Structural Basis of O-Glcnac Recognition by Mammalian 14-3-3 Proteins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Cross-Talk Between Methylation and Phosphorylation in Lymphoid-Specific Helicase Drives Cancer Stem-Like Properties
Signal Transduction and Targeted Therapy www.nature.com/sigtrans ARTICLE OPEN The cross-talk between methylation and phosphorylation in lymphoid-specific helicase drives cancer stem-like properties Na Liu1,2,3, Rui Yang1,2, Ying Shi1,2, Ling Chen1,2, Yating Liu1,2, Zuli Wang1,2, Shouping Liu1,2, Lianlian Ouyang4, Haiyan Wang1,2, Weiwei Lai1,2, Chao Mao1,2, Min Wang1,2, Yan Cheng5, Shuang Liu4, Xiang Wang6, Hu Zhou7, Ya Cao1,2, Desheng Xiao1 and Yongguang Tao1,2,6 Posttranslational modifications (PTMs) of proteins, including chromatin modifiers, play crucial roles in the dynamic alteration of various protein properties and functions including stem-cell properties. However, the roles of Lymphoid-specific helicase (LSH), a DNA methylation modifier, in modulating stem-like properties in cancer are still not clearly clarified. Therefore, exploring PTMs modulation of LSH activity will be of great significance to further understand the function and activity of LSH. Here, we demonstrate that LSH is capable to undergo PTMs, including methylation and phosphorylation. The arginine methyltransferase PRMT5 can methylate LSH at R309 residue, meanwhile, LSH could as well be phosphorylated by MAPK1 kinase at S503 residue. We further show that the accumulation of phosphorylation of LSH at S503 site exhibits downregulation of LSH methylation at R309 residue, which eventually promoting stem-like properties in lung cancer. Whereas, phosphorylation-deficient LSH S503A mutant promotes the accumulation of LSH methylation at R309 residue and attenuates stem-like properties, indicating the critical roles of LSH PTMs in modulating stem-like properties. Thus, our study highlights the importance of the crosstalk between LSH PTMs in determining its activity and function in lung cancer stem-cell maintenance. -

Tyrosine Kinase – Role and Significance in Cancer
Int. J. Med. Sci. 2004 1(2): 101-115 101 International Journal of Medical Sciences ISSN 1449-1907 www.medsci.org 2004 1(2):101-115 ©2004 Ivyspring International Publisher. All rights reserved Review Tyrosine kinase – Role and significance in Cancer Received: 2004.3.30 Accepted: 2004.5.15 Manash K. Paul and Anup K. Mukhopadhyay Published:2004.6.01 Department of Biotechnology, National Institute of Pharmaceutical Education and Research, Sector-67, S.A.S Nagar, Mohali, Punjab, India-160062 Abstract Tyrosine kinases are important mediators of the signaling cascade, determining key roles in diverse biological processes like growth, differentiation, metabolism and apoptosis in response to external and internal stimuli. Recent advances have implicated the role of tyrosine kinases in the pathophysiology of cancer. Though their activity is tightly regulated in normal cells, they may acquire transforming functions due to mutation(s), overexpression and autocrine paracrine stimulation, leading to malignancy. Constitutive oncogenic activation in cancer cells can be blocked by selective tyrosine kinase inhibitors and thus considered as a promising approach for innovative genome based therapeutics. The modes of oncogenic activation and the different approaches for tyrosine kinase inhibition, like small molecule inhibitors, monoclonal antibodies, heat shock proteins, immunoconjugates, antisense and peptide drugs are reviewed in light of the important molecules. As angiogenesis is a major event in cancer growth and proliferation, tyrosine kinase inhibitors as a target for anti-angiogenesis can be aptly applied as a new mode of cancer therapy. The review concludes with a discussion on the application of modern techniques and knowledge of the kinome as means to gear up the tyrosine kinase drug discovery process. -

