Graph Neural Representational Learning of Rna Secondary Structures for Predicting Rna-Protein Interactions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Large-Scale Analysis of Genome and Transcriptome Alterations in Multiple Tumors Unveils Novel Cancer-Relevant Splicing Networks
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Research Large-scale analysis of genome and transcriptome alterations in multiple tumors unveils novel cancer-relevant splicing networks Endre Sebestyén,1,5 Babita Singh,1,5 Belén Miñana,1,2 Amadís Pagès,1 Francesca Mateo,3 Miguel Angel Pujana,3 Juan Valcárcel,1,2,4 and Eduardo Eyras1,4 1Universitat Pompeu Fabra, E08003 Barcelona, Spain; 2Centre for Genomic Regulation, E08003 Barcelona, Spain; 3Program Against Cancer Therapeutic Resistance (ProCURE), Catalan Institute of Oncology (ICO), Bellvitge Institute for Biomedical Research (IDIBELL), E08908 L’Hospitalet del Llobregat, Spain; 4Catalan Institution for Research and Advanced Studies, E08010 Barcelona, Spain Alternative splicing is regulated by multiple RNA-binding proteins and influences the expression of most eukaryotic genes. However, the role of this process in human disease, and particularly in cancer, is only starting to be unveiled. We system- atically analyzed mutation, copy number, and gene expression patterns of 1348 RNA-binding protein (RBP) genes in 11 solid tumor types, together with alternative splicing changes in these tumors and the enrichment of binding motifs in the alter- natively spliced sequences. Our comprehensive study reveals widespread alterations in the expression of RBP genes, as well as novel mutations and copy number variations in association with multiple alternative splicing changes in cancer drivers and oncogenic pathways. Remarkably, the altered splicing patterns in several tumor types recapitulate those of undifferen- tiated cells. These patterns are predicted to be mainly controlled by MBNL1 and involve multiple cancer drivers, including the mitotic gene NUMA1. We show that NUMA1 alternative splicing induces enhanced cell proliferation and centrosome am- plification in nontumorigenic mammary epithelial cells. -

Overlap of Quantitative Trait Loci for Early Growth Rate, and for Body
REPRODUCTIONRESEARCH Functional reconstruction of NANOS3 expression in the germ cell lineage by a novel transgenic reporter reveals distinct subcellular localizations of NANOS3 Masashi Yamaji1, Takashi Tanaka2, Mayo Shigeta1, Shinichiro Chuma2, Yumiko Saga3 and Mitinori Saitou1,4,5 1Laboratory for Mammalian Germ Cell Biology, RIKEN Center for Developmental Biology, 2-2-3 Minatojima- Minamimachi, Chuo-ku, Kobe 650-0047, Japan, 2Department of Development and Differentiation, Institute for Frontier Medical Sciences, Kyoto University, Shogoin, Sakyo-ku, Kyoto 606-8507, Japan, 3Department of Genetics and Division of Mammalian Development, National Institute of Genetics, SOKENDAI, 1111 Yata, Mishima, Shizuoka 411-8540, Japan, 4Department of Anatomy and Cell Biology, Graduate School of Medicine, Kyoto University, Yoshida- Konoe-cho, Sakyo-ku, Kyoto 606-8501, Japan and 5JST, CREST, Yoshida-Konoe-cho, Sakyo-ku, Kyoto 606-8501, Japan Correspondence should be addressed to M Saitou at Department of Anatomy and Cell Biology, Graduate School of Medicine, Kyoto University; Email: [email protected] Abstract Mutations of RNA-binding proteins such as NANOS3, TIAL1, and DND1 in mice have been known to result in the failure of survival and/or proliferation of primordial germ cells (PGCs) soon after their fate is specified (around embryonic day (E) 8.0), leading to the infertility of these animals. However, the mechanisms of actions of these RNA-binding proteins remain largely unresolved. As a foundation to explore the role of these RNA-binding proteins in germ cells, we established a novel transgenic reporter strain that expresses NANOS3 fused with EGFP under the control of Nanos3 regulatory elements. NANOS3–EGFP exhibited exclusive expression in PGCs as early as E7.25, and continued to be expressed in female germ cells until around E14.5 and in male germ cells throughout the fetal period with declining expression levels after E16.5. -

