Microbial Partners in Health: Broadening Our Understanding of Host-Microbiome Relationships
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Unraveling the First Step in the Streptomyces Scabies Pathology: How Does the Pathogen Sense It Is in the Vicinity of a Living R
Unraveling the first step in the Streptomyces scabies pathology: How does the pathogen sense it is in the vicinity of a living root? By: Dianna Mojica A thesis submitted to the Department of Biology at California State University, Bakersfield in partial fulfilment for the degree of Master of Science in Biology Spring 2021 ii Copyright By Dianna Mojica 2021 i Table of Contents Acknowledgements v Abstract vi Keywords viii List of Figures ix Chapter 1 8 Introduction 8 Streptomyces, an unusual bacterial genus 8 Streptomyces as symbionts in interactions with plants and animals 10 Streptomyces scabies, a well-established and worldwide occurring plant pathogen 12 Literature Cited 16 Chapter 2 22 Abstract 22 Introduction 23 Materials and Methods 26 Bacterial strains and growth conditions used in this study 26 Thaxtomin A production in response to living or dead radish plant material 26 Thaxtomin A production in response to radish root exudates 28 Cellulase activity and thaxtomin A production in response to cellulose 29 S. scabies cannot distinguish between a living and a dead host plant to start thaxtomin A production 30 Literature Cited 39 Figures 43 Chapter 3 48 Conclusion 48 Literature Cited 50 iv Acknowledgements I thank my thesis advisor Dr. Francis for donating her time and sharing her knowledge on Streptomyces scabies as well as guiding me on how to write this thesis. I also thank Dr. Stokes and Dr. Keller for agreeing to be part of my thesis committee in such short notice. v Abstract Streptomyces is the largest genus within the phylum Actinobacteria. The genus contains Gram-positive bacteria with a complex life cycle resembling that of fungi and are distinctly recognized for their diverse secondary metabolites that have pharmaceutical, agricultural and industrial properties. -

Improved Taxonomy of the Genus Streptomyces
UNIVERSITEIT GENT Faculteit Wetenschappen Vakgroep Biochemie, Fysiologie & Microbiologie Laboratorium voor Microbiologie Improved taxonomy of the genus Streptomyces Benjamin LANOOT Scriptie voorgelegd tot het behalen van de graad van Doctor in de Wetenschappen (Biochemie) Promotor: Prof. Dr. ir. J. Swings Co-promotor: Dr. M. Vancanneyt Academiejaar 2004-2005 FACULTY OF SCIENCES ____________________________________________________________ DEPARTMENT OF BIOCHEMISTRY, PHYSIOLOGY AND MICROBIOLOGY UNIVERSITEIT LABORATORY OF MICROBIOLOGY GENT IMPROVED TAXONOMY OF THE GENUS STREPTOMYCES DISSERTATION Submitted in fulfilment of the requirements for the degree of Doctor (Ph D) in Sciences, Biochemistry December 2004 Benjamin LANOOT Promotor: Prof. Dr. ir. J. SWINGS Co-promotor: Dr. M. VANCANNEYT 1: Aerial mycelium of a Streptomyces sp. © Michel Cavatta, Academy de Lyon, France 1 2 2: Streptomyces coelicolor colonies © John Innes Centre 3: Blue haloes surrounding Streptomyces coelicolor colonies are secreted 3 4 actinorhodin (an antibiotic) © John Innes Centre 4: Antibiotic droplet secreted by Streptomyces coelicolor © John Innes Centre PhD thesis, Faculty of Sciences, Ghent University, Ghent, Belgium. Publicly defended in Ghent, December 9th, 2004. Examination Commission PROF. DR. J. VAN BEEUMEN (ACTING CHAIRMAN) Faculty of Sciences, University of Ghent PROF. DR. IR. J. SWINGS (PROMOTOR) Faculty of Sciences, University of Ghent DR. M. VANCANNEYT (CO-PROMOTOR) Faculty of Sciences, University of Ghent PROF. DR. M. GOODFELLOW Department of Agricultural & Environmental Science University of Newcastle, UK PROF. Z. LIU Institute of Microbiology Chinese Academy of Sciences, Beijing, P.R. China DR. D. LABEDA United States Department of Agriculture National Center for Agricultural Utilization Research Peoria, IL, USA PROF. DR. R.M. KROPPENSTEDT Deutsche Sammlung von Mikroorganismen & Zellkulturen (DSMZ) Braunschweig, Germany DR. -

