Site-Specific His/Asp Phosphoproteomic Analysis Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Cross-Talk Between Methylation and Phosphorylation in Lymphoid-Specific Helicase Drives Cancer Stem-Like Properties
Signal Transduction and Targeted Therapy www.nature.com/sigtrans ARTICLE OPEN The cross-talk between methylation and phosphorylation in lymphoid-specific helicase drives cancer stem-like properties Na Liu1,2,3, Rui Yang1,2, Ying Shi1,2, Ling Chen1,2, Yating Liu1,2, Zuli Wang1,2, Shouping Liu1,2, Lianlian Ouyang4, Haiyan Wang1,2, Weiwei Lai1,2, Chao Mao1,2, Min Wang1,2, Yan Cheng5, Shuang Liu4, Xiang Wang6, Hu Zhou7, Ya Cao1,2, Desheng Xiao1 and Yongguang Tao1,2,6 Posttranslational modifications (PTMs) of proteins, including chromatin modifiers, play crucial roles in the dynamic alteration of various protein properties and functions including stem-cell properties. However, the roles of Lymphoid-specific helicase (LSH), a DNA methylation modifier, in modulating stem-like properties in cancer are still not clearly clarified. Therefore, exploring PTMs modulation of LSH activity will be of great significance to further understand the function and activity of LSH. Here, we demonstrate that LSH is capable to undergo PTMs, including methylation and phosphorylation. The arginine methyltransferase PRMT5 can methylate LSH at R309 residue, meanwhile, LSH could as well be phosphorylated by MAPK1 kinase at S503 residue. We further show that the accumulation of phosphorylation of LSH at S503 site exhibits downregulation of LSH methylation at R309 residue, which eventually promoting stem-like properties in lung cancer. Whereas, phosphorylation-deficient LSH S503A mutant promotes the accumulation of LSH methylation at R309 residue and attenuates stem-like properties, indicating the critical roles of LSH PTMs in modulating stem-like properties. Thus, our study highlights the importance of the crosstalk between LSH PTMs in determining its activity and function in lung cancer stem-cell maintenance. -

Phosphorylation of Chicken Protein Tyrosine Phosphatase 1 by Casein Kinase II in Vitro
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 29, No 4, 229-233, December 1997 Phosphorylation of chicken protein tyrosine phosphatase 1 by casein kinase II in vitro Eun Joo Jung,1 Kee Ryeon Kang1 and Introduction Yoon-Se Kang1,2 The phosphorylation of protein tyrosine residues is an early event in signal transduction initiated by binding of 1 Department of Biochemistry and Gyeongsang Institute of Cancer growth factors and hormones to their cognate receptors Research, College of Medicine, Gyeongsang National University, and it leads to regulation of cellular activities which include Chinju 660-280, Korea proliferation, differentiation, and also malignant transfor- 2 Corresponding author mation of cells (Hunter, 1989; Ullirich and Schlessinger, Accepted 17 November 1997 1990; Cantley et al., 1991). Under normal conditions, the level of tyrosine phosphorylation within a cell is determined by a balance between the actions of protein tyrosine Abbreviations: CPTP, chicken protein tyrosine phosphatase; HPTP1B, human placenta kinases (PTKs) and protein tyrosine phosphatases (PTPs) protein tyrosine phosphatase 1B; CKII, casein kinase II; MAP kinase, mitogen-activated (Hunter, 1989; Fischer et al., 1991; Trowbridge, 1991). protein kinase; GST, glutathione S-transferase; pNPP, p-nitrophenyl phosphate; EGF, PTPs do not simply reverse the action of tyrosine kinases, epidermal growth factor but rather, PTP itself may play a central role in cellular regulation. PTPs are generally classified as transmem- brane (receptor-type) and nontransmembrane (nonrecep- tor-type) enzymes based on the presence or absence of extracellular and transmembrane portions of their predicted sequence (Fischer et al., 1991). Because the activity of Abstract tyrosine kinase can be controlled by phosphorylation, it has been postulated that PTP activity may be regulated The phosphorylation and dephosphorylation of by phosphorylation as well. -