A Comprehensive Analysis of 3' End Sequencing Data Sets Reveals Novel
bioRxiv preprint doi: https://doi.org/10.1101/033001; this version posted November 26, 2015. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A comprehensive analysis of 3’ end sequencing data sets reveals novel polyadenylation signals and the repressive role of heterogenous ribonucleoprotein C on cleavage and polyadenylation 1 1 1 1 1 Andreas J. Gruber , Ralf Schmidt , Andreas R. Gruber , Georges Martin , Souvik Ghosh , 1,2 1 1,§ Manuel Belmadani , Walter Keller , Mihaela Zavolan 1 Computational and Systems Biology, Biozentrum, University of Basel, Klingelbergstrasse 5070, 4056 Basel, Switzerland 2 Current address: University of British Columbia, 177 Michael Smith Laboratories, 2185 East Mall Vancouver BC V6T 1Z4 § To whom correspondence should be addressed ([email protected]) Abstract Alternative polyadenylation (APA) is a general mechanism of transcript diversification in mammals, which has been recently linked to proliferative states and cancer. Different 3’ untranslated region (3’ UTR) isoforms interact with different RNA binding proteins (RBPs), which modify the stability, translation, and subcellular localization of the corresponding transcripts. Although the heterogeneity of premRNA 3’ end processing has been established with highthroughput approaches, the mechanisms that underlie systematic changes in 3’ UTR lengths remain to be characterized. Through a uniform analysis of a large number of 3’ end sequencing data sets we have uncovered 18 signals, 6 of which novel, whose positioning with respect to premRNA cleavage sites indicates a role in premRNA 3’ end processing in both mouse and human. -

Genetic Variant in 3' Untranslated Region of the Mouse Pycard Gene
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.26.437184; this version posted March 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 2 3 Title: 4 Genetic Variant in 3’ Untranslated Region of the Mouse Pycard Gene Regulates Inflammasome 5 Activity 6 Running Title: 7 3’UTR SNP in Pycard regulates inflammasome activity 8 Authors: 9 Brian Ritchey1*, Qimin Hai1*, Juying Han1, John Barnard2, Jonathan D. Smith1,3 10 1Department of Cardiovascular & Metabolic Sciences, Lerner Research Institute, Cleveland Clinic, 11 Cleveland, OH 44195 12 2Department of Quantitative Health Sciences, Lerner Research Institute, Cleveland Clinic, Cleveland, OH 13 44195 14 3Department of Molecular Medicine, Cleveland Clinic Lerner College of Medicine of Case Western 15 Reserve University, Cleveland, OH 44195 16 *, These authors contributed equally to this study. 17 Address correspondence to Jonathan D. Smith: email [email protected]; ORCID ID 0000-0002-0415-386X; 18 mailing address: Cleveland Clinic, Box NC-10, 9500 Euclid Avenue, Cleveland, OH 44195, USA. 19 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.26.437184; this version posted March 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 20 Abstract 21 Quantitative trait locus mapping for interleukin-1 release after inflammasome priming and activation 22 was performed on bone marrow-derived macrophages (BMDM) from an AKRxDBA/2 strain intercross. -
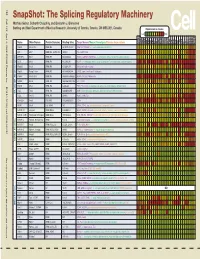
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