Evolution of the Streptomycin and Viomycin Biosynthetic Clusters and Resistance Genes
University of Warwick institutional repository: http://go.warwick.ac.uk/wrap A Thesis Submitted for the Degree of PhD at the University of Warwick http://go.warwick.ac.uk/wrap/2773 This thesis is made available online and is protected by original copyright. Please scroll down to view the document itself. Please refer to the repository record for this item for information to help you to cite it. Our policy information is available from the repository home page. Evolution of the streptomycin and viomycin biosynthetic clusters and resistance genes Paris Laskaris, B.Sc. (Hons.) A thesis submitted to the University of Warwick for the degree of Doctor of Philosophy. Department of Biological Sciences, University of Warwick, Coventry, CV4 7AL September 2009 Contents List of Figures ........................................................................................................................ vi List of Tables ....................................................................................................................... xvi Abbreviations ........................................................................................................................ xx Acknowledgements .............................................................................................................. xxi Declaration .......................................................................................................................... xxii Abstract ............................................................................................................................. -

26Sor Em2rray /Mmucd
D4006- Gut Microbiota Analysis UC Davis MMPC - Microbiome & Host Response Core Contents 1 Methods: 1 1.1 Sequencing . .1 1.2 Data processing . .1 2 Summary of Findings: 2 2.1 Sequencing analysis . .2 2.2 Microbial diversity . .2 2.2.1 Alpha Diversity . .2 2.2.2 Beta Diversity . 10 2.3 Data analysis using taxa abundance data . 13 2.3.1 Stacked bar graphs of Taxa abundances at each level . 14 A Appendix 1 (Taxa Abundance Tables) 30 B Appendix 2 (Taxa removed from filtered datasets at each level) 37 References: 43 Core Contacts: Helen E. Raybould, Ph.D., Core Leader ([email protected]) Trina A. Knotts, Ph.D., Core Co-Leader ([email protected]) Michael L. Goodson, Ph.D., Core Scientist ([email protected]) Client(s): Kent Lloyd, DVM PhD ;MMRRC; UC Davis Project #: MBP-2079 MMRRC strain ID: MMRRC_043514 Animal Information: The strain was donated to the MMRRC by Russell Ray at Baylor College of Medicine. Fecal samples were obtained from animals housed under the care of Russell Ray at Baylor College of Medicine. 1 Methods: Brief Project Description: MMRRC strains are often contributed to the MMRRC to fulfill the resource sharing aspects of NIH grants. Since transporting mice to another facilty often causes a microbiota shift, having a record of the original fecal microbiota from the donor institution where the original phenotyping or testing was performed may prove helpful if a phenotype is lost after transfer. Several MMRRC mouse lines were selected for fecal microbiota profiling of the microbiota. Table 1: Animal-Strain Information X.SampleID TreatmentGroup Animal_ID Genotype Line Sex MMRRC.043514.Mx054 MMRRC.043514_Het_M Mx054 Het MMRRC.043514 M MMRRC.043514.Mx415 MMRRC.043514_Het_M Mx415 Het MMRRC.043514 M 1 1.1 Sequencing Frozen fecal or regional gut samples were shipped on dry ice to UC Davis MMPC and Host Microbe Systems Biology Core. -

Study of Actinobacteria and Their Secondary Metabolites from Various Habitats in Indonesia and Deep-Sea of the North Atlantic Ocean
Study of Actinobacteria and their Secondary Metabolites from Various Habitats in Indonesia and Deep-Sea of the North Atlantic Ocean Von der Fakultät für Lebenswissenschaften der Technischen Universität Carolo-Wilhelmina zu Braunschweig zur Erlangung des Grades eines Doktors der Naturwissenschaften (Dr. rer. nat.) genehmigte D i s s e r t a t i o n von Chandra Risdian aus Jakarta / Indonesien 1. Referent: Professor Dr. Michael Steinert 2. Referent: Privatdozent Dr. Joachim M. Wink eingereicht am: 18.12.2019 mündliche Prüfung (Disputation) am: 04.03.2020 Druckjahr 2020 ii Vorveröffentlichungen der Dissertation Teilergebnisse aus dieser Arbeit wurden mit Genehmigung der Fakultät für Lebenswissenschaften, vertreten durch den Mentor der Arbeit, in folgenden Beiträgen vorab veröffentlicht: Publikationen Risdian C, Primahana G, Mozef T, Dewi RT, Ratnakomala S, Lisdiyanti P, and Wink J. Screening of antimicrobial producing Actinobacteria from Enggano Island, Indonesia. AIP Conf Proc 2024(1):020039 (2018). Risdian C, Mozef T, and Wink J. Biosynthesis of polyketides in Streptomyces. Microorganisms 7(5):124 (2019) Posterbeiträge Risdian C, Mozef T, Dewi RT, Primahana G, Lisdiyanti P, Ratnakomala S, Sudarman E, Steinert M, and Wink J. Isolation, characterization, and screening of antibiotic producing Streptomyces spp. collected from soil of Enggano Island, Indonesia. The 7th HIPS Symposium, Saarbrücken, Germany (2017). Risdian C, Ratnakomala S, Lisdiyanti P, Mozef T, and Wink J. Multilocus sequence analysis of Streptomyces sp. SHP 1-2 and related species for phylogenetic and taxonomic studies. The HIPS Symposium, Saarbrücken, Germany (2019). iii Acknowledgements Acknowledgements First and foremost I would like to express my deep gratitude to my mentor PD Dr. -

