Reactor Performance and Microbial Analysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Piscine Mycobacteriosis
Piscine Importance The genus Mycobacterium contains more than 150 species, including the obligate Mycobacteriosis pathogens that cause tuberculosis in mammals as well as environmental saprophytes that occasionally cause opportunistic infections. At least 20 species are known to Fish Tuberculosis, cause mycobacteriosis in fish. They include Mycobacterium marinum, some of its close relatives (e.g., M. shottsii, M. pseudoshottsii), common environmental Piscine Tuberculosis, organisms such as M. fortuitum, M. chelonae, M. abscessus and M. gordonae, and Swimming Pool Granuloma, less well characterized species such as M. salmoniphilum and M. haemophilum, Fish Tank Granuloma, among others. Piscine mycobacteriosis, which has a range of outcomes from Fish Handler’s Disease, subclinical infection to death, affects a wide variety of freshwater and marine fish. It Fish Handler’s Nodules has often been reported from aquariums, research laboratories and fish farms, but outbreaks also occur in free-living fish. The same organisms sometimes affect other vertebrates including people. Human infections acquired from fish are most often Last Updated: November 2020 characterized by skin lesions of varying severity, which occasionally spread to underlying joints and tendons. Some lesions may be difficult to cure, especially in those who are immunocompromised. Etiology Mycobacteriosis is caused by members of the genus Mycobacterium, which are Gram-positive, acid fast, pleomorphic rods in the family Mycobacteriaceae and order Actinomycetales. This genus is traditionally divided into two groups: the members of the Mycobacterium tuberculosis complex (e.g., M. tuberculosis, M. bovis, M. caprae, M. pinnipedii), which cause tuberculosis in mammals, and the nontuberculous mycobacteria. The organisms in the latter group include environmental saprophytes, which sometimes cause opportunistic infections, and other species such as M. -

Mycobacterium Kansasii, Strain 824 Catalog No. NR-44269 For
Product Information Sheet for NR-44269 Mycobacterium kansasii, Strain 824 at -60°C or colder immediately upon arrival. For long-term storage, the vapor phase of a liquid nitrogen freezer is recommended. Freeze-thaw cycles should be avoided. Catalog No. NR-44269 Growth Conditions: For research use only. Not for human use. Media: Middlebrook 7H9 broth with ADC enrichment or equivalent Contributor: Middlebrook 7H10 agar with OADC enrichment or Diane Ordway, Ph.D., Assistant Professor, Department of Lowenstein-Jensen agar or equivalent Microbiology, Immunology and Pathology, Colorado State Incubation: University, Fort Collins, Colorado, USA and National Institute Temperature: 37°C of Allergy and Infectious Diseases (NIAID), National Institutes Atmosphere: Aerobic with 5% CO2 of Health (NIH), Bethesda, Maryland, USA Propagation: 1. Keep vial frozen until ready for use; then thaw. Manufacturer: 2. Transfer the entire thawed aliquot into a single tube of BEI Resources broth. 3. Use several drops of the suspension to inoculate an agar Product Description: slant and/or plate. Bacteria Classification: Mycobacteriaceae, Mycobacterium 4. Incubate the tube, slant and/or plate at 37°C for 1 to 6 Species: Mycobacterium kansasii weeks. Strain: 824 Original Source: Mycobacterium kansasii (M. kansasii), strain Citation: 824 was isolated in 2012 from human sputum at the Acknowledgment for publications should read “The following University of Texas Health Science Center at Tyler, Tyler, reagent was obtained through BEI Resources, NIAID, NIH: 1 Texas, USA. Mycobacterium kansasii, Strain 824, NR-44269.” Comment: M. kansasii, strain 824 is part of the Top Priority Nontuberculous Mycobacteria Whole Genome Sequencing Biosafety Level: 2 Project at the Genomic Sequencing Center for Infectious Appropriate safety procedures should always be used with this Diseases (GSCID) at University of Maryland School of material. -