TIAL1 (NM 003252) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP760477 TIAL1 (NM_003252) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Purified recombinant protein of Human TIA1 cytotoxic granule-associated RNA binding protein-like 1 (TIAL1), transcript variant 1, full length, with N-terminal HIS tag, expressed in E.Coli, 50ug Species: Human Expression Host: E. coli Tag: N-His Predicted MW: 41.4 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25mM Tris, pH8.0, 150 mM NaCl, 10% glycerol, 1 % Sarkosyl. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_003243 Locus ID: 7073 UniProt ID: Q01085, Q49AS9 RefSeq Size: 1436 Cytogenetics: 10q26.11 RefSeq ORF: 1125 Synonyms: TCBP; TIAR Summary: The protein encoded by this gene is a member of a family of RNA-binding proteins, has three RNA recognition motifs (RRMs), and binds adenine and uridine-rich elements in mRNA and pre-mRNAs of a wide range of genes. It regulates various activities including translational control, splicing and apoptosis. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. The different isoforms have been show to function differently with respect to post-transcriptional silencing. [provided by RefSeq, Jul 2008] This product is to be used for laboratory only. -

The Devil Is in the Domain: Understanding Protein Recognition of Multiple RNA Targets
The devil is in the domain: understanding protein recognition of multiple RNA targets Glen R. Gronland1 and Andres Ramos1 1. Institute of Structural and Molecular Biology, University College London, London WC1E 6XA, UK Correspondence to Andres Ramos: [email protected] Abstract: RNA regulation provides a finely-tuned and highly-coordinated control of gene expression. Regulation is mediated by hundreds to thousands of multi-functional RNA-binding proteins which often interact with large sets of RNAs. In this brief review, we focus on recent work that highlights how the proteins use multiple RNA-binding domains to interact selectively with the different RNA targets. De-convoluting the molecular complexity of the RNA regulatory network is essential to understanding cell differentiation and function, and requires accurate models for protein-RNA recognition and protein target selectivity. We discuss that the structural and molecular understanding of the key determinant of recognition, together with the availability of methods to examine protein-RNA interactions at the transcriptome level, may provide an avenue to establish these models. Abbreviations list: CSD, Cold shock domain dsRBD, Double-stranded RNA-binding domain KH, hnRNP K-homology mRNA, Messenger RNA ncRNA, Non-coding RNA RBD, RNA-binding domain RBP, RNA-binding protein RRM, RNA-recognition motif ZnF, Zinc finger Introduction The combined regulation of the various steps in the metabolism and transport of messenger RNAs (mRNAs) and non-coding RNAs (ncRNAs) multiplies genomic potential, and allows cellular differentiation and the development of complex organisms. In the cell, functional RNAs are associated with a fluctuating assortment of multi-domain RNA-binding proteins (RBPs), forming integrated complexes, whose composition directs the fate of the transcript1,2. -

TIAL1 Rabbit Pab
Leader in Biomolecular Solutions for Life Science TIAL1 Rabbit pAb Catalog No.: A6075 Basic Information Background Catalog No. The protein encoded by this gene is a member of a family of RNA-binding proteins, has A6075 three RNA recognition motifs (RRMs), and binds adenine and uridine-rich elements in mRNA and pre-mRNAs of a wide range of genes. It regulates various activities including Observed MW translational control, splicing and apoptosis. Alternate transcriptional splice variants, 47kDa encoding different isoforms, have been characterized. The different isoforms have been show to function differently with respect to post-transcriptional silencing. Calculated MW 41kDa/43kDa Category Primary antibody Applications WB, IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 7073 Q01085 IHC 1:100 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 266-375 of human TIAL1 (NP_003243.1). Synonyms TIAL1;TCBP;TIAR Contact Product Information www.abclonal.com Source Isotype Purification Rabbit IgG Affinity purification Storage Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide,50% glycerol,pH7.3. Validation Data Western blot analysis of extracts of various cell lines, using TIAL1 antibody (A6075) at 1:3000 dilution. Secondary antibody: HRP Goat Anti-Rabbit IgG (H+L) (AS014) at 1:10000 dilution. Lysates/proteins: 25ug per lane. Blocking buffer: 3% nonfat dry milk in TBST. Detection: ECL Basic Kit (RM00020). Exposure time: 90s. Immunohistochemistry of paraffin- Immunohistochemistry of paraffin- Immunohistochemistry of paraffin- embedded rat lung using TIAL1 antibody embedded human appendix using TIAL1 embedded mouse spleen using TIAL1 (A6075) at dilution of 1:100 (40x lens). -