Principles of Protein Phosphorylation Biophysical Chemistry 1, Fall 2010
Principles of protein phosphorylation Biophysical Chemistry 1, Fall 2010 Signalling “cascades” Structural biology of phosphorylation Web assignment: http://pkr.genomics.purdue.edu Reversible protein phosphorylation Enzymatic reaction Posttranslational Control ΔG~12kcal/mol Kinases and phosphatases Phosphatase Kinase dephospho- phosphorylates rylates General Examples Signalling overview Cells are way too complex! MAPK/ERK Signaling Pathway RousExample:sarcoma virus Rous (RSV) sarcoma virus (RSV) gag - encodes capsid proteins pol - encodes reverse transcriptase env - encodes envelope proteins src - encodes a tyrosine kinase that attaches phosphate groups to the amino acid tyrosine in host cell proteins MutationsExample:, viruses Rous and cancer sarcoma virus (RSV) v-src lacks the C-terminal inhibitory phosphorylation site (tyrosine-527), and is therefore constitutively active as opposed to normal src (c-src) Continuous cell profileration tumor BiophysicsStructural of signalling Effect of Phosphorylation Phosphorylation is an important regulatory mechanism Can reversible attach/detach a phosphate and therefore switch “on”/”off” the function Effect of phosphorylation is manifold • Conformational change • Ordering/disordering • Electrostatic effects • Alternate binding behavior Signalling by reorientation Reorientation: A conformational switch Rmsd = 2.5Å DHP (red) and Z-Score = 4.6 DHPs74e (blue) (>3.6 same fold) Title: A phosphorylation-induced conformation change in dematin headpiece Author(s): Jiang ZHG, McKnight CJ Source: STRUCTURE Volume: 14 Issue: 2 Pages: 379-387 Published: FEB 2006 Order/disorderDisordering:transitions NtrC, a molecular switch upon phosphorylation Orange-yellow: unphosphorylated NtrC blue-cyan: phosphorylated NtrC Volkman et al., Science 2001, 291, 2429-33 Alternate Binding: SRC SH2 Src/SH2 interactions: binding vs release domain binding ExpectedCan conformationalwe understand effects (predict) the effect of phosphorylation Electrostatics Hydrogen bonding Size Mutational analysis Experimental data Lubman, O.Y. -

Phosphorylation of Chicken Protein Tyrosine Phosphatase 1 by Casein Kinase II in Vitro
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 29, No 4, 229-233, December 1997 Phosphorylation of chicken protein tyrosine phosphatase 1 by casein kinase II in vitro Eun Joo Jung,1 Kee Ryeon Kang1 and Introduction Yoon-Se Kang1,2 The phosphorylation of protein tyrosine residues is an early event in signal transduction initiated by binding of 1 Department of Biochemistry and Gyeongsang Institute of Cancer growth factors and hormones to their cognate receptors Research, College of Medicine, Gyeongsang National University, and it leads to regulation of cellular activities which include Chinju 660-280, Korea proliferation, differentiation, and also malignant transfor- 2 Corresponding author mation of cells (Hunter, 1989; Ullirich and Schlessinger, Accepted 17 November 1997 1990; Cantley et al., 1991). Under normal conditions, the level of tyrosine phosphorylation within a cell is determined by a balance between the actions of protein tyrosine Abbreviations: CPTP, chicken protein tyrosine phosphatase; HPTP1B, human placenta kinases (PTKs) and protein tyrosine phosphatases (PTPs) protein tyrosine phosphatase 1B; CKII, casein kinase II; MAP kinase, mitogen-activated (Hunter, 1989; Fischer et al., 1991; Trowbridge, 1991). protein kinase; GST, glutathione S-transferase; pNPP, p-nitrophenyl phosphate; EGF, PTPs do not simply reverse the action of tyrosine kinases, epidermal growth factor but rather, PTP itself may play a central role in cellular regulation. PTPs are generally classified as transmem- brane (receptor-type) and nontransmembrane (nonrecep- tor-type) enzymes based on the presence or absence of extracellular and transmembrane portions of their predicted sequence (Fischer et al., 1991). Because the activity of Abstract tyrosine kinase can be controlled by phosphorylation, it has been postulated that PTP activity may be regulated The phosphorylation and dephosphorylation of by phosphorylation as well. -

