Mouse Napb Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -
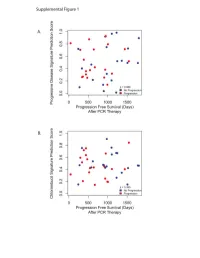
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

(12) United States Patent (10) Patent No.: US 7.873,482 B2 Stefanon Et Al
US007873482B2 (12) United States Patent (10) Patent No.: US 7.873,482 B2 Stefanon et al. (45) Date of Patent: Jan. 18, 2011 (54) DIAGNOSTIC SYSTEM FOR SELECTING 6,358,546 B1 3/2002 Bebiak et al. NUTRITION AND PHARMACOLOGICAL 6,493,641 B1 12/2002 Singh et al. PRODUCTS FOR ANIMALS 6,537,213 B2 3/2003 Dodds (76) Inventors: Bruno Stefanon, via Zilli, 51/A/3, Martignacco (IT) 33035: W. Jean Dodds, 938 Stanford St., Santa Monica, (Continued) CA (US) 90403 FOREIGN PATENT DOCUMENTS (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 WO WO99-67642 A2 12/1999 U.S.C. 154(b) by 158 days. (21)21) Appl. NoNo.: 12/316,8249 (Continued) (65) Prior Publication Data Swanson, et al., “Nutritional Genomics: Implication for Companion Animals'. The American Society for Nutritional Sciences, (2003).J. US 2010/O15301.6 A1 Jun. 17, 2010 Nutr. 133:3033-3040 (18 pages). (51) Int. Cl. (Continued) G06F 9/00 (2006.01) (52) U.S. Cl. ........................................................ 702/19 Primary Examiner—Edward Raymond (58) Field of Classification Search ................... 702/19 (74) Attorney, Agent, or Firm Greenberg Traurig, LLP 702/23, 182–185 See application file for complete search history. (57) ABSTRACT (56) References Cited An analysis of the profile of a non-human animal comprises: U.S. PATENT DOCUMENTS a) providing a genotypic database to the species of the non 3,995,019 A 1 1/1976 Jerome human animal Subject or a selected group of the species; b) 5,691,157 A 1 1/1997 Gong et al. -

The DNA Sequence and Comparative Analysis of Human Chromosome 20
articles The DNA sequence and comparative analysis of human chromosome 20 P. Deloukas, L. H. Matthews, J. Ashurst, J. Burton, J. G. R. Gilbert, M. Jones, G. Stavrides, J. P. Almeida, A. K. Babbage, C. L. Bagguley, J. Bailey, K. F. Barlow, K. N. Bates, L. M. Beard, D. M. Beare, O. P. Beasley, C. P. Bird, S. E. Blakey, A. M. Bridgeman, A. J. Brown, D. Buck, W. Burrill, A. P. Butler, C. Carder, N. P. Carter, J. C. Chapman, M. Clamp, G. Clark, L. N. Clark, S. Y. Clark, C. M. Clee, S. Clegg, V. E. Cobley, R. E. Collier, R. Connor, N. R. Corby, A. Coulson, G. J. Coville, R. Deadman, P. Dhami, M. Dunn, A. G. Ellington, J. A. Frankland, A. Fraser, L. French, P. Garner, D. V. Grafham, C. Grif®ths, M. N. D. Grif®ths, R. Gwilliam, R. E. Hall, S. Hammond, J. L. Harley, P. D. Heath, S. Ho, J. L. Holden, P. J. Howden, E. Huckle, A. R. Hunt, S. E. Hunt, K. Jekosch, C. M. Johnson, D. Johnson, M. P. Kay, A. M. Kimberley, A. King, A. Knights, G. K. Laird, S. Lawlor, M. H. Lehvaslaiho, M. Leversha, C. Lloyd, D. M. Lloyd, J. D. Lovell, V. L. Marsh, S. L. Martin, L. J. McConnachie, K. McLay, A. A. McMurray, S. Milne, D. Mistry, M. J. F. Moore, J. C. Mullikin, T. Nickerson, K. Oliver, A. Parker, R. Patel, T. A. V. Pearce, A. I. Peck, B. J. C. T. Phillimore, S. R. Prathalingam, R. W. Plumb, H. Ramsay, C. M. -

