The Novel Isoxazoline Ectoparasiticide Fluralaner
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Evaluation of Fluralaner and Afoxolaner Treatments to Control Flea
Dryden et al. Parasites & Vectors (2016) 9:365 DOI 10.1186/s13071-016-1654-7 RESEARCH Open Access Evaluation of fluralaner and afoxolaner treatments to control flea populations, reduce pruritus and minimize dermatologic lesions in naturally infested dogs in private residences in west central Florida USA Michael W. Dryden1*, Michael S. Canfield2, Kimberly Kalosy1, Amber Smith1, Lisa Crevoiserat1, Jennifer C. McGrady1, Kaitlin M. Foley1, Kathryn Green2, Chantelle Tebaldi2, Vicki Smith1, Tashina Bennett1, Kathleen Heaney3, Lisa Math3, Christine Royal3 and Fangshi Sun3 Abstract Background: A study was conducted to evaluate and compare the effectiveness of two different oral flea and tick products to control flea infestations, reduce pruritus and minimize dermatologic lesions over a 12 week period on naturally infested dogs in west central FL USA. Methods: Thirty-four dogs with natural flea infestations living in 17 homes were treated once with a fluralaner chew on study day 0. Another 27 dogs living in 17 different homes were treated orally with an afoxolaner chewable on day 0, once between days 28–30 and once again between days 54–60. All products were administered according to label directions by study investigators. Flea populations on pets were assessed using visual area counts and premise flea infestations were assessed using intermittent-light flea traps on days 0, 7, 14, 21, and once between days 28–30, 40–45, 54–60 and 82–86. Dermatologic assessments were conducted on day 0 and once monthly. Pruritus assessments were conducted by owners throughout the study. No concurrent treatments for existing skin disease (antibiotics, anti-inflammatories, anti-fungals) were allowed. -

B Commission Regulation (Eu)
02010R0037 — EN — 29.09.2018 — 035.001 — 1 This text is meant purely as a documentation tool and has no legal effect. The Union's institutions do not assume any liability for its contents. The authentic versions of the relevant acts, including their preambles, are those published in the Official Journal of the European Union and available in EUR-Lex. Those official texts are directly accessible through the links embedded in this document ►B COMMISSION REGULATION (EU) No 37/2010 of 22 December 2009 on pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin (Text with EEA relevance) (OJ L 15, 20.1.2010, p. 1) Amended by: Official Journal No page date ►M1 Commission Regulation (EU) No 758/2010 of 24 August 2010 L 223 37 25.8.2010 ►M2 Commission Regulation (EU) No 759/2010 of 24 August 2010 L 223 39 25.8.2010 ►M3 Commission Regulation (EU) No 761/2010 of 25 August 2010 L 224 1 26.8.2010 ►M4 Commission Regulation (EU) No 890/2010 of 8 October 2010 L 266 1 9.10.2010 ►M5 Commission Regulation (EU) No 914/2010 of 12 October 2010 L 269 5 13.10.2010 ►M6 Commission Regulation (EU) No 362/2011 of 13 April 2011 L 100 26 14.4.2011 ►M7 Commission Regulation (EU) No 363/2011 of 13 April 2011 L 100 28 14.4.2011 ►M8 Commission Implementing Regulation (EU) No 84/2012 of 1 L 30 1 2.2.2012 February 2012 ►M9 Commission Implementing Regulation (EU) No 85/2012 of 1 L 30 4 2.2.2012 February 2012 ►M10 Commission Implementing Regulation (EU) No 86/2012 of 1 L 30 6 2.2.2012 February 2012 ►M11 Commission -