Quantitative Trait Loci Mapping of Macrophage Atherogenic Phenotypes
QUANTITATIVE TRAIT LOCI MAPPING OF MACROPHAGE ATHEROGENIC PHENOTYPES BRIAN RITCHEY Bachelor of Science Biochemistry John Carroll University May 2009 submitted in partial fulfillment of requirements for the degree DOCTOR OF PHILOSOPHY IN CLINICAL AND BIOANALYTICAL CHEMISTRY at the CLEVELAND STATE UNIVERSITY December 2017 We hereby approve this thesis/dissertation for Brian Ritchey Candidate for the Doctor of Philosophy in Clinical-Bioanalytical Chemistry degree for the Department of Chemistry and the CLEVELAND STATE UNIVERSITY College of Graduate Studies by ______________________________ Date: _________ Dissertation Chairperson, Johnathan D. Smith, PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, David J. Anderson, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Baochuan Guo, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Stanley L. Hazen, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Renliang Zhang, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Aimin Zhou, PhD Department of Chemistry, Cleveland State University Date of Defense: October 23, 2017 DEDICATION I dedicate this work to my entire family. In particular, my brother Greg Ritchey, and most especially my father Dr. Michael Ritchey, without whose support none of this work would be possible. I am forever grateful to you for your devotion to me and our family. You are an eternal inspiration that will fuel me for the remainder of my life. I am extraordinarily lucky to have grown up in the family I did, which I will never forget. -

Inhibiting PARP1 Splicing Along with Inducing DNA Damage As Potential Breast Cancer Therapy
Reem Alsayed 3/26/21 03-545, S21 Professor Ihab Younis Inhibiting PARP1 Splicing along with Inducing DNA Damage as Potential Breast Cancer Therapy Student: Reem Alsayed Spring 2021 Professor: Ihab Younis 1 Reem Alsayed 3/26/21 03-545, S21 Professor Ihab Younis Abstract: Triple negative breast cancer is a deadly cancer and once it has metastasized it is deemed incurable. The need for an effective therapy is rising, and recent therapies include targeting the DNA damage response pathway. PARP1 is one of the first responders to DNA damage, and has been targeted for inhibition along with the stimulation of DNA damage as a treatment for breast cancer. However, such treatments lack in specificity, and only target one or two domains of the PARP1 protein, whereas PARP1 has other functions pertaining to multiple cancer hallmarks such as promoting angiogenesis, metastasis, inflammation, life cycle regulation, and regulation of tumorigenic genes. In this project, we hypothesize that by inhibiting the PARP1 protein production, we will be able to effectively inhibit all cancer hallmarks that are facilitated by PARP1, and we achieve this by inhibiting the splicing of PARP1. Splicing is the removal of intervening sequences (introns) in the pre-mRNA and the joining of the expressed sequences (exons). For PARP1, we blocked intron 22 splicing by introducing an Antisense Morpholino Oligonucleotide (AMO) that blocks the binding of the spliceosome. The results obtained demonstrate that 50uM PARP1 AMO inhibits PARP1 splicing >88%, as well as inhibits protein production. Additionally, the combination of PARP1 AMO and Doxorubicin lead to a loss in cell proliferation. -

