The Role of the NFAT Signalling Pathway in Diffuse Large B-Cell Lymphoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Metabolic Dysfunction Is Restricted to the Sciatic Nerve in Experimental
Page 1 of 255 Diabetes Metabolic dysfunction is restricted to the sciatic nerve in experimental diabetic neuropathy Oliver J. Freeman1,2, Richard D. Unwin2,3, Andrew W. Dowsey2,3, Paul Begley2,3, Sumia Ali1, Katherine A. Hollywood2,3, Nitin Rustogi2,3, Rasmus S. Petersen1, Warwick B. Dunn2,3†, Garth J.S. Cooper2,3,4,5* & Natalie J. Gardiner1* 1 Faculty of Life Sciences, University of Manchester, UK 2 Centre for Advanced Discovery and Experimental Therapeutics (CADET), Central Manchester University Hospitals NHS Foundation Trust, Manchester Academic Health Sciences Centre, Manchester, UK 3 Centre for Endocrinology and Diabetes, Institute of Human Development, Faculty of Medical and Human Sciences, University of Manchester, UK 4 School of Biological Sciences, University of Auckland, New Zealand 5 Department of Pharmacology, Medical Sciences Division, University of Oxford, UK † Present address: School of Biosciences, University of Birmingham, UK *Joint corresponding authors: Natalie J. Gardiner and Garth J.S. Cooper Email: [email protected]; [email protected] Address: University of Manchester, AV Hill Building, Oxford Road, Manchester, M13 9PT, United Kingdom Telephone: +44 161 275 5768; +44 161 701 0240 Word count: 4,490 Number of tables: 1, Number of figures: 6 Running title: Metabolic dysfunction in diabetic neuropathy 1 Diabetes Publish Ahead of Print, published online October 15, 2015 Diabetes Page 2 of 255 Abstract High glucose levels in the peripheral nervous system (PNS) have been implicated in the pathogenesis of diabetic neuropathy (DN). However our understanding of the molecular mechanisms which cause the marked distal pathology is incomplete. Here we performed a comprehensive, system-wide analysis of the PNS of a rodent model of DN. -

Liver Glucose Metabolism in Humans
Biosci. Rep. (2016) / 36 / art:e00416 / doi 10.1042/BSR20160385 Liver glucose metabolism in humans Mar´ıa M. Adeva-Andany*1, Noemi Perez-Felpete*,´ Carlos Fernandez-Fern´ andez*,´ Cristobal´ Donapetry-Garc´ıa* and Cristina Pazos-Garc´ıa* *Nephrology Division, Hospital General Juan Cardona, c/ Pardo Bazan´ s/n, 15406 Ferrol, Spain Synopsis Information about normal hepatic glucose metabolism may help to understand pathogenic mechanisms underlying obesity and diabetes mellitus. In addition, liver glucose metabolism is involved in glycosylation reactions and con- nected with fatty acid metabolism. The liver receives dietary carbohydrates directly from the intestine via the portal vein. Glucokinase phosphorylates glucose to glucose 6-phosphate inside the hepatocyte, ensuring that an adequate flow of glucose enters the cell to be metabolized. Glucose 6-phosphate may proceed to several metabolic path- ways. During the post-prandial period, most glucose 6-phosphate is used to synthesize glycogen via the formation of glucose 1-phosphate and UDP–glucose. Minor amounts of UDP–glucose are used to form UDP–glucuronate and UDP– galactose, which are donors of monosaccharide units used in glycosylation. A second pathway of glucose 6-phosphate metabolism is the formation of fructose 6-phosphate, which may either start the hexosamine pathway to produce UDP-N-acetylglucosamine or follow the glycolytic pathway to generate pyruvate and then acetyl-CoA. Acetyl-CoA may enter the tricarboxylic acid (TCA) cycle to be oxidized or may be exported to the cytosol to synthesize fatty acids, when excess glucose is present within the hepatocyte. Finally, glucose 6-phosphate may produce NADPH and ribose 5-phosphate through the pentose phosphate pathway. -

