Institution of Oceanography, UC San Diego, 9500 Gilman Drive, 0244, La Jolla, CA 92093-0244 b Grice and Hollings Marine Laboratories, College of Charleston, Charleston, South Carolina 29412, USA Published online: 16 Oct 2014. To cite this article: Tyler Cyronak, Erin O’Reilly, Peter A. Lee & Giacomo R. DiTullio (2014): In situ determination of cellular DMSP and pigment quotas in a Prorocentrum minimum bloom near the Falkland Islands, Advances in Oceanography and Limnology, DOI: 10.1080/19475721.2014.968620 To link to this article: http://dx.doi.org/10.1080/19475721.2014.968620 PLEASE SCROLL DOWN FOR ARTICLE Taylor & Francis makes every effort to ensure the accuracy of all the information (the “Content”) contained in the publications on our platform. However, Taylor & Francis, our agents, and our licensors make no representations or warranties whatsoever as to the accuracy, completeness, or suitability for any purpose of the Content. Any opinions and views expressed in this publication are the opinions and views of the authors, and are not the views of or endorsed by Taylor & Francis. -

Trait Changes Induced by Species Interactions in Two Phenotypically Distinct Strains of a Marine Dinoflagellate
The ISME Journal (2016) 10, 2658–2668 © 2016 International Society for Microbial Ecology All rights reserved 1751-7362/16 www.nature.com/ismej ORIGINAL ARTICLE Trait changes induced by species interactions in two phenotypically distinct strains of a marine dinoflagellate Sylke Wohlrab, Urban Tillmann, Allan Cembella and Uwe John Alfred Wegener Institute, Helmholtz Centre for Polar and Marine Research, Section Ecological Chemistry, Bremerhaven, Germany Populations of the toxigenic marine dinoflagellate Alexandrium are composed of multiple genotypes that display phenotypic variation for traits known to influence top-down processes, such as the ability to lyse co-occurring competitors and prospective grazers. We performed a detailed molecular analysis of species interactions to determine how different genotypes perceive and respond to other species. In a controlled laboratory culture study, we exposed two A. fundyense strains that differ in their capacity to produce lytic compounds to the dinoflagellate grazer Polykrikos kofoidii, and analyzed transcriptomic changes during this interaction. Approximately 5% of all analyzed genes were differentially expressed between the two Alexandrium strains under control conditions (without grazer presence) with fold-change differences that were proportionally higher than those observed in grazer treatments. Species interactions led to the genotype-specific expression of genes involved in endocytotic processes, cell cycle control and outer membrane properties, and signal transduction and gene expression -

University of Oklahoma
UNIVERSITY OF OKLAHOMA GRADUATE COLLEGE MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION SUBMITTED TO THE GRADUATE FACULTY in partial fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY By JOSHUA THOMAS COOPER Norman, Oklahoma 2017 MACRONUTRIENTS SHAPE MICROBIAL COMMUNITIES, GENE EXPRESSION AND PROTEIN EVOLUTION A DISSERTATION APPROVED FOR THE DEPARTMENT OF MICROBIOLOGY AND PLANT BIOLOGY BY ______________________________ Dr. Boris Wawrik, Chair ______________________________ Dr. J. Phil Gibson ______________________________ Dr. Anne K. Dunn ______________________________ Dr. John Paul Masly ______________________________ Dr. K. David Hambright ii © Copyright by JOSHUA THOMAS COOPER 2017 All Rights Reserved. iii Acknowledgments I would like to thank my two advisors Dr. Boris Wawrik and Dr. J. Phil Gibson for helping me become a better scientist and better educator. I would also like to thank my committee members Dr. Anne K. Dunn, Dr. K. David Hambright, and Dr. J.P. Masly for providing valuable inputs that lead me to carefully consider my research questions. I would also like to thank Dr. J.P. Masly for the opportunity to coauthor a book chapter on the speciation of diatoms. It is still such a privilege that you believed in me and my crazy diatom ideas to form a concise chapter in addition to learn your style of writing has been a benefit to my professional development. I’m also thankful for my first undergraduate research mentor, Dr. Miriam Steinitz-Kannan, now retired from Northern Kentucky University, who was the first to show the amazing wonders of pond scum. Who knew that studying diatoms and algae as an undergraduate would lead me all the way to a Ph.D. -