Diversity of Free-Living Nitrogen Fixing Bacteria in the Badlands of South Dakota Bibha Dahal South Dakota State University
South Dakota State University Open PRAIRIE: Open Public Research Access Institutional Repository and Information Exchange Theses and Dissertations 2016 Diversity of Free-living Nitrogen Fixing Bacteria in the Badlands of South Dakota Bibha Dahal South Dakota State University Follow this and additional works at: http://openprairie.sdstate.edu/etd Part of the Bacteriology Commons, and the Environmental Microbiology and Microbial Ecology Commons Recommended Citation Dahal, Bibha, "Diversity of Free-living Nitrogen Fixing Bacteria in the Badlands of South Dakota" (2016). Theses and Dissertations. 688. http://openprairie.sdstate.edu/etd/688 This Thesis - Open Access is brought to you for free and open access by Open PRAIRIE: Open Public Research Access Institutional Repository and Information Exchange. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of Open PRAIRIE: Open Public Research Access Institutional Repository and Information Exchange. For more information, please contact [email protected]. DIVERSITY OF FREE-LIVING NITROGEN FIXING BACTERIA IN THE BADLANDS OF SOUTH DAKOTA BY BIBHA DAHAL A thesis submitted in partial fulfillment of the requirements for the Master of Science Major in Biological Sciences Specialization in Microbiology South Dakota State University 2016 iii ACKNOWLEDGEMENTS “Always aim for the moon, even if you miss, you’ll land among the stars”.- W. Clement Stone I would like to express my profuse gratitude and heartfelt appreciation to my advisor Dr. Volker Brӧzel for providing me a rewarding place to foster my career as a scientist. I am thankful for his implicit encouragement, guidance, and support throughout my research. This research would not be successful without his guidance and inspiration. -

Ep 2434019 A1
(19) & (11) EP 2 434 019 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 28.03.2012 Bulletin 2012/13 C12N 15/82 (2006.01) C07K 14/395 (2006.01) C12N 5/10 (2006.01) G01N 33/50 (2006.01) (2006.01) (2006.01) (21) Application number: 11160902.0 C07K 16/14 A01H 5/00 C07K 14/39 (2006.01) (22) Date of filing: 21.07.2004 (84) Designated Contracting States: • Kamlage, Beate AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 12161, Berlin (DE) HU IE IT LI LU MC NL PL PT RO SE SI SK TR • Taman-Chardonnens, Agnes A. 1611, DS Bovenkarspel (NL) (30) Priority: 01.08.2003 EP 03016672 • Shirley, Amber 15.04.2004 PCT/US2004/011887 Durham, NC 27703 (US) • Wang, Xi-Qing (62) Document number(s) of the earlier application(s) in Chapel Hill, NC 27516 (US) accordance with Art. 76 EPC: • Sarria-Millan, Rodrigo 04741185.5 / 1 654 368 West Lafayette, IN 47906 (US) • McKersie, Bryan D (27) Previously filed application: Cary, NC 27519 (US) 21.07.2004 PCT/EP2004/008136 • Chen, Ruoying Duluth, GA 30096 (US) (71) Applicant: BASF Plant Science GmbH 67056 Ludwigshafen (DE) (74) Representative: Heistracher, Elisabeth BASF SE (72) Inventors: Global Intellectual Property • Plesch, Gunnar GVX - C 6 14482, Potsdam (DE) Carl-Bosch-Strasse 38 • Puzio, Piotr 67056 Ludwigshafen (DE) 9030, Mariakerke (Gent) (BE) • Blau, Astrid Remarks: 14532, Stahnsdorf (DE) This application was filed on 01-04-2011 as a • Looser, Ralf divisional application to the application mentioned 13158, Berlin (DE) under INID code 62. -