Lindane Lotion USP, 1% RX Only WARNINGS
Lindane Lotion USP, 1% RX Only WARNINGS: Lindane Lotion should only be used in patients who cannot tolerate or have failed first-line treatment with safer medications for the treatment of scabies. (See INDICATIONS AND USAGE.) Neurologic Toxicity Seizures and deaths have been reported following Lindane Lotion use with repeat or prolonged application, but also in rare cases following a single application used according to directions. Lindane lotion should be used with caution for infants, children, the elderly, and individuals with other skin conditions (e.g, atopic dermatitis, psoriasis) and in those who weigh < 110 lbs (50 kg) as they may be at risk of serious neurotoxicity. Contraindications Lindane Lotion is contraindicated in premature infants and individuals with known uncontrolled seizure disorders. Proper Use Instruct patients on the proper use of Lindane Lotion, the amount to apply, how long to leave it on, and avoiding re-treatment. Inform patients that itching occurs after the successful killing of scabies and is not necessarily an indication for re-treatment with Lindane Lotion. (See DOSAGE AND ADMINISTRATION.) DESCRIPTION Lindane Lotion USP, 1%, is an ectoparasiticide and ovicide. In addition to the active ingredient, lindane, it contains glycerol monostearate, cetyl alcohol, stearic acid, trolamine, carrageenan, 2- amino-2-methyl-1-propanol, methylparaben, butylparaben, perfume and water to form a lotion. Lindane is the gamma isomer of 1,2,3,4,5,6-hexachlorocyclohexane having the following structural formula: C6H6Cl6 M.W. 290.83 CLINICAL PHARMACOLOGY Lindane Lotion USP, 1%, is an ectoparasiticide and ovicide effective against Sarcoptes scabiei (scabies). Lindane exerts its parasiticidal action by being directly absorbed into the parasites and 1 their ova. -

Revised Use-Function Classification (2007)
INTERNATIONAL PROGRAMME ON CHEMICAL SAFETY IPCS INTOX Data Management System (INTOX DMS) Revised Use-Function Classification (2007) The Use-Function Classification is used in two places in the INTOX Data Management System: the Communication Record and the Agent/Product Record. The two records are linked: if there is an agent record for a Centre Agent that is the subject of a call, the appropriate Intended Use-Function can be selected automatically in the Communication Record. The Use-Function Classification is used when generating reports, both standard and customized, and for searching the case and agent databases. In particular, INTOX standard reports use the top level headings of the Intended Use-Functions that were selected for Centre Agents in the Communication Record (e.g. if an agent was classified as an Analgesic for Human Use in the Communication Record, it would be logged as a Pharmaceutical for Human Use in the report). The Use-Function classification is very important for ensuring harmonized data collection. In version 4.4 of the software, 5 new additions were made to the top levels of the classification provided with the system for the classification of organisms (items XIV to XVIII). This is a 'convenience' classification to facilitate searching of the Communications database. A taxonomic classification for organisms is provided within the INTOX DMS Agent Explorer. In May/June 2006 INTOX users were surveyed to find out whether they had made any changes to the Use-Function Classification. These changes were then discussed at the 4th and 5th Meetings of INTOX Users. Version 4.5 of the INTOX DMS includes the revised pesticides classification (shown in full below). -

Identification of Transcriptome and Fluralaner Responsive Genes in The
Jia et al. BMC Genomics (2020) 21:120 https://doi.org/10.1186/s12864-020-6533-0 RESEARCH ARTICLE Open Access Identification of transcriptome and fluralaner responsive genes in the common cutworm Spodoptera litura Fabricius, based on RNA-seq Zhong-Qiang Jia1, Di Liu1, Ying-Chuan Peng1,3, Zhao-Jun Han1, Chun-Qing Zhao1* and Tao Tang2* Abstract Background: Fluralaner is a novel isoxazoline insecticide with a unique action site on the γ-aminobutyric acid receptor (GABAR), shows excellent activity on agricultural pests including the common cutworm Spodoptera litura, and significantly influences the development and fecundity of S. litura at either lethal or sublethal doses. Herein, Illumina HiSeq Xten (IHX) platform was used to explore the transcriptome of S. litura and to identify genes responding to fluralaner exposure. Results: A total of 16,572 genes, including 451 newly identified genes, were observed in the S. litura transcriptome and annotated according to the COG, GO, KEGG and NR databases. These genes included 156 detoxification enzyme genes [107 cytochrome P450 enzymes (P450s), 30 glutathione S-transferases (GSTs) and 19 carboxylesterases (CarEs)] and 24 insecticide-targeted genes [5 ionotropic GABARs, 1 glutamate-gated chloride channel (GluCl), 2 voltage-gated sodium channels (VGSCs), 13 nicotinic acetylcholine receptors (nAChRs), 2 acetylcholinesterases (AChEs) and 1 ryanodine receptor (RyR)]. There were 3275 and 2491 differentially expressed genes (DEGs) in S. litura treated with LC30 or LC50 concentrations of fluralaner, respectively. Among the DEGs, 20 related to detoxification [16 P450s, 1 GST and 3 CarEs] and 5 were growth-related genes (1 chitin and 4 juvenile hormone synthesis genes). -