Citric Acid Cycle
CHEM464 / Medh, J.D. The Citric Acid Cycle Citric Acid Cycle: Central Role in Catabolism • Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylCoA • In aerobic organisms, citric acid cycle makes up the final stage of catabolism when acetyl CoA is completely oxidized to CO2. • Also called Krebs cycle or tricarboxylic acid (TCA) cycle. • It is a central integrative pathway that harvests chemical energy from biological fuel in the form of electrons in NADH and FADH2 (oxidation is loss of electrons). • NADH and FADH2 transfer electrons via the electron transport chain to final electron acceptor, O2, to form H2O. Entry of Pyruvate into the TCA cycle • Pyruvate is formed in the cytosol as a product of glycolysis • For entry into the TCA cycle, it has to be converted to Acetyl CoA. • Oxidation of pyruvate to acetyl CoA is catalyzed by the pyruvate dehydrogenase complex in the mitochondria • Mitochondria consist of inner and outer membranes and the matrix • Enzymes of the PDH complex and the TCA cycle (except succinate dehydrogenase) are in the matrix • Pyruvate translocase is an antiporter present in the inner mitochondrial membrane that allows entry of a molecule of pyruvate in exchange for a hydroxide ion. 1 CHEM464 / Medh, J.D. The Citric Acid Cycle The Pyruvate Dehydrogenase (PDH) complex • The PDH complex consists of 3 enzymes. They are: pyruvate dehydrogenase (E1), Dihydrolipoyl transacetylase (E2) and dihydrolipoyl dehydrogenase (E3). • It has 5 cofactors: CoASH, NAD+, lipoamide, TPP and FAD. CoASH and NAD+ participate stoichiometrically in the reaction, the other 3 cofactors have catalytic functions. -
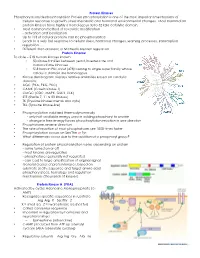
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Structural Basis of O-Glcnac Recognition by Mammalian 14-3-3 Proteins
Structural basis of O-GlcNAc recognition by mammalian 14-3-3 proteins Clifford A. Tolemana,1, Maria A. Schumachera,1, Seok-Ho Yub, Wenjie Zenga, Nathan J. Coxa, Timothy J. Smitha, Erik J. Soderblomc, Amberlyn M. Wandsb, Jennifer J. Kohlerb, and Michael Boycea,2 aDepartment of Biochemistry, Duke University School of Medicine, Durham, NC 27710; bDepartment of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390; and cDuke Proteomics and Metabolomics Core Facility, Center for Genomic and Computational Biology, Duke University, Durham, NC 27710 Edited by Carolyn R. Bertozzi, Stanford University, Stanford, CA, and approved April 23, 2018 (received for review December 24, 2017) O-GlcNAc is an intracellular posttranslational modification that gov- Results erns myriad cell biological processes and is dysregulated in human We developed a biochemical approach to test the hypothesis that diseases. Despite this broad pathophysiological significance, the O-GlcNAc is specifically recognized by mammalian reader pro- biochemical effects of most O-GlcNAcylation events remain unchar- teins. First, we derived a consensus O-GlcNAcylated peptide acterized. One prevalent hypothesis is that O-GlcNAc moieties may sequence by aligning 802 mapped Ser-O-GlcNAc sites (34–36) be recognized by “reader” proteins to effect downstream signaling. (Fig. 1A)(www.phosphosite.org). We noted that a Pro-Val-Ser However, no general O-GlcNAc readers have been identified, leav- tripeptide observed previously in smaller datasets (37, 38) also ing a considerable gap in the field. To elucidate O-GlcNAc signaling emerged in our sequence, suggesting that this motif may be mechanisms, we devised a biochemical screen for candidate O-GlcNAc important for O-GlcNAc modification and/or recognition. -