Citric Acid Cycle
CHEM464 / Medh, J.D. The Citric Acid Cycle Citric Acid Cycle: Central Role in Catabolism • Stage II of catabolism involves the conversion of carbohydrates, fats and aminoacids into acetylCoA • In aerobic organisms, citric acid cycle makes up the final stage of catabolism when acetyl CoA is completely oxidized to CO2. • Also called Krebs cycle or tricarboxylic acid (TCA) cycle. • It is a central integrative pathway that harvests chemical energy from biological fuel in the form of electrons in NADH and FADH2 (oxidation is loss of electrons). • NADH and FADH2 transfer electrons via the electron transport chain to final electron acceptor, O2, to form H2O. Entry of Pyruvate into the TCA cycle • Pyruvate is formed in the cytosol as a product of glycolysis • For entry into the TCA cycle, it has to be converted to Acetyl CoA. • Oxidation of pyruvate to acetyl CoA is catalyzed by the pyruvate dehydrogenase complex in the mitochondria • Mitochondria consist of inner and outer membranes and the matrix • Enzymes of the PDH complex and the TCA cycle (except succinate dehydrogenase) are in the matrix • Pyruvate translocase is an antiporter present in the inner mitochondrial membrane that allows entry of a molecule of pyruvate in exchange for a hydroxide ion. 1 CHEM464 / Medh, J.D. The Citric Acid Cycle The Pyruvate Dehydrogenase (PDH) complex • The PDH complex consists of 3 enzymes. They are: pyruvate dehydrogenase (E1), Dihydrolipoyl transacetylase (E2) and dihydrolipoyl dehydrogenase (E3). • It has 5 cofactors: CoASH, NAD+, lipoamide, TPP and FAD. CoASH and NAD+ participate stoichiometrically in the reaction, the other 3 cofactors have catalytic functions. -
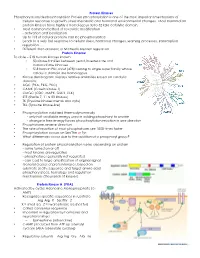
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Gene-Expression Signature Regulated by the KEAP1-NRF2-CUL3 Axis Is Associated with a Poor Prognosis in Head and Neck Squamous Cell Cancer Akhileshwar Namani1†, Md
Namani et al. BMC Cancer (2018) 18:46 DOI 10.1186/s12885-017-3907-z RESEARCH ARTICLE Open Access Gene-expression signature regulated by the KEAP1-NRF2-CUL3 axis is associated with a poor prognosis in head and neck squamous cell cancer Akhileshwar Namani1†, Md. Matiur Rahaman2†, Ming Chen2* and Xiuwen Tang1* Abstract Background: NRF2 is the key regulator of oxidative stress in normal cells and aberrant expression of the NRF2 pathway due to genetic alterations in the KEAP1 (Kelch-like ECH-associated protein 1)-NRF2 (nuclear factor erythroid 2 like 2)-CUL3 (cullin 3) axis leads to tumorigenesis and drug resistance in many cancers including head and neck squamous cell cancer (HNSCC). The main goal of this study was to identify specific genes regulated by the KEAP1-NRF2-CUL3 axis in HNSCC patients, to assess the prognostic value of this gene signature in different cohorts, and to reveal potential biomarkers. Methods: RNA-Seq V2 level 3 data from 279 tumor samples along with 37 adjacent normal samples from patients enrolled in the The Cancer Genome Atlas (TCGA)-HNSCC study were used to identify upregulated genes using two methods (altered KEAP1-NRF2-CUL3 versus normal, and altered KEAP1-NRF2-CUL3 versus wild-type). We then used a new approach to identify the combined gene signature by integrating both datasets and subsequently tested this signature in 4 independent HNSCC datasets to assess its prognostic value. In addition, functional annotation using the DAVID v6.8 database and protein-protein interaction (PPI) analysis using the STRING v10 databasewereperformedonthesignature. Results: A signature composed of a subset of 17 genes regulated by the KEAP1-NRF2-CUL3 axis was identified by overlapping both the upregulated genes of altered versus normal (251 genes) and altered versus wild-type (25 genes) datasets. -