Genomics of Atlantic Forest Mycobacteriaceae Strains Unravels a Mobilome Diversity with a Novel Integrative Conjugative Element and Plasmids Harbouring T7SS
RESEARCH ARTICLE Morgado and Paulo Vicente, Microbial Genomics 2020;6 DOI 10.1099/mgen.0.000382 Genomics of Atlantic Forest Mycobacteriaceae strains unravels a mobilome diversity with a novel integrative conjugative element and plasmids harbouring T7SS Sergio Mascarenhas Morgado* and Ana Carolina Paulo Vicente Abstract Mobile genetic elements (MGEs) are agents of bacterial evolution and adaptation. Genome sequencing provides an unbiased approach that has revealed an abundance of MGEs in prokaryotes, mainly plasmids and integrative conjugative elements. Nev- ertheless, many mobilomes, particularly those from environmental bacteria, remain underexplored despite their representing a reservoir of genes that can later emerge in the clinic. Here, we explored the mobilome of the Mycobacteriaceae family, focus- ing on strains from Brazilian Atlantic Forest soil. Novel Mycolicibacterium and Mycobacteroides strains were identified, with the former ones harbouring linear and circular plasmids encoding the specialized type- VII secretion system (T7SS) and mobility- associated genes. In addition, we also identified a T4SS- mediated integrative conjugative element (ICEMyc226) encoding two T7SSs and a number of xenobiotic degrading genes. Our study uncovers the diversity of the Mycobacteriaceae mobilome, pro- viding the evidence of an ICE in this bacterial family. Moreover, the presence of T7SS genes in an ICE, as well as plasmids, highlights the role of these mobile genetic elements in the dispersion of T7SS. Data SUMMARY Conjugative elements, such as conjugative plasmids and Genomic data analysed in this work are available in GenBank integrative conjugative elements (ICEs), mediate their own and listed in Tables S1 and S2 (available in the online version transfer to other organisms, often carrying cargo genes. -

(Gid ) Genes Coding for Putative Trna:M5u-54 Methyltransferases in 355 Bacterial and Archaeal Complete Genomes
Table S1. Taxonomic distribution of the trmA and trmFO (gid ) genes coding for putative tRNA:m5U-54 methyltransferases in 355 bacterial and archaeal complete genomes. Asterisks indicate the presence and the number of putative genes found. Genomes Taxonomic position TrmA Gid Archaea Crenarchaea Aeropyrum pernix_K1 Crenarchaeota; Thermoprotei; Desulfurococcales; Desulfurococcaceae Cenarchaeum symbiosum Crenarchaeota; Thermoprotei; Cenarchaeales; Cenarchaeaceae Pyrobaculum aerophilum_str_IM2 Crenarchaeota; Thermoprotei; Thermoproteales; Thermoproteaceae Sulfolobus acidocaldarius_DSM_639 Crenarchaeota; Thermoprotei; Sulfolobales; Sulfolobaceae Sulfolobus solfataricus Crenarchaeota; Thermoprotei; Sulfolobales; Sulfolobaceae Sulfolobus tokodaii Crenarchaeota; Thermoprotei; Sulfolobales; Sulfolobaceae Euryarchaea Archaeoglobus fulgidus Euryarchaeota; Archaeoglobi; Archaeoglobales; Archaeoglobaceae Haloarcula marismortui_ATCC_43049 Euryarchaeota; Halobacteria; Halobacteriales; Halobacteriaceae; Haloarcula Halobacterium sp Euryarchaeota; Halobacteria; Halobacteriales; Halobacteriaceae; Haloarcula Haloquadratum walsbyi Euryarchaeota; Halobacteria; Halobacteriales; Halobacteriaceae; Haloquadra Methanobacterium thermoautotrophicum Euryarchaeota; Methanobacteria; Methanobacteriales; Methanobacteriaceae Methanococcoides burtonii_DSM_6242 Euryarchaeota; Methanomicrobia; Methanosarcinales; Methanosarcinaceae Methanococcus jannaschii Euryarchaeota; Methanococci; Methanococcales; Methanococcaceae Methanococcus maripaludis_S2 Euryarchaeota; Methanococci; -

Mycobacterium Avium Subsp
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Veterinary and Biomedical Sciences, Papers in Veterinary and Biomedical Science Department of July 2001 Mycobacterium avium subsp. paratuberculosis in Veterinary Medicine N. Beth Harris University of Nebraska - Lincoln Raul G. Barletta University of Nebraska - Lincoln, [email protected] Follow this and additional works at: https://digitalcommons.unl.edu/vetscipapers Part of the Veterinary Medicine Commons Harris, N. Beth and Barletta, Raul G., "Mycobacterium avium subsp. paratuberculosis in Veterinary Medicine" (2001). Papers in Veterinary and Biomedical Science. 5. https://digitalcommons.unl.edu/vetscipapers/5 This Article is brought to you for free and open access by the Veterinary and Biomedical Sciences, Department of at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Papers in Veterinary and Biomedical Science by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. CLINICAL MICROBIOLOGY REVIEWS, July 2001, p. 489–512 Vol. 14, No. 3 0893-8512/01/$04.00ϩ0 DOI: 10.1128/CMR.14.3.489–512.2001 Copyright © 2001, American Society for Microbiology. All Rights Reserved. Mycobacterium avium subsp. paratuberculosis in Veterinary Medicine N. BETH HARRIS AND RAU´ L G. BARLETTA* Department of Veterinary and Biomedical Sciences, University of Nebraska—Lincoln, Lincoln, Nebraska 68583-0905 INTRODUCTION .......................................................................................................................................................489 -