Grazing Impacts of the Heterotrophic Dinoflagellate Polykrikos Kofoidii on a Bloom of Gymnodinium Catenatum
AQUATIC MICROBIAL ECOLOGY Published April 30 Aquat Microb Ecol NOTE Grazing impacts of the heterotrophic dinoflagellate Polykrikos kofoidii on a bloom of Gymnodinium catenatum Yukihiko Matsuyama'f*,Masahide Miyamoto2, Yuichi ~otani' 'National Research Institute of Fisheries and Environment of Inland Sea, Maruishi, Ohno, Saeki, Hiroshima 739-0452, Japan 2KumamotoAriake Fisheries Direction Office, Iwasaki, Tamana, Kumamoto 865-0016, Japan ABSTRACT: In 1998, a red tide of the paralytic shellfish an assessment of the natural population of G. catena- poisoning (PSP)-producing dinoflagellate Gymnodinium cate- turn coupled with a laboratory incubation experiment naturn Graham occurred in Yatsushiro Sea, western Japan. to evaluate the bloom fate. We present data showing The dramatic decline of dominant G. catenatum cells oc- curred during the field and laboratory assessments, accompa- considerable predation by the pseudocolonial hetero- nied with growth of the heterotrophic dinoflagellate Poly- trophic dinoflagellate Polykrikos kofoidii Chatton on knkos kofoidii Chatton. Microscopic observations on both the dominant G. catenatum population, and discuss field and laboratory cultured bloom water revealed that the ecological importance of the genus Polykrikos and >50% of P. kofoidii predated on the natural population of G. catenaturn, and 1 to 8 G. catenatum cells were found in its grazing impact on harmful algal blooms. food vacuoles of P. kofoidii pseudocolonies. Our results sug- Materials and methods. Filed population surveys: gest that predation by P. kofoidii contributes to the cessation The Gymnodinium catenatum bloom occurred from 19 of a G. catenatum bloom. January to 5 February in Miyano-Gawachi Bay, west- ern Yatsushiro Sea, Kyushu Island (Fig. 1). Five cruises KEY WORDS: PSP - Gymnodimurn catenatum . -

A Parasite of Marine Rotifers: a New Lineage of Dinokaryotic Dinoflagellates (Dinophyceae)
Hindawi Publishing Corporation Journal of Marine Biology Volume 2015, Article ID 614609, 5 pages http://dx.doi.org/10.1155/2015/614609 Research Article A Parasite of Marine Rotifers: A New Lineage of Dinokaryotic Dinoflagellates (Dinophyceae) Fernando Gómez1 and Alf Skovgaard2 1 Laboratory of Plankton Systems, Oceanographic Institute, University of Sao˜ Paulo, Prac¸a do Oceanografico´ 191, Cidade Universitaria,´ 05508-900 Butanta,˜ SP, Brazil 2Department of Veterinary Disease Biology, University of Copenhagen, Stigbøjlen 7, 1870 Frederiksberg C, Denmark Correspondence should be addressed to Fernando Gomez;´ [email protected] Received 11 July 2015; Accepted 27 August 2015 Academic Editor: Gerardo R. Vasta Copyright © 2015 F. Gomez´ and A. Skovgaard. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Dinoflagellate infections have been reported for different protistan and animal hosts. We report, for the first time, the association between a dinoflagellate parasite and a rotifer host, tentatively Synchaeta sp. (Rotifera), collected from the port of Valencia, NW Mediterranean Sea. The rotifer contained a sporangium with 100–200 thecate dinospores that develop synchronically through palintomic sporogenesis. This undescribed dinoflagellate forms a new and divergent fast-evolved lineage that branches amongthe dinokaryotic dinoflagellates. 1. Introduction form independent lineages with no evident relation to other dinoflagellates [12]. In this study, we describe a new lineage of The alveolates (or Alveolata) are a major lineage of protists an undescribed parasitic dinoflagellate that largely diverged divided into three main phyla: ciliates, apicomplexans, and from other known dinoflagellates. -