Evaluation of Fluralaner As an Oral Acaricide to Reduce Tick Infestation
Pelletier et al. Parasites Vectors (2020) 13:73 https://doi.org/10.1186/s13071-020-3932-7 Parasites & Vectors RESEARCH Open Access Evaluation of furalaner as an oral acaricide to reduce tick infestation in a wild rodent reservoir of Lyme disease Jérôme Pelletier1,2,3*, Jean‑Philippe Rocheleau2,4, Cécile Aenishaenslin1,2,3, Francis Beaudry5, Gabrielle Dimitri Masson1,2, L. Robbin Lindsay6, Nicholas H. Ogden2,7, Catherine Bouchard2,7 and Patrick A. Leighton1,2,3 Abstract Background: Lyme disease (LD) is an increasing public health threat in temperate zones of the northern hemisphere, yet relatively few methods exist for reducing LD risk in endemic areas. Disrupting the LD transmission cycle in nature is a promising avenue for risk reduction. This experimental study evaluated the efcacy of furalaner, a recent oral aca‑ ricide with a long duration of efect in dogs, for killing Ixodes scapularis ticks in Peromyscus maniculatus mice, a known wildlife reservoir for Borrelia burgdorferi in nature. Methods: We assigned 87 mice to 3 furalaner treatment groups (50 mg/kg, 12.5 mg/kg and untreated control) administered as a single oral treatment. Mice were then infested with 20 Ixodes scapularis larvae at 2, 28 and 45 days post‑treatment and we measured efcacy as the proportion of infesting larvae that died within 48 h. At each infesta‑ tion, blood from 3 mice in each treatment group was tested to obtain furalaner plasma concentrations (C p). Results: Treatment with 50 mg/kg and 12.5 mg/kg furalaner killed 97% and 94% of infesting larvae 2 days post‑treat‑ ment, but no signifcant efect of treatment on feeding larvae was observed 28 and 45 days post‑treatment. -

Antibiotics May Eradicate Gastrointestinal Immunodeficiency.8
1122 Letters to the Editor J Neurol Neurosurg Psychiatry: first published as 10.1136/jnnp.54.12.1122 on 1 December 1991. Downloaded from sitivity.? We present a case of OMM without pseudobulbar palsy. The association of vitro.8 Whipple's disease is associated with symptoms of Whipple's disease and with supranuclear gaze paresis with oculomas- immunodeficiency.8 Thus intestinal wall negative peroral jejunoileal biopsies, which ticatory myorhythmia, however, seems to be macrophages are ineffective in phagocytosing indicates the usefulness of laparotomy for pathognomonic of the effects of Whipple's intracellular gram positive bacilli, resulting in jejunoileal biopsies as an alternative to brain disease on the CNS.'2 This led us to perform inability to eliminate chronic infection.9 This biopsy to confirm Whipple's disease. surgical jejunal and mesenteric lymph node suggests that Whipple's disease may be con- A 47 year old woman was admitted in May biopsies rather than a brain biopsy despite sidered as a disease of macrophages.8 The 1988 in a depressed state. In June 1987 she negative endoscopic and numerous peroral periventricular and periaqueductal distribu- had noted progressive visual disturbance. distal jejunal and ileal biopsies. The nor- tion of the CNS involvement in Whipple's Rhythmic elevations of her right upper lip mality ofthe CT and MRI scans also suppor- disease consists of macrophagic infiltration appeared, and later, paroxysmal hypersomnia ted this decision. and subependymal nodules. Such a and considerable weight gain (10 kg). A nine The possibility of brain involvement with- "tumoural" involvement may explain why month course oftreatment for depression was out systemic manifestation in Whipple's dis- antibiotics with good BBB diffusion are not ineffective and her symptoms and signs ease should be kept in mind. -
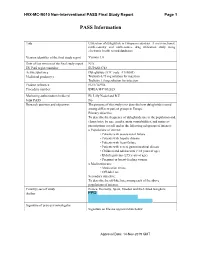
PASS Information
H9X-MC-B010 Non-interventional PASS Final Study Report Page 1 PASS Information Title Utilisation of dulaglutide in European countries: A cross-sectional, multi-country and multi-source drug utilisation study using electronic health record databases Version identifier of the final study report Version 1.0 Date of last version of the final study report N/A EU PAS register number EUPAS13783 Active substance Dulaglutide (ATC code: A10BJ05) Medicinal product(s): Trulicity 0.75 mg solution for injection Trulicity 1.5 mg solution for injection Product reference: EU/1/14/956 Procedure number: EMEA/H/C/002825 Marketing authorisation holder(s) Eli Lilly Nederland B.V. Joint PASS No Research question and objectives The purpose of this study is to describe how dulaglutide is used among different patient groups in Europe. Primary objective: To describe the frequency of dulaglutide use in the population and characterise by age, gender, main comorbidities, and main co- prescriptions overall and in the following subgroups of interest: o Populations of interest: • Patients with severe renal failure • Patients with hepatic disease • Patients with heart failure • Patients with severe gastrointestinal disease • Children and adolescents (<18 years of age) • Elderly patients (≥75 years of age) • Pregnant or breast-feeding women o Medication use: • Medication errors • Off-label use Secondary objective: To describe the off-label use among each of the above populations of interest. Country(-ies) of study France, Germany, Spain, Sweden and the United Kingdom Author PPD Signature of principal investigator Signature on file/see approval date below Approval Date: 14-Nov-2019 GMT H9X-MC-B010 Non-interventional PASS Final Study Report Page 2 Marketing Authorisation Holder Marketing authorisation holder (MAH) Eli Lilly Nederland B.V. -

