Supplementary Table 1. DNA Array1 Data of the Leukemic Clone from Patient C.2.2
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Np63 Activates the Fanconi Anemia DNA Repair Pathway and Limits the Efficacy of Cisplatin Treatment in Squamous Cell Carcinoma Anne Catherine Bretz1, Miriam P
3204–3218 Nucleic Acids Research, 2016, Vol. 44, No. 7 Published online 26 January 2016 doi: 10.1093/nar/gkw036 Np63 activates the Fanconi anemia DNA repair pathway and limits the efficacy of cisplatin treatment in squamous cell carcinoma Anne Catherine Bretz1, Miriam P. Gittler1,Joel¨ P. Charles1, Niklas Gremke1, Ines Eckhardt1, Marco Mernberger1, Robert Mandic2,Jurgen¨ Thomale3, Andrea Nist4, Michael Wanzel1,5 and Thorsten Stiewe1,4,5,* 1Institute of Molecular Oncology, Philipps-University Marburg, 35043 Marburg, Germany, 2Department of Otorhinolaryngology, Head & Neck Surgery, University Hospital Giessen and Marburg, Philipps-University, 35033 Marburg, Germany, 3Institute of Cell Biology, University Duisburg-Essen, 45122 Essen, Germany, 4Genomics Core Facility, Philipps-University Marburg, 35043 Marburg, Germany and 5Universities of Giessen and Marburg Lung Center, member of the German Center for Lung Research (DZL), 35043 Marburg, Germany Received September 03, 2015; Revised December 18, 2015; Accepted January 12, 2016 ABSTRACT INTRODUCTION TP63, a member of the p53 gene family gene, encodes Squamous cell carcinoma (SCC) is a common and lethal the Np63 protein and is one of the most frequently human cancer, and relatively little progress has been made amplified genes in squamous cell carcinomas (SCC) in improving outcomes for SCC due to a poor understand- of the head and neck (HNSCC) and lungs (LUSC). ing of its underlying molecular pathogenesis (1,2). While Using an epiallelic series of siRNAs with intrinsically SCCs typically lack somatic oncogene-activating mutations, they exhibit frequent amplification of the p53 gene fam- different knockdown abilities, we show that the com- ily member TP63 (1–6). Recent tumor genome sequencing plete loss of Np63 strongly impaired cell prolifera- studies have revealed TP63 amplification in 20–25% of head tion, whereas partial Np63 depletion rendered cells and neck SCC (HNSCC) and 25–50% of lung SCC (LUSC) hypersensitive to cisplatin accompanied by an ac- (4,6). -

Viewed Under 23 (B) Or 203 (C) fi M M Male Cko Mice, and Largely Unaffected Magni Cation; Scale Bars, 500 M (B) and 50 M (C)
BRIEF COMMUNICATION www.jasn.org Renal Fanconi Syndrome and Hypophosphatemic Rickets in the Absence of Xenotropic and Polytropic Retroviral Receptor in the Nephron Camille Ansermet,* Matthias B. Moor,* Gabriel Centeno,* Muriel Auberson,* † † ‡ Dorothy Zhang Hu, Roland Baron, Svetlana Nikolaeva,* Barbara Haenzi,* | Natalya Katanaeva,* Ivan Gautschi,* Vladimir Katanaev,*§ Samuel Rotman, Robert Koesters,¶ †† Laurent Schild,* Sylvain Pradervand,** Olivier Bonny,* and Dmitri Firsov* BRIEF COMMUNICATION *Department of Pharmacology and Toxicology and **Genomic Technologies Facility, University of Lausanne, Lausanne, Switzerland; †Department of Oral Medicine, Infection, and Immunity, Harvard School of Dental Medicine, Boston, Massachusetts; ‡Institute of Evolutionary Physiology and Biochemistry, St. Petersburg, Russia; §School of Biomedicine, Far Eastern Federal University, Vladivostok, Russia; |Services of Pathology and ††Nephrology, Department of Medicine, University Hospital of Lausanne, Lausanne, Switzerland; and ¶Université Pierre et Marie Curie, Paris, France ABSTRACT Tight control of extracellular and intracellular inorganic phosphate (Pi) levels is crit- leaves.4 Most recently, Legati et al. have ical to most biochemical and physiologic processes. Urinary Pi is freely filtered at the shown an association between genetic kidney glomerulus and is reabsorbed in the renal tubule by the action of the apical polymorphisms in Xpr1 and primary fa- sodium-dependent phosphate transporters, NaPi-IIa/NaPi-IIc/Pit2. However, the milial brain calcification disorder.5 How- molecular identity of the protein(s) participating in the basolateral Pi efflux remains ever, the role of XPR1 in the maintenance unknown. Evidence has suggested that xenotropic and polytropic retroviral recep- of Pi homeostasis remains unknown. Here, tor 1 (XPR1) might be involved in this process. Here, we show that conditional in- we addressed this issue in mice deficient for activation of Xpr1 in the renal tubule in mice resulted in impaired renal Pi Xpr1 in the nephron. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