Noelia Díaz Blanco
Effects of environmental factors on the gonadal transcriptome of European sea bass (Dicentrarchus labrax), juvenile growth and sex ratios Noelia Díaz Blanco Ph.D. thesis 2014 Submitted in partial fulfillment of the requirements for the Ph.D. degree from the Universitat Pompeu Fabra (UPF). This work has been carried out at the Group of Biology of Reproduction (GBR), at the Department of Renewable Marine Resources of the Institute of Marine Sciences (ICM-CSIC). Thesis supervisor: Dr. Francesc Piferrer Professor d’Investigació Institut de Ciències del Mar (ICM-CSIC) i ii A mis padres A Xavi iii iv Acknowledgements This thesis has been made possible by the support of many people who in one way or another, many times unknowingly, gave me the strength to overcome this "long and winding road". First of all, I would like to thank my supervisor, Dr. Francesc Piferrer, for his patience, guidance and wise advice throughout all this Ph.D. experience. But above all, for the trust he placed on me almost seven years ago when he offered me the opportunity to be part of his team. Thanks also for teaching me how to question always everything, for sharing with me your enthusiasm for science and for giving me the opportunity of learning from you by participating in many projects, collaborations and scientific meetings. I am also thankful to my colleagues (former and present Group of Biology of Reproduction members) for your support and encouragement throughout this journey. To the “exGBRs”, thanks for helping me with my first steps into this world. Working as an undergrad with you Dr. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Antigen-Specific Memory CD4 T Cells Coordinated Changes in DNA
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology The Journal of Immunology published online 18 March 2013 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2013/03/17/jimmun ol.1202267 Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto, Katsumi Ogoshi, Atsushi Sasaki, Jun Abe, Wei Qu, Yoichiro Nakatani, Budrul Ahsan, Kenshiro Oshima, Francis H. W. Shand, Akio Ametani, Yutaka Suzuki, Shuichi Kaneko, Takashi Wada, Masahira Hattori, Sumio Sugano, Shinichi Morishita and Kouji Matsushima J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Freely available online through http://www.jimmunol.org/content/suppl/2013/03/18/jimmunol.120226 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. Published March 18, 2013, doi:10.4049/jimmunol.1202267 The Journal of Immunology Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto,*,†,‡ Katsumi Ogoshi,* Atsushi Sasaki,† Jun Abe,* Wei Qu,† Yoichiro Nakatani,† Budrul Ahsan,x Kenshiro Oshima,† Francis H. -

Genomic Analyses of PMBL Reveal New Drivers and Mechanisms of Sensitivity to PD-1 Blockade
Regular Article LYMPHOID NEOPLASIA Genomic analyses of PMBL reveal new drivers and mechanisms of sensitivity to PD-1 blockade Bjoern Chapuy,1,2,* Chip Stewart,3,* Andrew J. Dunford,3,* Jaegil Kim,3 Kirsty Wienand,1 Atanas Kamburov,3 Gabriel K. Griffin,4 Pei-Hsuan Chen,4 Ana Lako,4 Robert A. Redd,5 Claire M. Cote,1 Matthew D. Ducar,6 Aaron R. Thorner,6 Scott J. Rodig,4 Gad Getz,3,7,8,† Downloaded from https://ashpublications.org/blood/article-pdf/134/26/2369/1549702/bloodbld2019002067.pdf by Margaret Shipp on 30 December 2019 and Margaret A. Shipp1,† 1Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA; 2Department of Hematology and Oncology, University Medical Center Gottingen,¨ Gottingen,¨ Germany; 3Broad Institute of Harvard and Massachusetts Institute of Technology, Cambridge, MA; 4Department of Pathology, Brigham and Women’s Hospital, Harvard Medical School, Boston, MA; 5Department of Data Sciences and 6Center for Cancer Genome Discovery, Dana-Farber Cancer Institute, Boston, MA; and 7Department of Pathology and 8Center for Cancer Research, Massachusetts General Hospital, Harvard Medical School, Boston, MA KEY POINTS Primary mediastinal large B-cell lymphomas (PMBLs) are aggressive tumors that typically present as large mediastinal masses in young women. PMBLs share clinical, transcriptional, l Comprehensive genomic analyses of and molecular features with classical Hodgkin lymphoma (cHL), including constitutive PMBL reveal new activation of nuclear factor kB (NF-kB), JAK/STAT signaling, and programmed cell death genetic drivers such protein 1 (PD-1)–mediated immune evasion. The demonstrated efficacy of PD-1 blockade in as ZNF217. relapsed/refractory PMBLs led to recent approval by the US Food and Drug Administration l High mutational and underscored the importance of characterizing targetable genetic vulnerabilities in this burden, MSI, and disease. -

