Oncoprotein AEG-1 Is an Endoplasmic Reticulum RNA Binding Protein Whose Interactome Is Enriched in Organelle Resident Protein-Encoding Mrnas
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Download on 20
bioRxiv preprint doi: https://doi.org/10.1101/850776; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Intramembrane protease RHBDL4 cleaves oligosaccharyltransferase subunits to target them for ER-associated degradation Julia D. Knopf1, Nina Landscheidt1, Cassandra L. Pegg2, Benjamin L. Schulz2, Nathalie Kühnle1, Chao-Wei Chao1, Simon Huck1 and Marius K. Lemberg1, # 1Centre for Molecular Biology of Heidelberg University (ZMBH), DKFZ-ZMBH Alliance, 69120 Heidelberg, Germany. 2School of Chemistry and Molecular Biosciences, ARC Training Centre for Biopharmaceutical Innovation, The University of Queensland, St Lucia QLD 4072, Australia. #Corresponding author: [email protected] Running title: RHBDL4 triggers ERAD of OST subunits Key words: Rhomboid serine protease, Rhbdd1, ubiquitin-dependent proteolysis, post- translational protein abundance control, N-linked glycosylation. Abbreviations ERAD, ER-associated degradation; OST, oligosacharyltransferase; TM, transmembrane; UIM, ubiquitin-interacting motif. Abstract The Endoplasmic Reticulum (ER)-resident intramembrane rhomboid protease RHBDL4 generates metastable protein fragments and together with the ER-associated degradation (ERAD) machinery provides a clearance mechanism for aberrant and surplus proteins. However, the endogenous substrate spectrum and with that the role of RHBDL4 in physiological ERAD is mainly unknown. Here, we use a substrate trapping approach in combination with quantitative proteomics to identify physiological RHBDL4 substrates. This revealed oligosacharyltransferase (OST) complex subunits such as the catalytic active subunit STT3A as substrates for the RHBDL4-dependent ERAD pathway. RHBDL4-catalyzed cleavage inactivates OST subunits by triggering dislocation into the cytoplasm and subsequent proteasomal degradation. -

S41467-020-18249-3.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-18249-3 OPEN Pharmacologically reversible zonation-dependent endothelial cell transcriptomic changes with neurodegenerative disease associations in the aged brain Lei Zhao1,2,17, Zhongqi Li 1,2,17, Joaquim S. L. Vong2,3,17, Xinyi Chen1,2, Hei-Ming Lai1,2,4,5,6, Leo Y. C. Yan1,2, Junzhe Huang1,2, Samuel K. H. Sy1,2,7, Xiaoyu Tian 8, Yu Huang 8, Ho Yin Edwin Chan5,9, Hon-Cheong So6,8, ✉ ✉ Wai-Lung Ng 10, Yamei Tang11, Wei-Jye Lin12,13, Vincent C. T. Mok1,5,6,14,15 &HoKo 1,2,4,5,6,8,14,16 1234567890():,; The molecular signatures of cells in the brain have been revealed in unprecedented detail, yet the ageing-associated genome-wide expression changes that may contribute to neurovas- cular dysfunction in neurodegenerative diseases remain elusive. Here, we report zonation- dependent transcriptomic changes in aged mouse brain endothelial cells (ECs), which pro- minently implicate altered immune/cytokine signaling in ECs of all vascular segments, and functional changes impacting the blood–brain barrier (BBB) and glucose/energy metabolism especially in capillary ECs (capECs). An overrepresentation of Alzheimer disease (AD) GWAS genes is evident among the human orthologs of the differentially expressed genes of aged capECs, while comparative analysis revealed a subset of concordantly downregulated, functionally important genes in human AD brains. Treatment with exenatide, a glucagon-like peptide-1 receptor agonist, strongly reverses aged mouse brain EC transcriptomic changes and BBB leakage, with associated attenuation of microglial priming. We thus revealed tran- scriptomic alterations underlying brain EC ageing that are complex yet pharmacologically reversible. -
Download Validation Data
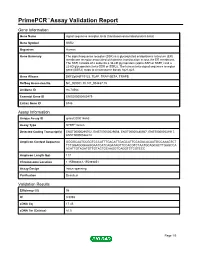
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