Genomic and Phylogenomic Insights Into the Family Streptomycetaceae Lead to Proposal of Charcoactinosporaceae Fam. Nov. and 8 No
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.08.193797; this version posted July 8, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Genomic and phylogenomic insights into the family Streptomycetaceae 2 lead to proposal of Charcoactinosporaceae fam. nov. and 8 novel genera 3 with emended descriptions of Streptomyces calvus 4 Munusamy Madhaiyan1, †, * Venkatakrishnan Sivaraj Saravanan2, † Wah-Seng See-Too3, † 5 1Temasek Life Sciences Laboratory, 1 Research Link, National University of Singapore, 6 Singapore 117604; 2Department of Microbiology, Indira Gandhi College of Arts and Science, 7 Kathirkamam 605009, Pondicherry, India; 3Division of Genetics and Molecular Biology, 8 Institute of Biological Sciences, Faculty of Science, University of Malaya, Kuala Lumpur, 9 Malaysia 10 *Corresponding author: Temasek Life Sciences Laboratory, 1 Research Link, National 11 University of Singapore, Singapore 117604; E-mail: [email protected] 12 †All these authors have contributed equally to this work 13 Abstract 14 Streptomycetaceae is one of the oldest families within phylum Actinobacteria and it is large and 15 diverse in terms of number of described taxa. The members of the family are known for their 16 ability to produce medically important secondary metabolites and antibiotics. In this study, 17 strains showing low 16S rRNA gene similarity (<97.3 %) with other members of 18 Streptomycetaceae were identified and subjected to phylogenomic analysis using 33 orthologous 19 gene clusters (OGC) for accurate taxonomic reassignment resulted in identification of eight 20 distinct and deeply branching clades, further average amino acid identity (AAI) analysis showed 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.07.08.193797; this version posted July 8, 2020. -

Induction of Secondary Metabolism Across Actinobacterial Genera
Induction of secondary metabolism across actinobacterial genera A thesis submitted for the award Doctor of Philosophy at Flinders University of South Australia Rio Risandiansyah Department of Medical Biotechnology Faculty of Medicine, Nursing and Health Sciences Flinders University 2016 TABLE OF CONTENTS TABLE OF CONTENTS ............................................................................................ ii TABLE OF FIGURES ............................................................................................. viii LIST OF TABLES .................................................................................................... xii SUMMARY ......................................................................................................... xiii DECLARATION ...................................................................................................... xv ACKNOWLEDGEMENTS ...................................................................................... xvi Chapter 1. Literature review ................................................................................. 1 1.1 Actinobacteria as a source of novel bioactive compounds ......................... 1 1.1.1 Natural product discovery from actinobacteria .................................... 1 1.1.2 The need for new antibiotics ............................................................... 3 1.1.3 Secondary metabolite biosynthetic pathways in actinobacteria ........... 4 1.1.4 Streptomyces genetic potential: cryptic/silent genes .......................... -
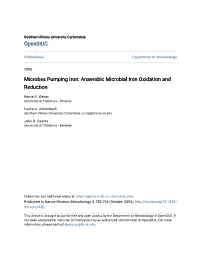
Fe(II) Fe(III)
Southern Illinois University Carbondale OpenSIUC Publications Department of Microbiology 2006 Microbes Pumping Iron: Anaerobic Microbial Iron Oxidation and Reduction Karrie A. Weber University of California - Berkeley Laurie A. Achenbach Southern Illinois University Carbondale, [email protected] John D. Coates University of California - Berkeley Follow this and additional works at: https://opensiuc.lib.siu.edu/micro_pubs Published in Nature Reviews Microbiology 4, 752-764 (October 2006), http://dx.doi.org/10.1038/ nrmicro1490. This Article is brought to you for free and open access by the Department of Microbiology at OpenSIUC. It has been accepted for inclusion in Publications by an authorized administrator of OpenSIUC. For more information, please contact [email protected]. 1 Microbes Pumping Iron: 2 Anaerobic Microbial Iron Oxidation and Reduction 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 Karrie A. Weber1, Laurie A. Achenbach2, and John D. Coates1* 18 19 20 21 22 23 1Department of Plant and Microbial Biology 24 271 Koshland Hall 25 University of California, Berkeley 26 Berkeley, CA 94720 27 28 29 2Department of Microbiology 30 Southern Illinois University 31 Carbondale, IL 62901 32 33 34 35 Corresponding Author 36 Tel: (510) 643-8455 37 Fax: (510) 642-4995 38 e-mail: [email protected] 39 1 1 Abstract 2 Iron (Fe) has long been a recognized physiological requirement for life, yet its role for a 3 broad diversity of microorganisms persisting in water, soils, and sediments extends well-beyond 4 a nutritional necessity. Microorganisms from within both the domain Archaea and Bacteria are 5 capable of metabolically exploiting the favorable redox potential between the Fe(III)/Fe(II) 6 couple. -