Transcriptome Analysis of Human Diabetic Kidney Disease
ORIGINAL ARTICLE Transcriptome Analysis of Human Diabetic Kidney Disease Karolina I. Woroniecka,1 Ae Seo Deok Park,1 Davoud Mohtat,2 David B. Thomas,3 James M. Pullman,4 and Katalin Susztak1,5 OBJECTIVE—Diabetic kidney disease (DKD) is the single cases, mild and then moderate mesangial expansion can be leading cause of kidney failure in the U.S., for which a cure has observed. In general, diabetic kidney disease (DKD) is not yet been found. The aim of our study was to provide an considered a nonimmune-mediated degenerative disease unbiased catalog of gene-expression changes in human diabetic of the glomerulus; however, it has long been noted that kidney biopsy samples. complement and immunoglobulins sometimes can be de- — tected in diseased glomeruli, although their role and sig- RESEARCH DESIGN AND METHODS Affymetrix expression fi arrays were used to identify differentially regulated transcripts in ni cance is not clear (4). 44 microdissected human kidney samples. The DKD samples were The understanding of DKD has been challenged by multi- significant for their racial diversity and decreased glomerular ple issues. First, the diagnosis of DKD usually is made using filtration rate (~20–30 mL/min). Stringent statistical analysis, using clinical criteria, and kidney biopsy often is not performed. the Benjamini-Hochberg corrected two-tailed t test, was used to According to current clinical practice, the development of identify differentially expressed transcripts in control and diseased albuminuria in patients with diabetes is sufficient to make the glomeruli and tubuli. Two different Web-based algorithms were fi diagnosis of DKD (5). We do not understand the correlation used to de ne differentially regulated pathways. -
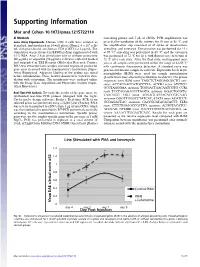
Supporting Information
Supporting Information Mor and Cohen 10.1073/pnas.1215722110 SI Methods containing primer and 5 μL of cDNA. PCR amplification was Gene Array Experiments. Human CD4 T cells were isolated as preceded by incubation of the mixture for 10 min at 95 °C, and described, and incubated in 24-well plates (Nunc), 4 × 106 cells/ the amplification step consisted of 45 cycles of denaturation, mL with plate-bound anti-human CD3 (OKT3) at 2 μg/mL. The annealing, and extension. Denaturation was performed for 15 s stimulation was performed in RPMI medium supplemented with at 95 °C; annealing was performed in 60 °C; and the extension 0.1% BSA. After 2 h of stimulation with or without cefuroxime was performed at 72 °C for 20 s, with fluorescence detection at (50 μg/mL) or ampicillin (50 μg/mL), cells were collected washed 72 °C after each cycle. After the final cycle, melting-point anal- and suspended in TRI Reagent (Molecular Research Center). yses of all samples were performed within the range of 62–95 °C RNA was extracted from samples and used to prepare probes for with continuous fluorescence detection. A standard curve was gene array in accord with the manufacturer’s instructions (Super- generated from one sample in each run. Expression levels of β2- Array Bioscience). Adequate labeling of the probes was tested microglobulin (B2M) were used for sample normalization before hybridization. Three healthy donors were tested in stim- (β-actin levels were affected by cefuroxime treatment). The primer ulation with cefuroxime. The membranes were analyzed online sequences were B2M sense TAGCTCTAGGAGGGCTG anti- with the Image Data Acquisition and Expression Analysis (Super- sense ACCACAACCATGCCTTA; ACVR2 sense ATCTCC- Array Bioscience). -

Genome-Wide Identification of Genes Regulating DNA Methylation Using Genetic Anchors for Causal Inference
bioRxiv preprint doi: https://doi.org/10.1101/823807; this version posted October 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Genome-wide identification of genes regulating DNA methylation using genetic anchors for causal inference Paul J. Hop1,2, René Luijk1, Lucia Daxinger3, Maarten van Iterson1, Koen F. Dekkers1, Rick Jansen4, BIOS Consortium5, Joyce B.J. van Meurs6, Peter A.C. ’t Hoen 7, M. Arfan Ikram8, Marleen M.J. van Greevenbroek9,10, Dorret I. Boomsma11, P. Eline Slagboom1, Jan H. Veldink2, Erik W. van Zwet12, Bastiaan T. Heijmans1* 1 Molecular Epidemiology, Department of Biomedical Data Sciences, Leiden University Medical Center, Leiden, 2333 ZC, The Netherlands 2 Department of Neurology, UMC Utrecht Brain Center, University Medical Centre Utrecht, Utrecht University, Utrecht 3584 CG, Netherlands 3 Department of Human Genetics, Leiden University Medical Center, Leiden, 2333 ZC, The Netherlands 4 Department of Psychiatry, Amsterdam UMC, Vrije Universiteit Amsterdam, Amsterdam Neuroscience, Amsterdam, 1081 HV, The Netherlands 5 Biobank-based Integrated Omics Study Consortium. For a complete list of authors, see the acknowledgements. 6 Department of Internal Medicine, Erasmus Medical Centre, Rotterdam, The Netherlands. 7 Centre for Molecular and Biomolecular Informatics, Radboud Institute for Molecular Life Sciences, Radboud University -