BEI Resources Product Information Sheet Catalog No. NR-44267
Product Information Sheet for NR-44267 Mycobacterium intracellulare, Strain 1956 Growth Conditions: Media: Catalog No. NR-44267 Middlebrook 7H9 broth with ADC enrichment or equivalent Middlebrook 7H10 agar with OADC enrichment or Lowenstein- Jensen agar or equivalent For research use only. Not for human use. Incubation: Temperature: 37°C Contributor: Atmosphere: Aerobic with 5% CO2 Diane Ordway, Ph.D., Assistant Professor, Department of Propagation: Microbiology, Immunology and Pathology, Colorado State 1. Keep vial frozen until ready for use; then thaw. University, Fort Collins, Colorado, USA and National Institute 2. Transfer the entire thawed aliquot into a single tube of of Allergy and Infectious Diseases (NIAID), National Institutes broth. of Health (NIH), Bethesda, Maryland, USA 3. Use several drops of the suspension to inoculate an agar slant and/or plate. Manufacturer: 4. Incubate the tube, slant and/or plate at 37°C for 2 to 8 BEI Resources weeks. Product Description: Citation: Bacteria Classification: Mycobacteriaceae, Mycobacterium Acknowledgment for publications should read “The following Species: Mycobacterium intracellulare reagent was obtained through BEI Resources, NIAID, NIH: Strain: 1956 Mycobacterium intracellulare, Strain 1956, NR-44267.” Original Source: Mycobacterium intracellulare (M. intracellulare), strain 1956 was isolated in 2011 from 1 Biosafety Level: 2 human sputum at NIAID, NIH, Bethesda, Maryland, USA. Appropriate safety procedures should always be used with this Comment: M. intracellulare, strain 1956 is part of the Top material. Laboratory safety is discussed in the following Priority Nontuberculous Mycobacteria Whole Genome publication: U.S. Department of Health and Human Services, Sequencing Project at the Genomic Sequencing Center for Public Health Service, Centers for Disease Control and Infectious Diseases (GSCID) at University of Maryland Prevention, and National Institutes of Health. -

Bibliography
Bibliography Abella, C.A., X.P. Cristina, A. Martinez, I. Pibernat and X. Vila. 1998. on moderate concentrations of acetate: production of single cells. Two new motile phototrophic consortia: "Chlorochromatium lunatum" Appl. Microbiol. Biotechnol. 35: 686-689. and "Pelochromatium selenoides". Arch. Microbiol. 169: 452-459. Ahring, B.K, P. Westermann and RA. Mah. 1991b. Hydrogen inhibition Abella, C.A and LJ. Garcia-Gil. 1992. Microbial ecology of planktonic of acetate metabolism and kinetics of hydrogen consumption by Me filamentous phototrophic bacteria in holomictic freshwater lakes. Hy thanosarcina thermophila TM-I. Arch. Microbiol. 157: 38-42. drobiologia 243-244: 79-86. Ainsworth, G.C. and P.H.A Sheath. 1962. Microbial Classification: Ap Acca, M., M. Bocchetta, E. Ceccarelli, R Creti, KO. Stetter and P. Cam pendix I. Symp. Soc. Gen. Microbiol. 12: 456-463. marano. 1994. Updating mass and composition of archaeal and bac Alam, M. and D. Oesterhelt. 1984. Morphology, function and isolation terial ribosomes. Archaeal-like features of ribosomes from the deep of halobacterial flagella. ]. Mol. Biol. 176: 459-476. branching bacterium Aquifex pyrophilus. Syst. Appl. Microbiol. 16: 629- Albertano, P. and L. Kovacik. 1994. Is the genus LeptolynglYya (Cyano 637. phyte) a homogeneous taxon? Arch. Hydrobiol. Suppl. 105: 37-51. Achenbach-Richter, L., R Gupta, KO. Stetter and C.R Woese. 1987. Were Aldrich, H.C., D.B. Beimborn and P. Schönheit. 1987. Creation of arti the original eubacteria thermophiles? Syst. Appl. Microbiol. 9: 34- factual internal membranes during fixation of Methanobacterium ther 39. moautotrophicum. Can.]. Microbiol. 33: 844-849. Adams, D.G., D. Ashworth and B. -