Supplementary Material Impacts of Radiation on the Bacterial And
Supplementary Material Impacts of radiation on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone Rachael E. Antwis, Nicholas A. Beresford, Joseph A. Jackson, Ross Fawkes, Catherine L. Barnett, Elaine Potter, Lee Walker, Sergey Gaschak, Michael D. Wood Table S1 Sample sizes of host species used in the study, and associated sex, total absorbed dose rate, burn and site categories for these samples. Animals estimated to receive total absorbed dose rates of <4 µGy h-1 were assigned ‘low’, those with estimated dose rates of 4-42 µGy h-1 assigned ‘medium’, and those >42 µGy h-1 assigned to the ‘high’ category. All samples from 2017 were collected within the Red Forest but from areas that had experienced different degrees of damage from forest fires. Samples from 2018 were either collected within or outside the Red Forest. Sampling information Sex Total absorbed dose category Burn category Site category Year Sample Host species Total n Female Male Unidentified Low Medium High Burnt Burnt Unburnt Inside Red Outside type (regrowth) (minimal Forest Red regrowth) Forest 2017 Faeces Bank vole 22 5 16 1 0 0 22 3 1 18 All - 2017 Faeces Striped field mouse 29 10 18 1 0 7 22 15 0 14 All - 2017 Faeces Wood mouse 27 6 20 1 0 0 27 2 25 0 All - 2017 Faeces Yellow-necked mouse 58 28 29 1 0 7 51 19 17 22 All - 2018 Gut Bank vole 142 59 83 0 42 64 36 - - - 34 108 (caecum) RESULTS Differences in community composition between host species and sample types Wood mice and yellow-necked mice had a similar bacterial community composition to one another (Figure 1a), with the exception of the family Helicobacteraceae (Z = 2.982, p = 0.023; Figure 1c). -

Exploring the Potential of Antibiotic Production from Rare Actinobacteria by Whole-Genome Sequencing and Guided MS/MS Analysis
fmicb-11-01540 July 27, 2020 Time: 14:51 # 1 ORIGINAL RESEARCH published: 15 July 2020 doi: 10.3389/fmicb.2020.01540 Exploring the Potential of Antibiotic Production From Rare Actinobacteria by Whole-Genome Sequencing and Guided MS/MS Analysis Dini Hu1,2, Chenghang Sun3, Tao Jin4, Guangyi Fan4, Kai Meng Mok2, Kai Li1* and Simon Ming-Yuen Lee5* 1 School of Ecology and Nature Conservation, Beijing Forestry University, Beijing, China, 2 Department of Civil and Environmental Engineering, Faculty of Science and Technology, University of Macau, Macau, China, 3 Institute of Medicinal Biotechnology, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing, China, 4 Beijing Genomics Institute, Shenzhen, China, 5 State Key Laboratory of Quality Research in Chinese Medicine, Institute of Chinese Medical Sciences, University of Macau, Macau, China Actinobacteria are well recognized for their production of structurally diverse bioactive Edited by: secondary metabolites, but the rare actinobacterial genera have been underexploited Sukhwan Yoon, for such potential. To search for new sources of active compounds, an experiment Korea Advanced Institute of Science combining genomic analysis and tandem mass spectrometry (MS/MS) screening and Technology, South Korea was designed to isolate and characterize actinobacterial strains from a mangrove Reviewed by: Hui Li, environment in Macau. Fourteen actinobacterial strains were isolated from the collected Jinan University, China samples. Partial 16S sequences indicated that they were from six genera, including Baogang Zhang, China University of Geosciences, Brevibacterium, Curtobacterium, Kineococcus, Micromonospora, Mycobacterium, and China Streptomyces. The isolate sp.01 showing 99.28% sequence similarity with a reference *Correspondence: rare actinobacterial species Micromonospora aurantiaca ATCC 27029T was selected for Kai Li whole genome sequencing.