Glycolysis Citric Acid Cycle Oxidative Phosphorylation Calvin Cycle Light
Stage 3: RuBP regeneration Glycolysis Ribulose 5- Light-Dependent Reaction (Cytosol) phosphate 3 ATP + C6H12O6 + 2 NAD + 2 ADP + 2 Pi 3 ADP + 3 Pi + + 1 GA3P 6 NADP + H Pi NADPH + ADP + Pi ATP 2 C3H4O3 + 2 NADH + 2 H + 2 ATP + 2 H2O 3 CO2 Stage 1: ATP investment ½ glucose + + Glucose 2 H2O 4H + O2 2H Ferredoxin ATP Glyceraldehyde 3- Ribulose 1,5- Light Light Fx iron-sulfur Sakai-Kawada, F Hexokinase phosphate bisphosphate - 4e + center 2016 ADP Calvin Cycle 2H Stroma Mn-Ca cluster + 6 NADP + Light-Independent Reaction Phylloquinone Glucose 6-phosphate + 6 H + 6 Pi Thylakoid Tyr (Stroma) z Fe-S Cyt f Stage 1: carbon membrane Phosphoglucose 6 NADPH P680 P680* PQH fixation 2 Plastocyanin P700 P700* D-(+)-Glucose isomerase Cyt b6 1,3- Pheophytin PQA PQB Fructose 6-phosphate Bisphosphoglycerate ATP Lumen Phosphofructokinase-1 3-Phosphoglycerate ADP Photosystem II P680 2H+ Photosystem I P700 Stage 2: 3-PGA Photosynthesis Fructose 1,6-bisphosphate reduction 2H+ 6 ADP 6 ATP 6 CO2 + 6 H2O C6H12O6 + 6 O2 H+ + 6 Pi Cytochrome b6f Aldolase Plastoquinol-plastocyanin ATP synthase NADH reductase Triose phosphate + + + CO2 + H NAD + CoA-SH isomerase α-Ketoglutarate + Stage 2: 6-carbonTwo 3- NAD+ NADH + H + CO2 Glyceraldehyde 3-phosphate Dihydroxyacetone phosphate carbons Isocitrate α-Ketoglutarate dehydogenase dehydrogenase Glyceraldehyde + Pi + NAD Isocitrate complex 3-phosphate Succinyl CoA Oxidative Phosphorylation dehydrogenase NADH + H+ Electron Transport Chain GDP + Pi 1,3-Bisphosphoglycerate H+ Succinyl CoA GTP + CoA-SH Aconitase synthetase -

The Origins of Protein Phosphorylation
historical perspective The origins of protein phosphorylation Philip Cohen The reversible phosphorylation of proteins is central to the regulation of most aspects of cell func- tion but, even after the first protein kinase was identified, the general significance of this discovery was slow to be appreciated. Here I review the discovery of protein phosphorylation and give a per- sonal view of the key findings that have helped to shape the field as we know it today. he days when protein phosphorylation was an abstruse backwater, best talked Tabout between consenting adults in private, are over. My colleagues no longer cringe on hearing that “phosphorylase kinase phosphorylates phosphorylase” and their eyes no longer glaze over when a “”kinase kinase kinase” is mentioned. This is because protein phosphorylation has gradu- ally become an integral part of all the sys- tems they are studying themselves. Indeed it would be difficult to find anyone today who would disagree with the statement that “the reversible phosphorylation of proteins regu- lates nearly every aspect of cell life”. Phosphorylation and dephosphorylation, catalysed by protein kinases and protein phosphatases, can modify the function of a protein in almost every conceivable way; for Carl and Gerty Cori, the 1947 Nobel Laureates. Picture: Science Photo Library. example by increasing or decreasing its bio- logical activity, by stabilizing it or marking it for destruction, by facilitating or inhibiting movement between subcellular compart- so long before its general significance liver enzyme that catalysed the phosphory- ments, or by initiating or disrupting pro- was appreciated? lation of casein3. Soon after, Fischer and tein–protein interactions. -