RHOBTB1 CRISPR/Cas9 KO Plasmid (H): Sc-418424
SANTA CRUZ BIOTECHNOLOGY, INC. RHOBTB1 CRISPR/Cas9 KO Plasmid (h): sc-418424 BACKGROUND APPLICATIONS The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and RHOBTB1 CRISPR/Cas9 KO Plasmid (h) is recommended for the disruption of CRISPR-associated protein (Cas9) system is an adaptive immune response gene expression in human cells. defense mechanism used by archea and bacteria for the degradation of foreign genetic material (4,6). This mechanism can be repurposed for other 20 nt non-coding RNA sequence: guides Cas9 functions, including genomic engineering for mammalian systems, such as to a specific target location in the genomic DNA gene knockout (KO) (1,2,3,5). CRISPR/Cas9 KO Plasmid products enable the U6 promoter: drives gRNA scaffold: helps Cas9 identification and cleavage of specific genes by utilizing guide RNA (gRNA) expression of gRNA bind to target DNA sequences derived from the Genome-scale CRISPR Knock-Out (GeCKO) v2 library developed in the Zhang Laboratory at the Broad Institute (3,5). Termination signal Green Fluorescent Protein: to visually REFERENCES verify transfection CRISPR/Cas9 Knockout Plasmid CBh (chicken β-Actin 1. Cong, L., et al. 2013. Multiplex genome engineering using CRISPR/Cas hybrid) promoter: drives expression of Cas9 systems. Science 339: 819-823. 2A peptide: allows production of both Cas9 and GFP from the 2. Mali, P., et al. 2013. RNA-guided human genome engineering via Cas9. same CBh promoter Science 339: 823-826. Nuclear localization signal 3. Ran, F.A., et al. 2013. Genome engineering using the CRISPR-Cas9 system. Nuclear localization signal SpCas9 ribonuclease Nat. Protoc. 8: 2281-2308. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

High Throughput Strategies Aimed at Closing the GAP in Our Knowledge of Rho Gtpase Signaling
cells Review High Throughput strategies Aimed at Closing the GAP in Our Knowledge of Rho GTPase Signaling Manel Dahmene 1, Laura Quirion 2 and Mélanie Laurin 1,3,* 1 Oncology Division, CHU de Québec–Université Laval Research Center, Québec, QC G1V 4G2, Canada; [email protected] 2 Montréal Clinical Research Institute (IRCM), Montréal, QC H2W 1R7, Canada; [email protected] 3 Université Laval Cancer Research Center, Québec, QC G1R 3S3, Canada * Correspondence: [email protected] Received: 21 May 2020; Accepted: 7 June 2020; Published: 9 June 2020 Abstract: Since their discovery, Rho GTPases have emerged as key regulators of cytoskeletal dynamics. In humans, there are 20 Rho GTPases and more than 150 regulators that belong to the RhoGEF, RhoGAP, and RhoGDI families. Throughout development, Rho GTPases choregraph a plethora of cellular processes essential for cellular migration, cell–cell junctions, and cell polarity assembly. Rho GTPases are also significant mediators of cancer cell invasion. Nevertheless, to date only a few molecules from these intricate signaling networks have been studied in depth, which has prevented appreciation for the full scope of Rho GTPases’ biological functions. Given the large complexity involved, system level studies are required to fully grasp the extent of their biological roles and regulation. Recently, several groups have tackled this challenge by using proteomic approaches to map the full repertoire of Rho GTPases and Rho regulators protein interactions. These studies have provided in-depth understanding of Rho regulators specificity and have contributed to expand Rho GTPases’ effector portfolio. Additionally, new roles for understudied family members were unraveled using high throughput screening strategies using cell culture models and mouse embryos. -

Open Questions: What About the 'Other' Rho Gtpases?