C 2011 by Daniel T. Wright. All Rights Reserved. INFORMATION and the EVOLUTION of CODON BIAS
c 2011 by Daniel T. Wright. All rights reserved. INFORMATION AND THE EVOLUTION OF CODON BIAS BY DANIEL T. WRIGHT DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Library and Information Science in the Graduate College of the University of Illinois at Urbana-Champaign, 2011 Urbana, Illinois Doctoral Committee: Professor Les Gasser, Chair Professor Allen Renear Professor Zan Luthey-Schulten Assistant Professor Miles Efron Abstract The informational properties of biological systems are the subject of much debate and re- search. I present a general argument in favor of the existence and central importance of information in organisms, followed by a case study of the genetic code (specifically, codon bias) and the translation system from the perspective of information. The codon biases of 831 Bacteria and Archeae are analyzed and modeled as points in a 64-dimensional statistical space. The major results are that (1) codon bias evolution does not follow canonical patterns, and (2) the use of coding space in organsims is a subset of the total possible coding space. These findings imply that codon bias is a unique adaptive mechanism that owes its exis- tence to organisms' use of information in representing genes, and that there is a particularly biological character to the resulting biased coding and information use. ii For my family, without whose love and support none of this would have been possible. Or even begun! iii Acknowledgments This project owes its existence to many people. First, I would like to express my deep thanks and appreciation to the faculty and staff of the Graduate School of Library and Information Science for many years of support and encouragement. -
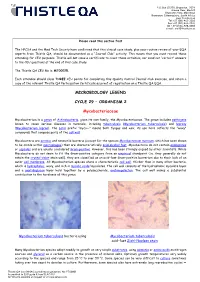
Mycobacteriaceae
P.O. Box 131375, Bryanston, 2074 Ground Floor, Block 5 Bryanston Gate, Main Road Bryanston, Johannesburg, South Africa www.thistle.co.za Tel: +27 (011) 463-3260 Fax: +27 (011) 463-3036 OR + 27 (0) 86-538-4484 e-mail : [email protected] Please read this section first The HPCSA and the Med Tech Society have confirmed that this clinical case study, plus your routine review of your EQA reports from Thistle QA, should be documented as a “Journal Club” activity. This means that you must record those attending for CEU purposes. Thistle will not issue a certificate to cover these activities, nor send out “correct” answers to the CEU questions at the end of this case study. The Thistle QA CEU No is: MT00025. Each attendee should claim THREE CEU points for completing this Quality Control Journal Club exercise, and retain a copy of the relevant Thistle QA Participation Certificate as proof of registration on a Thistle QA EQA. MICROBIOLOGY LEGEND CYCLE 29 – ORGANISM 2 Mycobacteriaceae Mycobacterium is a genus of Actinobacteria, given its own family, the Mycobacteriaceae. The genus includes pathogens known to cause serious diseases in mammals, including tuberculosis (Mycobacterium tuberculosis) and leprosy (Mycobacterium leprae). The Latin prefix "myco—" means both fungus and wax; its use here reflects the "waxy" compounds that compose parts of the cell wall. Mycobacteria are aerobic and nonmotile bacteria (except for the species Mycobacterium marinum, which has been shown to be motile within macrophages) that are characteristically acid-alcohol fast. Mycobacteria do not contain endospores or capsules and are usually considered Gram-positive. -