The Tip of the Iceberg: RNA-Binding Proteins with Prion-Like Domains in Neurodegenerative Disease
BRES-42024; No. of pages: 20; 4C: 7, 8, 10, 12, 13 BRAIN RESEARCH XX (2012) XXX– XXX Available online at www.sciencedirect.com www.elsevier.com/locate/brainres Review The tip of the iceberg: RNA-binding proteins with prion-like domains in neurodegenerative disease Oliver D. Kinga,⁎, Aaron D. Gitlerb,⁎⁎, James Shorterc,⁎⁎⁎ aBoston Biomedical Research Institute, 64 Grove St., Watertown, MA 02472, USA bDepartment of Genetics, Stanford University School of Medicine, 300 Pasteur Drive, M322 Alway Building, Stanford, CA 94305-5120, USA cDepartment of Biochemistry and Biophysics, University of Pennsylvania School of Medicine, 805b Stellar-Chance Laboratories, 422 Curie Boulevard, Philadelphia, PA 19104, USA ARTICLE INFO ABSTRACT Article history: Prions are self-templating protein conformers that are naturally transmitted between individ- Accepted 7 January 2012 uals and promote phenotypic change. In yeast, prion-encoded phenotypes can be beneficial, neutral or deleterious depending upon genetic background and environmental conditions. A dis- Keywords: tinctive and portable ‘prion domain’ enriched in asparagine, glutamine, tyrosine and glycine res- Prion idues unifies the majority of yeast prion proteins. Deletion of this domain precludes RNA-binding protein prionogenesis and appending this domain to reporter proteins can confer prionogenicity. An al- TDP-43 gorithm designed to detect prion domains has successfully identified 19 domains that can confer FUS prion behavior. Scouring the human genome with this algorithm enriches a select group of RNA- TAF15 binding proteins harboring a canonical RNA recognition motif (RRM) and a putative prion do- EWSR1 main. Indeed, of 210 human RRM-bearing proteins, 29 have a putative prion domain, and 12 of Amyotrophic lateral sclerosis these are in the top 60 prion candidates in the entire genome. -

The Neurodegenerative Diseases ALS and SMA Are Linked at The
Nucleic Acids Research, 2019 1 doi: 10.1093/nar/gky1093 The neurodegenerative diseases ALS and SMA are linked at the molecular level via the ASC-1 complex Downloaded from https://academic.oup.com/nar/advance-article-abstract/doi/10.1093/nar/gky1093/5162471 by [email protected] on 06 November 2018 Binkai Chi, Jeremy D. O’Connell, Alexander D. Iocolano, Jordan A. Coady, Yong Yu, Jaya Gangopadhyay, Steven P. Gygi and Robin Reed* Department of Cell Biology, Harvard Medical School, 240 Longwood Ave. Boston MA 02115, USA Received July 17, 2018; Revised October 16, 2018; Editorial Decision October 18, 2018; Accepted October 19, 2018 ABSTRACT Fused in Sarcoma (FUS) and TAR DNA Binding Protein (TARDBP) (9–13). FUS is one of the three members of Understanding the molecular pathways disrupted in the structurally related FET (FUS, EWSR1 and TAF15) motor neuron diseases is urgently needed. Here, we family of RNA/DNA binding proteins (14). In addition to employed CRISPR knockout (KO) to investigate the the RNA/DNA binding domains, the FET proteins also functions of four ALS-causative RNA/DNA binding contain low-complexity domains, and these domains are proteins (FUS, EWSR1, TAF15 and MATR3) within the thought to be involved in ALS pathogenesis (5,15). In light RNAP II/U1 snRNP machinery. We found that each of of the discovery that mutations in FUS are ALS-causative, these structurally related proteins has distinct roles several groups carried out studies to determine whether the with FUS KO resulting in loss of U1 snRNP and the other two members of the FET family, TATA-Box Bind- SMN complex, EWSR1 KO causing dissociation of ing Protein Associated Factor 15 (TAF15) and EWS RNA the tRNA ligase complex, and TAF15 KO resulting in Binding Protein 1 (EWSR1), have a role in ALS.