Ridley BMC Biology (2016) 14:64 DOI 10.1186/s12915-016-0289-7 COMMENT Open Access Open questions: what about the ‘other’ Rho GTPases? Anne J. Ridley Abstract know if they interact with and/or regulate the activity of other family members. Indeed, by studying only RhoA, Rho GTPases have many and diverse roles in cell Rac1 and Cdc42, we are likely to be missing the real physiology, and some family members are very well functions of many GEFs and GAPs because their targets studied, including RhoA, Rac1 and Cdc42. But many in cells are among the other Rho GTPases. are relatively neglected, and fundamental questions Interestingly, four family members—Rnd1, Rnd2, Rnd3 about their mechanisms and functions remain open. and RhoH—are ‘atypical’, in that they are known to be constitutively GTP-bound and do not hydrolyse GTP: much less is known about how these family members Rho GTPases are household names for anyone who are regulated. RhoU and RhoV have high intrinsic GDP/ works on eukaryotic cell migration and their functions GTP exchange rates, so are unlikely to need GEFs for in cell migration, cell division and cell polarity are de- activation but could still be turned off by GAPs [1, 4]. scribed in most textbooks on cell biology. Yet most of For two other members, RhoBTB1 and RhoBTB2, the what we know about Rho GTPases comes from studying Rho domains are quite divergent in sequence from other a small subset of the many different family members, family members and they are unlikely to be regulated by namely RhoA, Rac1 and Cdc42. -

Rho Guanine Nucleotide Exchange Factors: Regulators of Rho Gtpase Activity in Development and Disease
Oncogene (2014) 33, 4021–4035 & 2014 Macmillan Publishers Limited All rights reserved 0950-9232/14 www.nature.com/onc REVIEW Rho guanine nucleotide exchange factors: regulators of Rho GTPase activity in development and disease DR Cook1, KL Rossman2,3 and CJ Der1,2,3 The aberrant activity of Ras homologous (Rho) family small GTPases (20 human members) has been implicated in cancer and other human diseases. However, in contrast to the direct mutational activation of Ras found in cancer and developmental disorders, Rho GTPases are activated most commonly in disease by indirect mechanisms. One prevalent mechanism involves aberrant Rho activation via the deregulated expression and/or activity of Rho family guanine nucleotide exchange factors (RhoGEFs). RhoGEFs promote formation of the active GTP-bound state of Rho GTPases. The largest family of RhoGEFs is comprised of the Dbl family RhoGEFs with 70 human members. The multitude of RhoGEFs that activate a single Rho GTPase reflects the very specific role of each RhoGEF in controlling distinct signaling mechanisms involved in Rho activation. In this review, we summarize the role of Dbl RhoGEFs in development and disease, with a focus on Ect2 (epithelial cell transforming squence 2), Tiam1 (T-cell lymphoma invasion and metastasis 1), Vav and P-Rex1/2 (PtdIns(3,4,5)P3 (phosphatidylinositol (3,4,5)-triphosphate)-dependent Rac exchanger). Oncogene (2014) 33, 4021–4035; doi:10.1038/onc.2013.362; published online 16 September 2013 Keywords: Rac1; RhoA; Cdc42; guanine nucleotide exchange factors; cancer; -

Post-Transcriptionally Impaired De Novo Mutations Contribute to The