A Clearer Picture of the ER Translocon Complex Max Gemmer and Friedrich Förster*
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs231340. doi:10.1242/jcs.231340 REVIEW A clearer picture of the ER translocon complex Max Gemmer and Friedrich Förster* ABSTRACT et al., 1986). SP-equivalent N-terminal transmembrane helices that The endoplasmic reticulum (ER) translocon complex is the main gate are not cleaved off can also target proteins to the ER through the into the secretory pathway, facilitating the translocation of nascent same mechanism. In this SRP-dependent co-translational ER- peptides into the ER lumen or their integration into the lipid membrane. targeting mode, ribosomes associate with the ER membrane via ER Protein biogenesis in the ER involves additional processes, many of translocon complexes. These membrane protein complexes them occurring co-translationally while the nascent protein resides at translocate nascent soluble proteins into the ER, integrate nascent the translocon complex, including recruitment of ER-targeted membrane proteins into the ER membrane, mediate protein folding ribosome–nascent-chain complexes, glycosylation, signal peptide and membrane protein topogenesis, and modify them chemically. In cleavage, membrane protein topogenesis and folding. To perform addition to co-translational protein import and translocation, distinct such varied functions on a broad range of substrates, the ER ER translocon complexes enable post-translational translocation and translocon complex has different accessory components that membrane integration. This post-translational pathway is widespread associate with it either stably or transiently. Here, we review recent in yeast (Panzner et al., 1995), whereas higher eukaryotes primarily structural and functional insights into this dynamically constituted use it for relatively short peptides (Schlenstedt and Zimmermann, central hub in the ER and its components. -

Regulation of Hematopoietic Activity Involving New Interacting Partners (RRAGC & PSMC2, CKAP4 & MANF and CTR9 & CNTNAP2)
CellBio, 2020, 9, 123-141 https://www.scirp.org/journal/cellbio ISSN Online: 2325-7792 ISSN Print: 2325-7776 Regulation of Hematopoietic Activity Involving New Interacting Partners (RRAGC & PSMC2, CKAP4 & MANF and CTR9 & CNTNAP2) Swati Sharma1, Gurudutta U. Gangenahalli2*, Upma Singh3 1Department of Pharmacology, All India Institute of Medical Science (AIIMS), New Delhi, India 2Stem Cell & Gene Therapy Research Group, Institute of Nuclear Medicine & Allied Sciences, New Delhi, India 3Department of Applied Chemistry, School of Vocational Studies & Applied Sciences, Gautam Buddha University, Greater Noida, India How to cite this paper: Sharma, S., Gan- Abstract genahalli, G.U. and Singh, U. (2020) Regu- lation of Hematopoietic Activity Involving Hematopoietic stem cells (HSCs) are tissue-specific cells giving rise to all New Interacting Partners (RRAGC & PSMC2, mature blood cell types regulated by a diverse group of hematopoietic cyto- CKAP4 & MANF and CTR9 & CNTNAP2). kines and growth factors that influences the survival & proliferation of early CellBio, 9, 123-141. https://doi.org/10.4236/cellbio.2020.93007 progenitors and differentiation mechanisms by modulating the functional ac- tivities of HSCs. In this study, the functional yet distinctive role of three novel Received: August 13, 2020 combinations of gene pairs RRAGC & PSMC2; CKAP4 & MANF; and CTR9 Accepted: September 27, 2020 & CNTNAP2 have been newly identified. These novel combinations of genes Published: September 30, 2020 were confirmed and expressed in K562 human leukemic cell line in the pres- Copyright © 2020 by author(s) and ence of cytokine combination (IL-3, FLT-3 and SCF) using RT-PCR and Scientific Research Publishing Inc. siRNA-mediated gene knock down strategy. -