(Alveolata) As Inferred from Hsp90 and Actin Phylogenies1
J. Phycol. 40, 341–350 (2004) r 2004 Phycological Society of America DOI: 10.1111/j.1529-8817.2004.03129.x EARLY EVOLUTIONARY HISTORY OF DINOFLAGELLATES AND APICOMPLEXANS (ALVEOLATA) AS INFERRED FROM HSP90 AND ACTIN PHYLOGENIES1 Brian S. Leander2 and Patrick J. Keeling Canadian Institute for Advanced Research, Program in Evolutionary Biology, Departments of Botany and Zoology, University of British Columbia, Vancouver, British Columbia, Canada Three extremely diverse groups of unicellular The Alveolata is one of the most biologically diverse eukaryotes comprise the Alveolata: ciliates, dino- supergroups of eukaryotic microorganisms, consisting flagellates, and apicomplexans. The vast phenotypic of ciliates, dinoflagellates, apicomplexans, and several distances between the three groups along with the minor lineages. Although molecular phylogenies un- enigmatic distribution of plastids and the economic equivocally support the monophyly of alveolates, and medical importance of several representative members of the group share only a few derived species (e.g. Plasmodium, Toxoplasma, Perkinsus, and morphological features, such as distinctive patterns of Pfiesteria) have stimulated a great deal of specula- cortical vesicles (syn. alveoli or amphiesmal vesicles) tion on the early evolutionary history of alveolates. subtending the plasma membrane and presumptive A robust phylogenetic framework for alveolate pinocytotic structures, called ‘‘micropores’’ (Cavalier- diversity will provide the context necessary for Smith 1993, Siddall et al. 1997, Patterson -
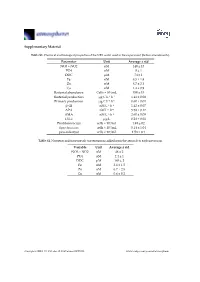
Supplementary Material Parameter Unit Average ± Std NO3 + NO2 Nm
Supplementary Material Table S1. Chemical and biological properties of the NRS water used in the experiment (before amendments). Parameter Unit Average ± std NO3 + NO2 nM 140 ± 13 PO4 nM 8 ± 1 DOC μM 74 ± 1 Fe nM 8.5 ± 1.8 Zn nM 8.7 ± 2.1 Cu nM 1.4 ± 0.9 Bacterial abundance Cells × 104/mL 350 ± 15 Bacterial production μg C L−1 h−1 1.41 ± 0.08 Primary production μg C L−1 h−1 0.60 ± 0.01 β-Gl nM L−1 h−1 1.42 ± 0.07 APA nM L−1 h−1 5.58 ± 0.17 AMA nM L−1·h−1 2.60 ± 0.09 Chl-a μg/L 0.28 ± 0.01 Prochlorococcus cells × 104/mL 1.49 ± 02 Synechococcus cells × 104/mL 5.14 ± 1.04 pico-eukaryot cells × 103/mL 1.58 × 0.1 Table S2. Nutrients and trace metals concentrations added from the aerosols to each mesocosm. Variable Unit Average ± std NO3 + NO2 nM 48 ± 2 PO4 nM 2.4 ± 1 DOC μM 165 ± 2 Fe nM 2.6 ± 1.5 Zn nM 6.7 ± 2.5 Cu nM 0.6 ± 0.2 Atmosphere 2019, 10, 358; doi:10.3390/atmos10070358 www.mdpi.com/journal/atmosphere Atmosphere 2019, 10, 358 2 of 6 Table S3. ANOVA test results between control, ‘UV-treated’ and ‘live-dust’ treatments at 20 h or 44 h, with significantly different values shown in bold. ANOVA df Sum Sq Mean Sq F Value p-value Chl-a 20 H 2, 6 0.03, 0.02 0.02, 0 4.52 0.0634 44 H 2, 6 0.02, 0 0.01, 0 23.13 0.002 Synechococcus Abundance 20 H 2, 7 8.23 × 107, 4.11 × 107 4.11 × 107, 4.51 × 107 0.91 0.4509 44 H 2, 7 5.31 × 108, 6.97 × 107 2.65 × 108, 1.16 × 107 22.84 0.0016 Prochlorococcus Abundance 20 H 2, 8 4.22 × 107, 2.11 × 107 2.11 × 107, 2.71 × 106 7.77 0.0216 44 H 2, 8 9.02 × 107, 1.47 × 107 4.51 × 107, 2.45 × 106 18.38 0.0028 Pico-eukaryote -