A Saprolegnia Parasitica Challenge System for Rainbow Trout: Assessment of Pyceze As an Anti-Fungal Agent for Both Fish and Ova
DISEASES OF AQUATIC ORGANISMS Published May 12 Dis Aquat Org l A Saprolegnia parasitica challenge system for rainbow trout: assessment of Pyceze as an anti-fungal agent for both fish and ova T. G.Pottinger*, J. G. Day NERC Institute of Freshwater Ecology, Windermere Laboratory, Far Sawrey, Ambleside, Cumbria, LA22 OLP, United Kingdom ABSTRACT: A reproducible Saprolegnia parasitica spore delivery system was developed and demon- strated to be effective in providing a sustained spore challenge for up to 10 d. Treatment of rainbow trout with slow-release intraperitoneal implants containing cortisol resulted in chronically elevated blood cortisol levels and rendered the fish susceptible to infection by S. parasitica when exposed to the spore challenge. Sham-implanted fish were not susceptible to infect~on.Bronopol (2-bromo-2-nitro- propane-1,3-dol), formulated as Pyceze, was effective in protecting predisposed fish from infection by S. parasitica when administered as a daily bathlflush treatment at concentrations of 15 mg I-' and greater. Pyceze was also demonstrated to protect fertilised rainbow trout ova from S. parasitica chal- lenge when administered as a daily bath/flush treatment at concentrations of between 30 and 100 mg 1-l. Pyceze appears to qualify as a safe and effective replacement for malachite green and formalin in the prevention of fungal infections in the aquaculture environment. KEY WORDS: Saprolegnia . Fungal infection . Salmonid . Bronopol - Pyceze . Cortisol INTRODUCTION chite green in combating mycotic infections of fish and fish eggs, but is safer for the operator, the fish, and Mycotic infections of farmed fish, primarily by water the environment. Among the alternative compounds moulds or pseudofungi of the genus Saprolegnia, rep- tested for antifungal activity are sodium chloride resent a significant economic and welfare problem. -

A Single Amino Acid Substitution in the Third Transmembrane Region Has Opposite Impacts on the Selectivity of the Parasiticides
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2017/09/08/mol.117.109413.DC1 1521-0111/92/5/546–555$25.00 https://doi.org/10.1124/mol.117.109413 MOLECULAR PHARMACOLOGY Mol Pharmacol 92:546–555, November 2017 Copyright ª 2017 by The American Society for Pharmacology and Experimental Therapeutics A Single Amino Acid Substitution in the Third Transmembrane Region Has Opposite Impacts on the Selectivity of the Parasiticides Fluralaner and Ivermectin for Ligand-Gated Chloride Channels s Yunosuke Nakata, Toshinori Fuse, Kohei Yamato, Miho Asahi, Kunimitsu Nakahira, Fumiyo Ozoe, and Yoshihisa Ozoe Faculty of Life and Environmental Science, Shimane University, Matsue, Shimane, Japan (Y.N., T.F., K.Y, F.O., Y.O.); and Downloaded from Biological Research Laboratories, Nissan Chemical Industries, Ltd., Saitama, Japan (M.A., K.N.) Received May 16, 2017; accepted September 9, 2017 ABSTRACT Fluralaner (Bravecto) is a recently marketed isoxazoline ectopar- transmembrane region (TM3) with an aromatic amino acid molpharm.aspetjournals.org asiticide. This compound potently inhibits GABA-gated chloride dramatically enhanced the potency of fluralaner in the GluCls. channels (GABACls) and less potently glutamate-gated chloride In stark contrast to the enhancement of fluralaner potency, this channels (GluCls) in insects. The mechanism underlying this mutation eliminated the activation of currents and the potenti- selectivity is unknown. Therefore, we sought to identify the ation but not the antagonism of glutamate responses that are amino acid residues causing the low potency of fluralaner toward otherwise all elicited by the macrolide parasiticide ivermectin GluCls. We examined the fluralaner sensitivity of mutant housefly (IVM). -