Cytoskeleton-Associated Protein 4: Functions Beyond the Endoplasmic Reticulum in Physiology and Disease
International Scholarly Research Network ISRN Cell Biology Volume 2012, Article ID 142313, 11 pages doi:10.5402/2012/142313 Review Article Cytoskeleton-Associated Protein 4: Functions Beyond the Endoplasmic Reticulum in Physiology and Disease Kevin M. Tuffy and Sonia Lobo Planey Department of Basic Sciences, The Commonwealth Medical College, Scranton, PA 18509, USA Correspondence should be addressed to Sonia Lobo Planey, [email protected] Received 21 August 2012; Accepted 19 September 2012 Academic Editors: A. Hergovich and C. M. Wells Copyright © 2012 K. M. Tuffy and S. L. Planey. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Cytoskeleton-associated protein 4 (CKAP4; also known as p63, CLIMP-63, or ERGIC-63) is a 63 kDa, reversibly palmitoylated and phosphorylated, type II transmembrane (TM) protein, originally identified as a resident of the endoplasmic reticulum (ER)/Golgi intermediate compartment (ERGIC). When localized to the ER, a major function of CKAP4 is to anchor rough ER to microtubules, organizing the overall structure of ER with respect to the microtubule network. There is also steadily accumulating evidence for diverse roles for CKAP4 localized outside the ER, including data demonstrating functionality of cell surface forms of CKAP4 in various cell types and of CKAP4 in the nucleus. We will review the recent studies that provide evidence for the existence of CKAP4 in multiple cellular compartments (i.e., ER, plasma membrane, and the nucleus) and discuss CKAP4’s role in the regulation of various physiological and pathological processes, such as interstitial cystitis, drug-induced cytotoxicity, pericullar proteolytic activity, and lung lipid homeostasis. -

Common Variant Near the Endothelin Receptor Type a (EDNRA)
Common variant near the endothelin receptor type A(EDNRA) gene is associated with intracranial aneurysm risk Katsuhito Yasunoa,b,c, Mehmet Bakırcıog˘ lua,b,c, Siew-Kee Lowd, Kaya Bilgüvara,b,c, Emília Gaála,b,c,e, Ynte M. Ruigrokf, Mika Niemeläe, Akira Hatag, Philippe Bijlengah, Hidetoshi Kasuyai, Juha E. Jääskeläinenj, Dietmar Krexk, Georg Auburgerl, Matthias Simonm, Boris Krischekn, Ali K. Ozturka,b,c, Shrikant Maneo, Gabriel J. E. Rinkelf, Helmuth Steinmetzl, Juha Hernesniemie, Karl Schallerh, Hitoshi Zembutsud, Ituro Inouep, Aarno Palotieq, François Cambienr, Yusuke Nakamurad, Richard P. Liftonc,s,t,1, and Murat Günela,b,c,1 Departments of aNeurosurgery, bNeurobiology, and cGenetics, Yale Program on Neurogenetics, Yale Center for Human Genetics and Genomics, and sDepartment of Internal Medicine and tHoward Hughes Medical Institute, Yale University School of Medicine, New Haven, CT 06510; dHuman Genome Center, Institute of Medical Science, University of Tokyo, Tokyo 108-8639, Japan; eDepartment of Neurosurgery, Helsinki University Central Hospital, Helsinki, FI-00029 HUS, Finland; fDepartment of Neurology, Rudolf Magnus Institute of Neuroscience, University Medical Center Utrecht, 3584 CX Utrecht, The Netherlands; gDepartment of Public Health, School of Medicine, Chiba University, Chiba 260-8670, Japan; hService de Neurochirurgie, Department of Clinical Neurosciences, Geneva University Hospital, 1211 Geneva 4, Switzerland; iDepartment of Neurosurgery, Medical Center East, Tokyo Women’s University, Tokyo 116-8567, Japan; jDepartment -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

A Crosstalk Between the RNA Binding Protein Smaug and the Hedgehog Pathway Links Cell Signaling to Mrna Regulation in Drosophila Lucía Bruzzone
A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila Lucía Bruzzone To cite this version: Lucía Bruzzone. A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila. Cellular Biology. Université Sorbonne Paris Cité, 2018. English. NNT : 2018USPCC234. tel-02899776 HAL Id: tel-02899776 https://tel.archives-ouvertes.fr/tel-02899776 Submitted on 15 Jul 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparée à l’Université Paris Diderot Ecole doctorale HOB n° 561 Institut Jacques Monod / Equipe Développement, Signalisation et Trafic A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in Drosophila Lucía Bruzzone Thèse de doctorat de Biologie Dirigée par Anne Plessis Présentée et soutenue publiquement à Paris le 19 mars 2018 Président du jury: Alain Zider / Professeur Université Paris Diderot -