Variation in Protein Coding Genes Identifies Information Flow
bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 1 1 2 3 4 5 Variation in protein coding genes identifies information flow as a contributor to 6 animal complexity 7 8 Jack Dean, Daniela Lopes Cardoso and Colin Sharpe* 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Institute of Biological and Biomedical Sciences 25 School of Biological Science 26 University of Portsmouth, 27 Portsmouth, UK 28 PO16 7YH 29 30 * Author for correspondence 31 [email protected] 32 33 Orcid numbers: 34 DLC: 0000-0003-2683-1745 35 CS: 0000-0002-5022-0840 36 37 38 39 40 41 42 43 44 45 46 47 48 49 Abstract bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 2 1 Across the metazoans there is a trend towards greater organismal complexity. How 2 complexity is generated, however, is uncertain. Since C.elegans and humans have 3 approximately the same number of genes, the explanation will depend on how genes are 4 used, rather than their absolute number. -

Chromatin Conformation Links Distal Target Genes to CKD Loci
BASIC RESEARCH www.jasn.org Chromatin Conformation Links Distal Target Genes to CKD Loci Maarten M. Brandt,1 Claartje A. Meddens,2,3 Laura Louzao-Martinez,4 Noortje A.M. van den Dungen,5,6 Nico R. Lansu,2,3,6 Edward E.S. Nieuwenhuis,2 Dirk J. Duncker,1 Marianne C. Verhaar,4 Jaap A. Joles,4 Michal Mokry,2,3,6 and Caroline Cheng1,4 1Experimental Cardiology, Department of Cardiology, Thoraxcenter Erasmus University Medical Center, Rotterdam, The Netherlands; and 2Department of Pediatrics, Wilhelmina Children’s Hospital, 3Regenerative Medicine Center Utrecht, Department of Pediatrics, 4Department of Nephrology and Hypertension, Division of Internal Medicine and Dermatology, 5Department of Cardiology, Division Heart and Lungs, and 6Epigenomics Facility, Department of Cardiology, University Medical Center Utrecht, Utrecht, The Netherlands ABSTRACT Genome-wide association studies (GWASs) have identified many genetic risk factors for CKD. However, linking common variants to genes that are causal for CKD etiology remains challenging. By adapting self-transcribing active regulatory region sequencing, we evaluated the effect of genetic variation on DNA regulatory elements (DREs). Variants in linkage with the CKD-associated single-nucleotide polymorphism rs11959928 were shown to affect DRE function, illustrating that genes regulated by DREs colocalizing with CKD-associated variation can be dysregulated and therefore, considered as CKD candidate genes. To identify target genes of these DREs, we used circular chro- mosome conformation capture (4C) sequencing on glomerular endothelial cells and renal tubular epithelial cells. Our 4C analyses revealed interactions of CKD-associated susceptibility regions with the transcriptional start sites of 304 target genes. Overlap with multiple databases confirmed that many of these target genes are involved in kidney homeostasis. -

Quantitative Trait Loci Mapping of Macrophage Atherogenic Phenotypes
QUANTITATIVE TRAIT LOCI MAPPING OF MACROPHAGE ATHEROGENIC PHENOTYPES BRIAN RITCHEY Bachelor of Science Biochemistry John Carroll University May 2009 submitted in partial fulfillment of requirements for the degree DOCTOR OF PHILOSOPHY IN CLINICAL AND BIOANALYTICAL CHEMISTRY at the CLEVELAND STATE UNIVERSITY December 2017 We hereby approve this thesis/dissertation for Brian Ritchey Candidate for the Doctor of Philosophy in Clinical-Bioanalytical Chemistry degree for the Department of Chemistry and the CLEVELAND STATE UNIVERSITY College of Graduate Studies by ______________________________ Date: _________ Dissertation Chairperson, Johnathan D. Smith, PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, David J. Anderson, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Baochuan Guo, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Stanley L. Hazen, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Renliang Zhang, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Aimin Zhou, PhD Department of Chemistry, Cleveland State University Date of Defense: October 23, 2017 DEDICATION I dedicate this work to my entire family. In particular, my brother Greg Ritchey, and most especially my father Dr. Michael Ritchey, without whose support none of this work would be possible. I am forever grateful to you for your devotion to me and our family. You are an eternal inspiration that will fuel me for the remainder of my life. I am extraordinarily lucky to have grown up in the family I did, which I will never forget. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -

Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and Their Roles in Complex Disease Jeremy J
The University of Maine DigitalCommons@UMaine Electronic Theses and Dissertations Fogler Library 12-2013 Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and their Roles in Complex Disease Jeremy J. Jay Follow this and additional works at: http://digitalcommons.library.umaine.edu/etd Part of the Computer Sciences Commons Recommended Citation Jay, Jeremy J., "Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and their Roles in Complex Disease" (2013). Electronic Theses and Dissertations. 2140. http://digitalcommons.library.umaine.edu/etd/2140 This Open-Access Dissertation is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Electronic Theses and Dissertations by an authorized administrator of DigitalCommons@UMaine. CONTEXTUAL ANALYSIS OF LARGE-SCALE BIOMEDICAL ASSOCIATIONS FOR THE ELUCIDATION AND PRIORITIZATION OF GENES AND THEIR ROLES IN COMPLEX DISEASE By Jeremy J. Jay B.S.I. Baylor University, 2006 M.S. University of Tennessee, 2009 A DISSERTATION Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy (in Computer Science) The Graduate School The University of Maine December 2013 Advisory Committee: George Markowsky, Professor, Advisor Elissa J Chesler, Associate Professor, The Jackson Laboratory Erich J Baker, Associate Professor, Baylor University Judith Blake, Associate Professor, The Jackson Laboratory James Fastook, Professor DISSERTATION ACCEPTANCE STATEMENT On behalf of the Graduate Committee for Jeremy J. Jay, I affirm that this manuscript is the final and accepted dissertation. Signatures of all committee members are on file with the Graduate School at the University of Maine, 42 Stodder Hall, Orono, Maine. -