Recent Dinoflagellate Cysts from the Chesapeake Estuary (Maryland and Virginia, U.S.A.): Taxonomy and Ecological Preferences
Recent dinoflagellate cysts from the Chesapeake estuary (Maryland and Virginia, U.S.A.): taxonomy and ecological preferences. Tycho Van Hauwaert Academic year 2015–2016 Master’s dissertation submitted in partial fulfillment of the requirements for the degree of Master in Science in Geology Promotor: Prof. Dr. S. Louwye Co-promotor: Dr. K. Mertens Tutor: P. Gurdebeke Jury: Dr. T. Verleye, Dr. E. Verleyen Picture on the cover An exceptionally dense bloom of Alexandrium monilatum was observed in lower Chesapeake Bay along the north shore of the York River between Sarah's Creek and the Perrin River on 17 August 2015. Credit: W. Vogelbein/VIMS ii ACKNOWLEDGEMENTS First of all I want to thank my promoters, Prof. Dr. S. Louwye and Dr. K. Mertens. They introduced me into the wonderful world of dinoflagellates and the dinocysts due to the course Advanced Micropaleontology. I did not have hesitated long to choose a subject within the research unit of paleontology. Thank you for the proofreading, help with identification and many discussions. A special mention for Pieter Gurdebeke. This appreciation you can imagine as a 22-minutes standing ovation for the small talks and jokes only! If you include the assistance in the thesis, I would not dare to calculate the time of applause. I remember when we were discussing the subject during the fieldtrip to the Alps in September. We have come a long way and I am pleased with the result. Thank you very much for helping me with the preparation of slides, identification of dinocysts, some computer programs, proofreading of the different chapters and many more! When I am back from my trip to Canada, I would like to discuss it with a (small) bottle of beer. -

The Mitochondrial Genome and Transcriptome of the Basal
View metadata, citation and similar papers at core.ac.uk brought to you by CORE GBEprovided by PubMed Central The Mitochondrial Genome and Transcriptome of the Basal Dinoflagellate Hematodinium sp.: Character Evolution within the Highly Derived Mitochondrial Genomes of Dinoflagellates C. J. Jackson, S. G. Gornik, and R. F. Waller* School of Botany, University of Melbourne, Australia *Corresponding author: E-mail: [email protected]. Accepted: 12 November 2011 Abstract The sister phyla dinoflagellates and apicomplexans inherited a drastically reduced mitochondrial genome (mitochondrial DNA, mtDNA) containing only three protein-coding (cob, cox1, and cox3) genes and two ribosomal RNA (rRNA) genes. In apicomplexans, single copies of these genes are encoded on the smallest known mtDNA chromosome (6 kb). In dinoflagellates, however, the genome has undergone further substantial modifications, including massive genome amplification and recombination resulting in multiple copies of each gene and gene fragments linked in numerous combinations. Furthermore, protein-encoding genes have lost standard stop codons, trans-splicing of messenger RNAs (mRNAs) is required to generate complete cox3 transcripts, and extensive RNA editing recodes most genes. From taxa investigated to date, it is unclear when many of these unusual dinoflagellate mtDNA characters evolved. To address this question, we investigated the mitochondrial genome and transcriptome character states of the deep branching dinoflagellate Hematodinium sp. Genomic data show that like later-branching dinoflagellates Hematodinium sp. also contains an inflated, heavily recombined genome of multicopy genes and gene fragments. Although stop codons are also lacking for cox1 and cob, cox3 still encodes a conventional stop codon. Extensive editing of mRNAs also occurs in Hematodinium sp.