bioRxiv preprint doi: https://doi.org/10.1101/175844; this version posted November 26, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Post-transcriptionally impaired de novo mutations 2 contribute to the genetic etiology of four neuropsychiatric 3 disorders 4 5 Fengbiao Mao1,2¶, Lu Wang3¶, Xiaolu Zhao2, Zhongshan Li4, Luoyuan Xiao5, 6 Rajesh C. Rao2, Jinchen Li4, Huajing Teng1*, Xin He6*, and Zhong Sheng Sun1,4* 7 8 1 Beijing Institutes of Life Science, Chinese Academy of Sciences, Beijing 100101, 9 China. 10 2 Department of Pathology, University of Michigan, Ann Arbor, MI 48109, USA. 11 3 Institute of Life Science, Southeast University, Nanjing 210096, China. 12 4 Institute of Genomic Medicine, Wenzhou Medical University, Wenzhou 325027, 13 China 14 5 Department of Computer Science and Technology, Tsinghua University, Beijing 15 100084, China. 16 6 Department of Human Genetics, University of Chicago, Chicago, IL, USA. 17 18 ¶These authors contributed equally to this work 19 * Corresponding authors 20 E-mail: 21 [email protected] (Z.S.S.) 22 [email protected] (X.H.) 23 [email protected] (H.T.) 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/175844; this version posted November 26, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Rho Gtpases of the Rhobtb Subfamily and Tumorigenesis1
Acta Pharmacol Sin 2008 Mar; 29 (3): 285–295 Invited review Rho GTPases of the RhoBTB subfamily and tumorigenesis1 Jessica BERTHOLD2, Kristína SCHENKOVÁ2, Francisco RIVERO2,3,4 2Centers for Biochemistry and Molecular Medicine, University of Cologne, Cologne, Germany; 3The Hull York Medical School, University of Hull, Hull HU6 7RX, UK Key words Abstract Rho guanosine triphosphatase; RhoBTB; RhoBTB proteins constitute a subfamily of atypical members within the Rho fa- DBC2; cullin; neoplasm mily of small guanosine triphosphatases (GTPases). Their most salient feature 1This work was supported by grants from the is their domain architecture: a GTPase domain (in most cases, non-functional) Center for Molecular Medicine Cologne, the is followed by a proline-rich region, a tandem of 2 broad-complex, tramtrack, Deutsche Forschungsgemeinschaft, and the bric à brac (BTB) domains, and a conserved C-terminal region. In humans, Köln Fortune Program of the Medical Faculty, University of Cologne. the RhoBTB subfamily consists of 3 isoforms: RhoBTB1, RhoBTB2, and RhoBTB3. Orthologs are present in several other eukaryotes, such as Drosophi- 4 Correspondence to Dr Francisco RIVERO. la and Dictyostelium, but have been lost in plants and fungi. Interest in RhoBTB Phn 49-221-478-6987. Fax 49-221-478-6979. arose when RHOBTB2 was identified as the gene homozygously deleted in E-mail [email protected] breast cancer samples and was proposed as a candidate tumor suppressor gene, a property that has been extended to RHOBTB1. The functions of RhoBTB pro- Received 2007-11-17 Accepted 2007-12-16 teins have not been defined yet, but may be related to the roles of BTB domains in the recruitment of cullin3, a component of a family of ubiquitin ligases. -

It Was Hypothesized That Hybrid Synthetic/Adenoviral Nanoparticles