Phosphorylation of Ubiquitin-Activating Enzyme in Cultured Cells
Proc. Natl. Acad. Sci. USA Vol. 92, pp. 3454-3457, April 1995 Biochemistry Phosphorylation of ubiquitin-activating enzyme in cultured cells JAMES C. COOK* AND P. BOON CHOCKt Laboratory of Biochemistry, Section on Metabolic Regulation, National Heart, Lung, and Blood Institute, National Institutes of Health, Building 3, Room 202, Bethesda, MD 20892 Communicated by Earl R. Stadtman, National Institutes of Health, Bethesda, MD, January 3, 1995 ABSTRACT Ubiquitin-activating enzyme, El, is the first protein ligase. Isoforms of all three enzymes have been enzyme in the pathway leading to formation of ubiquitin- described (11-14, 19). protein conjugates. El exists as two isoforms in human cells Obviously, ubiquitin conjugation must be a regulated pro- which are separable by electrophoresis. These isoforms mi- cess. Only certain proteins are recognized by the conjugating grate with apparent molecular sizes of 110 kDa and 117 kDa enzymes, and some are targets for conjugation only under in SDS/polyacrylamide gels. Immunoprecipitation ofEl from certain environmental conditions (15, 16) or during specific lysates of HeLa cells metabolically labeled with [32P]phos- phases of the cell cycle (17). Yet little is known about how this phate indicated the presence of a phosphorylated form of El regulation is accomplished. which migrates at 117 kDa. Phospho amino acid analysis Previous reports demonstrated that El can undergo in vitro identified serine as the phosphorylated residue in El. Phos- phosphorylation (18) and that it exists as two isoforms in phorylated El was also detected in normal and transformed human cells (19). These were designated "ElllOkDa" and cells from another human cell line. -

1,2,3,5,6,8,10,11,16,20,21 Citric Acid Cycle Reactions
Overview of the citric acid cycle, AKA the krebs cycle AKA tricarboxylic acid AKA TCA cycle Suggested problems from the end of chapter 19: 1,2,3,5,6,8,10,11,16,20,21 Glycogen is broken down into glucose. The reactions of glycolysis result in pyruvate, which is then fed into the citric acid cycle in the form of acetyl CoA. The products of the citric acid cycles are 2 CO2, 3 NADH, 1 FADH2, and 1 ATP or GTP. After pyruvate is generated, it is transported into the mitochondrion, an organelle that contains the citric acid cycle enzymes and the oxidative phosphorylation enzymes. In E. coli, where there are neither mitochondria nor other organelles, these enzymes also seem to be concentrated in certain regions in the cell. Citric acid cycle reactions Overall, there are 8 reactions that result in oxidation of the metabolic fuel. This results in reduction of NAD+ and FAD. NADH and FADH2 will transfer their electrons to oxygen during oxidative phosphorylation. •In 1936 Carl Martius and Franz Knoop showed that citrate can be formed non-enzymaticly from Oxaloacetate and pyruvate. •In 1937 Hans Krebs used this information for biochemical experiments that resulted in his suggestion that citrate is processed in an ongoing circle, into which pyruvate is “fed.” •In 1951 it was shown that it was acetyl Coenzyme-A that condenses with oxaloacetate to form citrate. 1 The pre-citric acid reaction- pyruvate dehydrogenase Pyruvate dehydrogenase is a multi-subunit complex, containing three enzymes that associate non-covalently and catalyze 5 reaction. The enzymes are: (E1) pyruvate dehydrogenase (E2) dihydrolipoyl transacetylase (E3) dihydrolipoyl dehydrogenase What are the advantages for arranging enzymes in complexes? E. -

O-Glcnacylation Is a Key Regulator of Multiple Cellular Metabolic Pathways
O-GlcNAcylation is a key regulator of multiple cellular metabolic pathways Hongshuo Zhang, Zhen Li, Yufei Wang and Ying Kong Core Laboratory of Glycobiology and Glycoengineering, College of Basic Medical Sciences, Dalian Medical University, Dalian, Liaoning, China ABSTRACT O-GlcNAcylation modifies proteins in serine or threonine residues in the nucleus, cytoplasm, and mitochondria. It regulates a variety of cellular biological processes and abnormal O-GlcNAcylation is associated with diabetes, cancer, cardiovascular disease, and neurodegenerative diseases. Recent evidence has suggested that O-GlcNAcylation acts as a nutrient sensor and signal integrator to regulate metabolic signaling, and that dysregulation of its metabolism may be an important indicator of pathogenesis in disease. Here, we review the literature focusing on O-GlcNAcylation regulation in major metabolic processes, such as glucose metabolism, mitochondrial oxidation, lipid metabolism, and amino acid metabolism. We discuss its role in physiological processes, such as cellular nutrient sensing and homeostasis maintenance. O-GlcNAcylation acts as a key regulator in multiple metabolic processes and pathways. Our review will provide a better understanding of how O-GlcNAcylation coordinates metabolism and integrates molecular networks. Subjects Biochemistry, Cell Biology, Metabolic Sciences Keywords O-GlcNAc, Nutrient sensing, Glucose uptake, Glycolysis, Mitochondria, Lipid metabolism, Glutamine, Signaling pathway, Homeostasis, TCA INTRODUCTION Increasing evidence has suggested -