Study Protocol
STUDY PROTOCOL FOR COMPASSIONATE AQUACULTURE INVESTIGATIONAL NEW ANIMAL DRUG (INAD) EXEMPTION FOR 35% PEROX-AID® (HYDROGEN PEROXIDE) FOR THE CONTROL OF ECTOPARASITES (INAD #11-669) Sponsor: U.S. Fish and Wildlife Service, Division of the National Fish Hatchery System ______________________ ___________________ Sponsor Signature Date Approved Manufacturer: Eka Chemicals Inc. 1775 West Oak Commons Court Marietta, Georgia 30062-2254 Facility for Coordination of 35% PEROX AID® INAD: Aquatic Animal Drug Approval Partnership Program 4050 Bridger Canyon Road Bozeman, Mt 59715 Proposed Starting Date: December 1, 2007 Proposed Ending Date: November 30, 2012 Study Director: Mr. Jim Bowker _________________________ ________________ Study Director Signature Date Clinical Field Trial Location and Trial Number: Facility ______________________________________________________________ Type or Print Name Investigator____________________________________________________________ Type or Print Name __________________________________________ ________________________ Investigator Signature Date 1 I. STUDY ID AND TITLE 3 II. SPONSOR 3 III. INVESTIGATORS/FACILITIES 4 IV. PROPOSED STARTING AND COMPLETION DATES: 4 V. BACKGROUND/PURPOSE 4 VI. SPECIFIC OBJECTIVES 6 VII. MATERIALS 7 VIII. EXPERIMENTAL UNIT 10 IX. ENTRANCE CRITERIA 10 X. TREATMENT GROUPS 11 XI. TREATMENT SCHEDULES 12 XII. TREATMENT RESPONSE PARAMETERS 15 XIII. FORMS FOR DATA COLLECTION 15 XIV. RECORD KEEPING PROCEDURES 16 XV. DISPOSITION OF INVESTIGATIONAL ANIMALS 16 XVI. DISPOSITION OF INVESTIGATIONAL -

Cover Memo Isoxazolines Inquires
Name: Isoxazoline inquiries DATE:10/1/2018 This serves as the response to your Freedom of Information Act (FOIA) request for records regarding adverse event reports received for afoxolaner, fluralaner, lotilaner and sarolaner. A search of CVM’s Adverse Drug Event (ADE) database was performed on 10-01-2018. The search parameters were: Active ingredient(s): afoxolaner, fluralaner, lotilaner and sarolaner Reports received: From 09-04-2013 through 07-31-2018 Case type: Spontaneous ADE report Species: All Route of administration: All For each drug (active ingredient), we have provided the ‘CVM ADE Comprehensive Clinical Detail Report Listing’, which is a cumulative listing of adverse experiences in reports submitted to CVM. General Information about CVM’s ADE Database The primary purpose for maintaining the CVM ADE database is to provide an early warning or signaling system to CVM for adverse effects not detected during pre-market testing of FDA- approved animal drugs and for monitoring the performance of drugs not approved for use in animals. Information from these ADE reports is received and coded in an electronic FDA/CVM ADE database. CVM scientists use the ADE database to make decisions about product safety which may include changes to the label or other regulatory action. CVM’s ADE reporting system depends on detection and voluntary reporting of adverse clinical events by veterinarians and animal owners. The Center's ADE review process is complex, and for each report takes into consideration confounding factors such as: Dosage Concomitant drug use The medical and physical condition of animals at the time of treatment Environmental and management information Product defects Name: Isoxazoline inquiries DATE:10/1/2018 Extra-label (off label) uses The specifics of these complex factors cannot be addressed in the CVM ADE Comprehensive Clinical Detail Report Listing.