Electronic Supplementary Material (ESI) for Nanoscale. This journal is © The Royal Society of Chemistry 2020 SUPPORTING INFORMATION Lana Papafilippou,a Andrew Claxton,b Paul Dark,b,c Kostas Kostarelos*a,d and Marilena Hadjidemetriou*a aNanomedicine Lab, Faculty of Biology, Medicine & Health, AV Hill Building, The University of Manchester, Manchester, M13 9PT, UK bCentre for Acute Care Trauma, Manchester Academic Health Science Centre, Health Innovation Manchester, Division of Critical Care, Salford Royal NHS Foundation Trust, Greater Manchester, UK cDivision of Infection, Immunity and Respiratory Medicine, Faculty of Biology, Medicine & Health, AV Hill Building, The University of Manchester, Manchester, M13 9PT, UK. d Catalan Institute of Nanoscience and Nanotechnology (ICN2), Campus UAB, Bellaterra, 08193 Barcelona, Spain. _______________________________________ * Correspondence should be addressed to: [email protected],uk; [email protected] 1 Supporting Figure 1 Figure S1: Physicochemical characterization of corona-coated Amphotericin B-intercalated liposomes (AmBisome®). Mean hydrodynamic diameter (nm) and ζ-potential (mV) distributions are depicted for corona-coated liposomal formulation AmBisome® recovered post-incubation with human plasma from 12 healthy volunteers, 7 SIRS patients and 12 sepsis patients. 2 Supporting Figure 2 Figure S2: Proteomic analysis of corona profiles. (A) Heatmap of normalized abundance values of all corona proteins identified in healthy controls, SIRS patients and sepsis patients, as identified by LC-MS/MS (Progenesis QI). Protein columns are sorted according to the abundance values (from highest to lowest) of the first sample. The list of proteins shown in the heatmap, their respective accession numbers and their mean normalized abundance values are shown in Table S5; (B) Volcano plot represents the potential protein biomarkers differentially abundant between healthy donors and sepsis patients (n=135) identified in corona samples. -
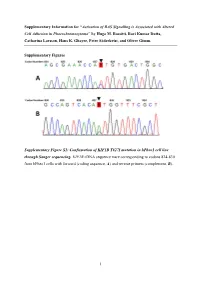
Activation of RAS Signalling Is Associated with Altered Cell Adhesion in Phaeochromocytoma” by Hugo M
Supplementary Information for “Activation of RAS Signalling is Associated with Altered Cell Adhesion in Phaeochromocytoma” by Hugo M. Rossitti, Ravi Kumar Dutta, Catharina Larsson, Hans K. Ghayee, Peter Söderkvist, and Oliver Gimm. Supplementary Figures Supplementary Figure S1: Confirmation of KIF1B T827I mutation in hPheo1 cell line through Sanger sequencing. KIF1B cDNA sequence trace corresponding to codons 824-830 from hPheo1 cells with forward (coding sequence, A) and reverse primers (complement, B). 1 Supplementary Figure S2: Confirmation of NRAS Q61K mutation in hPheo1 cell line through Sanger sequencing. NRAS cDNA sequence trace corresponding to codons 56-67 from hPheo1 cells with forward (coding sequence, A) and reverse primers (complement, B). 2 Supplementary Figure S3: CCND1 gene expression and hPheo1 proliferation. A: Expression of CCND1 mRNA assessed by RT-qPCR and presented as fold change (2-ΔΔCT, mean ± standard error of the mean). B: Cell counts at 1, 2, and 3 days after plating (corresponding to 72, 96 and 120 hours posttransfection, respectively) of control- or siNRAS#1-transfected hPheo1 cells expressed as fold change of the number of cells plated at day 0 (48 hours posttransfection; mean ± standard deviation). All results are from three independent siRNA experiments. 3 Supplementary Tables Supplementary Table S1: List of transcript cluster IDs significantly upregulated in hPheo1 by siNRAS treatment (comparison: siNRAS versus control-transfected hPheo1; ANOVA p < 0.05, FDR < 0.25, fold change < -1.5 or > 1.5). Transcript