Glycan Engineering for Cell and Developmental Biology
View metadata, citation and similar papers at core.ac.uk brought to you by CORE HHS Public Access provided by Caltech Authors - Main Author manuscript Author ManuscriptAuthor Manuscript Author Cell Chem Manuscript Author Biol. Author Manuscript Author manuscript; available in PMC 2016 May 05. Published in final edited form as: Cell Chem Biol. 2016 January 21; 23(1): 108–121. doi:10.1016/j.chembiol.2015.12.007. Glycan Engineering for Cell and Developmental Biology Matthew E. Griffin1 and Linda C. Hsieh-Wilson1,* 1Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125, USA Abstract Cell-surface glycans are a diverse class of macromolecules that participate in many key biological processes, including cell-cell communication, development, and disease progression. Thus, the ability to modulate the structures of glycans on cell surfaces provides a powerful means not only to understand fundamental processes but also to direct activity and elicit desired cellular responses. Here, we describe methods to sculpt glycans on cell surfaces and highlight recent successes in which artificially engineered glycans have been employed to control biological outcomes such as the immune response and stem cell fate. Introduction Cell-surface glycans participate in many important processes throughout the lifespan of an organism, ranging from cell migration and tissue patterning to the immune response, disease progression, and cell death (Fuster and Esko, 2005; Häcker et al., 2005; Lichtenstein and Rabinovich, 2013; van Kooyk and Rabinovich, 2008). At a molecular level, glycans are often the first points of contact between cells, and they function by facilitating a variety of interactions both in cis (on the same cell) and in trans (on different cells). -

The Roles of Zdhhcs in Cancer and As Potential Targets
S3181561: William Suelmann 1-7-2019 Supervisor:Lotteke Ziel-Swier The roles of zDHHCs in cancer and as potential targets Abstract: zDHHCs are proteins that are involved in multiple pathways in the cell. Palmitoylation of proteins happens when forming a thioester bond of palmitoyl-CoA and cysteine residue in DHHC-CR domain. There are multiple zDHHCs that are involved in multiple cancers, both as oncogenic and tumor suppressor. However, for targeting zDHHCs as potential targets in cancer there is more research needed. Introduction: Discovery and mechanism of zDHHCs The process of palmitoylation was first described in 1951[10], however the enzymes that catalyze palmitoylation are a more recent discovery. The first major breakthrough in finding this enzyme came through an experiment on yeast, also known as Saccharomyces cerevisiae. An Erf2p-Erf4p complex had been identified to be involved in localising and palmitoylation of Ras2 protein towards the plasma membrane. This complex had shown to increase Ras2 palmitoylation[1], where deletion of Erf2 or Erf4 would lead to either mislocalization of the Ras protein or a decrease in palmitoylation[2]. Another protein, identified as Akr1p(ankyrin-repeat-containing protein 1) was also identified in yeast, as it was shown that Akr1p mediated the palmitoylation of Yck2 (yeast casein kinase 2)[3]. Interestingly, Erf2 and Akr1p both were integral membrane proteins that contained the aspartate-histidine-histidine-cysteine (DHHC) cysteine-rich (CR) conserved domain and multiple transmembrane domains, where the DHHC domain was located between the second and third transmembrane domains[4][5]. zDHHC protein was acknowledged as a palmitoyltransferase through these studies in yeast, which showed how important the DHHC-CR domain is for activity within the cell. -

RRBP1 (NM 001042576) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC218816L3 RRBP1 (NM_001042576) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: RRBP1 (NM_001042576) Human Tagged ORF Clone Tag: Myc-DDK Symbol: RRBP1 Synonyms: ES/130; ES130; hES; p180; RRp Vector: pLenti-C-Myc-DDK-P2A-Puro (PS100092) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(RC218816). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_001042576 ORF Size: 2931 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 RRBP1 (NM_001042576) Human Tagged ORF Clone – RC218816L3 OTI Disclaimer: Due to the inherent nature of this plasmid, standard methods to replicate additional amounts of DNA in E. coli are highly likely to result in mutations and/or rearrangements. Therefore, OriGene does not guarantee the capability to replicate this plasmid DNA. Additional amounts of DNA can be purchased from OriGene with batch-specific, full-sequence verification at a reduced cost. Please contact our customer care team at [email protected] or by calling 301.340.3188 option 3 for pricing and delivery. The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g.