Β-Catenin Signaling Dynamics Regulate Cell Fate in Differentiating Neural Stem Cells
β-Catenin signaling dynamics regulate cell fate in differentiating neural stem cells Alyssa B. Rosenblooma, Marcin Tarczynski a, Nora Lama, Ravi S. Kaneb,1, Lukasz J. Bugaja,c,1, and David V. Schaffera,d,e,f,1 aDepartment of Bioengineering, University of California, Berkeley, CA 94720; bSchool of Chemical & Biomolecular Engineering, Georgia Institute of Technology, Atlanta, GA 30332; cDepartment of Bioengineering, University of Pennsylvania, Philadelphia, PA 19104; dDepartment of Chemical and Biomolecular Engineering, University of California, Berkeley, CA 94720; eDepartment of Molecular and Cell Biology, University of California, Berkeley, CA 94720; and fHelen Wills Neuroscience Institute, University of California, Berkeley, CA 94720 Edited by Randall T. Moon, University of Washington, Seattle, WA, and approved September 21, 2020 (received for review May 4, 2020) Stem cells undergo differentiation in complex and dynamic environ- of how signaling dynamics impact cellular function. Optogenetics ments wherein instructive signals fluctuate on various timescales. has recently emerged as a field in which light—which can readily Thus, cells must be equipped to properly respond to the timing of be varied in intensity, space, and time—is harnessed to precisely signals, for example, to distinguish sustained signaling from transient modulate cell-signaling pathways. In this approach, light-sensitive noise. However, how stem cells respond to dynamic variations in proteins are engineered to interface with specific signaling path- differentiation cues is not well characterized. Here, we use optoge- ways, and the subsequent introduction of such an optogenetic netic activation of β-catenin signaling to probe the dynamic responses construct into cells renders the signaling pathway responsive to of differentiating adult neural stem cells (NSCs). -

Glycolysis Citric Acid Cycle Oxidative Phosphorylation Calvin Cycle Light
Stage 3: RuBP regeneration Glycolysis Ribulose 5- Light-Dependent Reaction (Cytosol) phosphate 3 ATP + C6H12O6 + 2 NAD + 2 ADP + 2 Pi 3 ADP + 3 Pi + + 1 GA3P 6 NADP + H Pi NADPH + ADP + Pi ATP 2 C3H4O3 + 2 NADH + 2 H + 2 ATP + 2 H2O 3 CO2 Stage 1: ATP investment ½ glucose + + Glucose 2 H2O 4H + O2 2H Ferredoxin ATP Glyceraldehyde 3- Ribulose 1,5- Light Light Fx iron-sulfur Sakai-Kawada, F Hexokinase phosphate bisphosphate - 4e + center 2016 ADP Calvin Cycle 2H Stroma Mn-Ca cluster + 6 NADP + Light-Independent Reaction Phylloquinone Glucose 6-phosphate + 6 H + 6 Pi Thylakoid Tyr (Stroma) z Fe-S Cyt f Stage 1: carbon membrane Phosphoglucose 6 NADPH P680 P680* PQH fixation 2 Plastocyanin P700 P700* D-(+)-Glucose isomerase Cyt b6 1,3- Pheophytin PQA PQB Fructose 6-phosphate Bisphosphoglycerate ATP Lumen Phosphofructokinase-1 3-Phosphoglycerate ADP Photosystem II P680 2H+ Photosystem I P700 Stage 2: 3-PGA Photosynthesis Fructose 1,6-bisphosphate reduction 2H+ 6 ADP 6 ATP 6 CO2 + 6 H2O C6H12O6 + 6 O2 H+ + 6 Pi Cytochrome b6f Aldolase Plastoquinol-plastocyanin ATP synthase NADH reductase Triose phosphate + + + CO2 + H NAD + CoA-SH isomerase α-Ketoglutarate + Stage 2: 6-carbonTwo 3- NAD+ NADH + H + CO2 Glyceraldehyde 3-phosphate Dihydroxyacetone phosphate carbons Isocitrate α-Ketoglutarate dehydogenase dehydrogenase Glyceraldehyde + Pi + NAD Isocitrate complex 3-phosphate Succinyl CoA Oxidative Phosphorylation dehydrogenase NADH + H+ Electron Transport Chain GDP + Pi 1,3-Bisphosphoglycerate H+ Succinyl CoA GTP + CoA-SH Aconitase synthetase -