Initial Findings of Striatum Tripartite Model in OCD Brain Samples Based
www.nature.com/scientificreports OPEN Initial fndings of striatum tripartite model in OCD brain samples based on transcriptome analysis Received: 1 May 2018 Bianca C. G. Lisboa1, Katia C. Oliveira1, Ana Carolina Tahira 1, André Rocha Barbosa1,2, Accepted: 17 December 2018 Arthur Sant’Anna Feltrin3, Gisele Gouveia1, Luzia Lima1, Ana Cecília Feio dos Santos 1, Published: xx xx xxxx David Correa Martins Jr.3, Renato David Puga4, Ariane Cristine Moretto1, Carlos Alberto De Bragança Pereira2, Beny Lafer1, Renata Elaine Paraizo Leite1, Renata Eloah De Lucena Ferretti-Rebustini1, Jose Marcelo Farfel1, Lea Tenenholz Grinberg1,5, Wilson Jacob-Filho1, Euripedes Constantino Miguel1, Marcelo Queiroz Hoexter1 & Helena Brentani1,2 Obsessive-compulsive disorder (OCD) is a psychiatric disorder characterized by obsessions and/or compulsions. Diferent striatal subregions belonging to the cortico-striato-thalamic circuitry (CSTC) play an important role in the pathophysiology of OCD. The transcriptomes of 3 separate striatal areas (putamen (PT), caudate nucleus (CN) and accumbens nucleus (NAC)) from postmortem brain tissue were compared between 6 OCD and 8 control cases. In addition to network connectivity deregulation, diferent biological processes are specifc to each striatum region according to the tripartite model of the striatum and contribute in various ways to OCD pathophysiology. Specifcally, regulation of neurotransmitter levels and presynaptic processes involved in chemical synaptic transmission were shared between NAC and PT. The Gene Ontology terms cellular response to chemical stimulus, response to external stimulus, response to organic substance, regulation of synaptic plasticity, and modulation of synaptic transmission were shared between CN and PT. Most genes harboring common and/or rare variants previously associated with OCD that were diferentially expressed or part of a least preserved coexpression module in our study also suggest striatum subregion specifcity. -
![MSF (SEPT9) Mouse Monoclonal Antibody [Clone ID: OTI2B12] Product Data](https://docslib.b-cdn.net/cover/9390/msf-sept9-mouse-monoclonal-antibody-clone-id-oti2b12-product-data-4319390.webp)
MSF (SEPT9) Mouse Monoclonal Antibody [Clone ID: OTI2B12] Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for CF812819 MSF (SEPT9) Mouse Monoclonal Antibody [Clone ID: OTI2B12] Product data: Product Type: Primary Antibodies Clone Name: OTI2B12 Applications: WB Recommended Dilution: WB 1:500~2000 Reactivity: Human Host: Mouse Isotype: IgG1 Clonality: Monoclonal Immunogen: Full length human recombinant protein of human SEPT9 (NP_006631) produced in HEK293T cell. Formulation: Lyophilized powder (original buffer 1X PBS, pH 7.3, 8% trehalose) Reconstitution Method: For reconstitution, we recommend adding 100uL distilled water to a final antibody concentration of about 1 mg/mL. To use this carrier-free antibody for conjugation experiment, we strongly recommend performing another round of desalting process. (OriGene recommends Zeba Spin Desalting Columns, 7KMWCO from Thermo Scientific) Purification: Purified from mouse ascites fluids or tissue culture supernatant by affinity chromatography (protein A/G) Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 63.5 kDa Gene Name: Homo sapiens septin 9 (SEPTIN9), transcript variant 3, mRNA. Database Link: NP_006631 Entrez Gene 10801 Human Q9UHD8 This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 MSF (SEPT9) Mouse Monoclonal Antibody [Clone ID: OTI2B12] – CF812819 Background: This gene is a member of the septin family involved in cytokinesis and cell cycle control. This gene is a candidate for the ovarian tumor suppressor gene. -

Identification of Three Novel Ca Channel Subunit Genes Reveals
Downloaded from genome.cshlp.org on October 4, 2021 - Published by Cold Spring Harbor Laboratory Press Letter Identification of Three Novel Ca2+ Channel ␥ Subunit Genes Reveals Molecular Diversification by Tandem and Chromosome Duplication Daniel L. Burgess,1,2 Caleb F. Davis,1 Lisa A. Gefrides,1 and Jeffrey L. Noebels1 1Department of Neurology, Baylor College of Medicine, Houston, Texas 77030 USA Gene duplication is believed to be an important evolutionary mechanism for generating functional diversity within genomes. The accumulated products of ancient duplication events can be readily observed among the genes encoding voltage-dependent Ca2+ ion channels. Ten paralogous genes have been identified that encode isoforms of the ␣1 subunit, four that encode  subunits, and three that encode ␣2␦ subunits. Until recently, only a single gene encoding a muscle-specific isoform of the Ca2+ channel ␥ subunit (CACNG1) was known. Expression of a distantly related gene in the brain was subsequently demonstrated upon isolation of the Cacng2 gene, which is mutated in the mouse neurological mutant stargazer (stg). In this study, we sought to identify additional genes that encoded ␥ subunits. Because gene duplication often generates paralogs that remain in close syntenic proximity (tandem duplication) or are copied onto related daughter chromosomes (chromosome or whole-genome duplication), we hypothesized that the known positions of CACNG1 and CACNG2 could be used to predict the likely locations of additional ␥ subunit genes. Low-stringency genomic sequence analysis of targeted regions led to the identification of three novel Ca2+ channel ␥ subunit genes, CACNG3, CACNG4, and CACNG5,on chromosomes 16 and 17. These results demonstrate the value of genome evolution models for the identification of distantly related members of gene families.