Workshop on Mycobacterium Avium Subsp
Workshop on Mycobacterium avium subsp. paratuberculosis in North American Bison (Bison bison) PROCEEDINGS AND WORKSHOP REPORT Alberta Cooperative Conservation Research Unit National (Canadian) Wood Bison (ACCRU) Recovery Team Editors: Murray Woodbury, Elena Garde, Helen Schwantje, John Nishi, and Brett Elkin with contributions from Michelle Oakley, Jenny Powers, Jennifer Sibley and workshop participants. Department of Resources, Wildlife and Economic Development Government of the Northwest Territories Yellowknife, NT 2006 Manuscript Report No. 170 The contents of this paper are the sole responsibility of the author ii iii ABSTRACT Many of the emerging diseases worldwide are shared between domestic livestock, free ranging wildlife, and humans. Infection by mycobacterial organisms is not a recent or newly emerging issue, but the increasing frequency of disease caused by Mycobacterium species, where wildlife, domestic livestock, and humans are epidemiologically linked is cause for concern for wildlife management, agricultural and public health agencies alike. Human tuberculosis, including that caused by Mycobacterium bovis, once thought to be a well understood disease of lifestyle and socioeconomic consequences is once again a global concern. Johne’s disease (Mycobacterium avium subspecies paratuberculosis infection), previously considered a production limiting disease important only in domestic ruminants, has recently been implicated in the pathogenesis of Crohn’s disease in humans. Furthermore, wildlife and livestock reservoirs of infection -

Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln US Department of Energy Publications U.S. Department of Energy 2010 Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota Gurdeep Rastogi South Dakota School of Mines and Technology Shariff Osman Lawrence Berkeley National Laboratory Ravi K. Kukkadapu Pacific Northwest National Laboratory, [email protected] Mark Engelhard Pacific Northwest National Laboratory Parag A. Vaishampayan California Institute of Technology See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/usdoepub Part of the Bioresource and Agricultural Engineering Commons Rastogi, Gurdeep; Osman, Shariff; Kukkadapu, Ravi K.; Engelhard, Mark; Vaishampayan, Parag A.; Andersen, Gary L.; and Sani, Rajesh K., "Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota" (2010). US Department of Energy Publications. 170. https://digitalcommons.unl.edu/usdoepub/170 This Article is brought to you for free and open access by the U.S. Department of Energy at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in US Department of Energy Publications by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Gurdeep Rastogi, Shariff Osman, Ravi K. Kukkadapu, Mark Engelhard, Parag A. Vaishampayan, Gary L. Andersen, and Rajesh K. Sani This article is available at DigitalCommons@University of Nebraska - Lincoln: https://digitalcommons.unl.edu/ usdoepub/170 Microb Ecol (2010) 60:539–550 DOI 10.1007/s00248-010-9657-y SOIL MICROBIOLOGY Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota Gurdeep Rastogi & Shariff Osman & Ravi Kukkadapu & Mark Engelhard & Parag A. -

Supplemental Tables for Plant-Derived Benzoxazinoids Act As Antibiotics and Shape Bacterial Communities
Supplemental Tables for Plant-derived benzoxazinoids act as antibiotics and shape bacterial communities Niklas Schandry, Katharina Jandrasits, Ruben Garrido-Oter, Claude Becker Contents Table S1. Syncom strains 2 Table S2. PERMANOVA 5 Table S3. ANOVA: observed taxa 6 Table S4. Observed diversity means and pairwise comparisons 7 Table S5. ANOVA: Shannon Diversity 9 Table S6. Shannon diversity means and pairwise comparisons 10 1 Table S1. Syncom strains Strain Genus Family Order Class Phylum Mixed Root70 Acidovorax Comamonadaceae Burkholderiales Betaproteobacteria Proteobacteria Root236 Aeromicrobium Nocardioidaceae Propionibacteriales Actinomycetia Actinobacteria Root100 Aminobacter Phyllobacteriaceae Rhizobiales Alphaproteobacteria Proteobacteria Root239 Bacillus Bacillaceae Bacillales Bacilli Firmicutes Root483D1 Bosea Bradyrhizobiaceae Rhizobiales Alphaproteobacteria Proteobacteria Root342 Caulobacter Caulobacteraceae Caulobacterales Alphaproteobacteria Proteobacteria Root137 Cellulomonas Cellulomonadaceae Actinomycetales Actinomycetia Actinobacteria Root1480D1 Duganella Oxalobacteraceae Burkholderiales Gammaproteobacteria Proteobacteria Root231 Ensifer Rhizobiaceae Rhizobiales Alphaproteobacteria Proteobacteria Root420 Flavobacterium Flavobacteriaceae Flavobacteriales Bacteroidia Bacteroidetes Root268 Hoeflea Phyllobacteriaceae Rhizobiales Alphaproteobacteria Proteobacteria Root209 Hydrogenophaga Comamonadaceae Burkholderiales Gammaproteobacteria Proteobacteria Root107 Kitasatospora Streptomycetaceae Streptomycetales Actinomycetia Actinobacteria