Anti-KEAP1 Antibody (ARG66711)
Product datasheet [email protected] ARG66711 Package: 100 μg anti-KEAP1 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes KEAP1 Tested Reactivity Hu Tested Application IHC-P, WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name KEAP1 Species Human Immunogen Synthetic peptide within aa. 411-460 of Human KEAP1. Conjugation Un-conjugated Alternate Names KLHL19; Cytosolic inhibitor of Nrf2; INrf2; Kelch-like protein 19; Kelch-like ECH-associated protein 1 Application Instructions Application table Application Dilution IHC-P 1:100 - 1:300 WB 1:500 - 1:2000 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Positive Control K562 Observed Size ~ 70 kDa Properties Form Liquid Purification Affinity purification with immunogen. Buffer PBS, 0.02% Sodium azide, 50% Glycerol and 0.5% BSA. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol and 0.5% BSA Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. www.arigobio.com 1/3 Bioinformation Gene Symbol KEAP1 Gene Full Name kelch-like ECH-associated protein 1 Background This gene encodes a protein containing KELCH-1 like domains, as well as a BTB/POZ domain. Kelch-like ECH-associated protein 1 interacts with NF-E2-related factor 2 in a redox-sensitive manner and the dissociation of the proteins in the cytoplasm is followed by transportation of NF-E2-related factor 2 to the nucleus. -

The Origins of Protein Phosphorylation
historical perspective The origins of protein phosphorylation Philip Cohen The reversible phosphorylation of proteins is central to the regulation of most aspects of cell func- tion but, even after the first protein kinase was identified, the general significance of this discovery was slow to be appreciated. Here I review the discovery of protein phosphorylation and give a per- sonal view of the key findings that have helped to shape the field as we know it today. he days when protein phosphorylation was an abstruse backwater, best talked Tabout between consenting adults in private, are over. My colleagues no longer cringe on hearing that “phosphorylase kinase phosphorylates phosphorylase” and their eyes no longer glaze over when a “”kinase kinase kinase” is mentioned. This is because protein phosphorylation has gradu- ally become an integral part of all the sys- tems they are studying themselves. Indeed it would be difficult to find anyone today who would disagree with the statement that “the reversible phosphorylation of proteins regu- lates nearly every aspect of cell life”. Phosphorylation and dephosphorylation, catalysed by protein kinases and protein phosphatases, can modify the function of a protein in almost every conceivable way; for Carl and Gerty Cori, the 1947 Nobel Laureates. Picture: Science Photo Library. example by increasing or decreasing its bio- logical activity, by stabilizing it or marking it for destruction, by facilitating or inhibiting movement between subcellular compart- so long before its general significance liver enzyme that catalysed the phosphory- ments, or by initiating or disrupting pro- was appreciated? lation of casein3. Soon after, Fischer and tein–protein interactions. -

Protein Tyrosine Kinases: Their Roles and Their Targeting in Leukemia
cancers Review Protein Tyrosine Kinases: Their Roles and Their Targeting in Leukemia Kalpana K. Bhanumathy 1,*, Amrutha Balagopal 1, Frederick S. Vizeacoumar 2 , Franco J. Vizeacoumar 1,3, Andrew Freywald 2 and Vincenzo Giambra 4,* 1 Division of Oncology, College of Medicine, University of Saskatchewan, Saskatoon, SK S7N 5E5, Canada; [email protected] (A.B.); [email protected] (F.J.V.) 2 Department of Pathology and Laboratory Medicine, College of Medicine, University of Saskatchewan, Saskatoon, SK S7N 5E5, Canada; [email protected] (F.S.V.); [email protected] (A.F.) 3 Cancer Research Department, Saskatchewan Cancer Agency, 107 Wiggins Road, Saskatoon, SK S7N 5E5, Canada 4 Institute for Stem Cell Biology, Regenerative Medicine and Innovative Therapies (ISBReMIT), Fondazione IRCCS Casa Sollievo della Sofferenza, 71013 San Giovanni Rotondo, FG, Italy * Correspondence: [email protected] (K.K.B.); [email protected] (V.G.); Tel.: +1-(306)-716-7456 (K.K.B.); +39-0882-416574 (V.G.) Simple Summary: Protein phosphorylation is a key regulatory mechanism that controls a wide variety of cellular responses. This process is catalysed by the members of the protein kinase su- perfamily that are classified into two main families based on their ability to phosphorylate either tyrosine or serine and threonine residues in their substrates. Massive research efforts have been invested in dissecting the functions of tyrosine kinases, revealing their importance in the initiation and progression of human malignancies. Based on these investigations, numerous tyrosine kinase inhibitors have been included in clinical protocols and proved to be effective in targeted therapies for various haematological malignancies.