Estrogen Receptor Phosphorylation Deborah A
Steroids 68 (2003) 1–9 Review Estrogen receptor phosphorylation Deborah A. Lannigan∗ Center for Cell Signaling, Health Sciences Center, University of Virginia, Hospital West, Room 7041, Box 800577, Charlottesville, VA 22908-0577, USA Received 30 April 2002; accepted 13 June 2002 Abstract Estrogen receptor ␣ (ER␣) is phosphorylated on multiple amino acid residues. For example, in response to estradiol binding, human ER␣ is predominately phosphorylated on Ser-118 and to a lesser extent on Ser-104 and Ser-106. In response to activation of the mitogen-activated protein kinase pathway, phosphorylation occurs on Ser-118 and Ser-167. These serine residues are all located within the activation function 1 region of the N-terminal domain of ER␣. In contrast, activation of protein kinase A increases the phosphorylation of Ser-236, which is located in the DNA-binding domain. The in vivo phosphorylation status of Tyr-537, located in the ligand-binding domain, remains controversial. In this review, I present evidence that these phosphorylations occur, and identify the kinases thought to be responsible. Additionally, the functional importance of ER␣ phosphorylation is discussed. © 2002 Elsevier Science Inc. All rights reserved. Keywords: Estrogen receptor; Phosphorylation; Transcription 1. Overview There are two known ER isoforms, ␣ and , which dif- fer in their ligand specificities and physiological functions This review will focus on the major phosphorylation sites [17–19]. There are also a number of splice variants for each in estrogen receptor ␣ (ER␣) that occur in response to ei- of the isoforms, some of which influence the activity of the ther estradiol or through the activation of second messen- wild type receptor [20–23]. -

The Crosstalk Between O-Glcnacylation and Phosphorylation in Insulin Signaling
THE CROSSTALK BETWEEN O-GLCNACYLATION AND PHOSPHORYLATION IN INSULIN SIGNALING by Ping Hu A dissertation submitted to Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland December, 2015 © Ping Hu All Rights Reserved ABSTRACT Extensive crosstalk between phosphorylation and O-GlcNAcylation is involved in regulating insulin signaling. In addition, many of the key regulators of insulin signaling have been shown to be O-GlcNAcylated. In this study, we first explored the crosstalk between phosphorylation and O-GlcNAcylation globally, and then focused on an upstream signaling regulator key to insulin signaling, insulin receptor substrate 1, IRS1. Through regulating cellular O-GlcNAcylation levels in differentiated adipocyte 3T3-L1 cells by specific OGT and OGA inhibitors, we measured the affects of O-GlcNAcylation on the phosphorylation status of key molecules. We observed that O-GlcNAcylation regulates the flux through the two main divergent cellular pathways of insulin signaling, one regulating glucose metabolism and the other regulating cellular proliferation via MAP kinase. Decreased O-GlcNAcylation was associated with increased glucose utilization, and increased O-GlcNAcylation was associated with increased insulin-induced cellular proliferation pathways. Since IRS1 is early in the insulin signaling pathway and is both highly O-GlcNAcylated and highly phosphorylated, we focused on the crosstalk of those two PTMs on IRS1. The phosphorylation spectrum of IRS1 revealed by 2D electrophoretic analyses, partially explained why decreased O-GlcNAcylation facilitated glucose uptake. Two new O- GlcNAcylation sites were mapped through CID/ETD mass spectrometry. With help of protein truncation and site mutagenesis techniques, we propose a mechanism suggesting that the O-GlcNAcylation of Ser635, may be a negative regulator of the C-terminal O- GlcNAcylations on IRS1.