The Effect of Rbfox2 Modulation on Retinal Transcriptome and Visual
www.nature.com/scientificreports OPEN The efect of Rbfox2 modulation on retinal transcriptome and visual function Lei Gu1, Riki Kawaguchi2, Joseph Caprioli1,3 & Natik Piri1,3* Rbfox proteins regulate alternative splicing, mRNA stability and translation. These proteins are involved in neurogenesis and have been associated with various neurological conditions. Here, we analyzed Rbfox2 expression in adult and developing mouse retinas and the efect of its downregulation on visual function and retinal transcriptome. In adult rodents, Rbfox2 is expressed in all retinal ganglion cell (RGC) subtypes, horizontal cells, as well as GABAergic amacrine cells (ACs). Among GABAergic AC subtypes, Rbfox2 was colocalized with cholinergic starburst ACs, NPY (neuropeptide Y)- and EBF1 (early B-cell factor 1)-positive ACs. In diferentiating retinal cells, Rbfox2 expression was observed as early as E12 and, unlike Rbfox1, which changes its subcellular localization from cytoplasmic to predominantly nuclear at around P0, Rbfox2 remains nuclear throughout retinal development. Rbfox2 knockout in adult animals had no detectable efect on retinal gross morphology. However, the visual clif test revealed a signifcant abnormality in the depth perception of Rbfox2- defcient animals. Gene set enrichment analysis identifed genes regulating the RNA metabolic process as a top enriched class of genes in Rbfox2-defcient retinas. Pathway analysis of the top 100 diferentially expressed genes has identifed Rbfox2-regulated genes associated with circadian rhythm and entrainment, glutamatergic/cholinergic/dopaminergic synaptic function, calcium and PI3K-AKT signaling. Te family of RNA binding protein, fox-1 (Rbfox) homolog includes three evolutionarily conserved members, Rbfox1, Rbfox2 and Rbfox3, that regulate cell- or tissue-specifc alternative splicing at diferent developmental stages. -

Post-Translational Modification of MRE11: Its Implication in DDR And
G C A T T A C G G C A T genes Review Post-Translational Modification of MRE11: Its Implication in DDR and Diseases Ruiqing Lu 1,† , Han Zhang 2,† , Yi-Nan Jiang 1, Zhao-Qi Wang 3,4, Litao Sun 5,* and Zhong-Wei Zhou 1,* 1 School of Medicine, Sun Yat-Sen University, Shenzhen 518107, China; [email protected] (R.L.); [email protected] (Y.-N.J.) 2 Institute of Medical Biology, Chinese Academy of Medical Sciences and Peking Union Medical College; Kunming 650118, China; [email protected] 3 Leibniz Institute on Aging–Fritz Lipmann Institute (FLI), 07745 Jena, Germany; zhao-qi.wang@leibniz-fli.de 4 Faculty of Biological Sciences, Friedrich-Schiller-University of Jena, 07745 Jena, Germany 5 School of Public Health (Shenzhen), Sun Yat-Sen University, Shenzhen 518107, China * Correspondence: [email protected] (L.S.); [email protected] (Z.-W.Z.) † These authors contributed equally to this work. Abstract: Maintaining genomic stability is vital for cells as well as individual organisms. The meiotic recombination-related gene MRE11 (meiotic recombination 11) is essential for preserving genomic stability through its important roles in the resection of broken DNA ends, DNA damage response (DDR), DNA double-strand breaks (DSBs) repair, and telomere maintenance. The post-translational modifications (PTMs), such as phosphorylation, ubiquitination, and methylation, regulate directly the function of MRE11 and endow MRE11 with capabilities to respond to cellular processes in promptly, precisely, and with more diversified manners. Here in this paper, we focus primarily on the PTMs of MRE11 and their roles in DNA response and repair, maintenance of genomic stability, as well as their Citation: Lu, R.; Zhang, H.; Jiang, association with diseases such as cancer. -

Recurrent Genetic Mutations in Lymphoid Malignancies
Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1312 Recurrent Genetic Mutations in Lymphoid Malignancies EMMA YOUNG ACTA UNIVERSITATIS UPSALIENSIS ISSN 1651-6206 ISBN 978-91-554-9850-4 UPPSALA urn:nbn:se:uu:diva-314956 2017 Dissertation presented at Uppsala University to be publicly examined in Rudbecksalen, Dag Hammarskjölds Väg 20, Uppsala, Friday, 5 May 2017 at 13:15 for the degree of Doctor of Philosophy (Faculty of Medicine). The examination will be conducted in English. Faculty examiner: Professor Dimitar Efremov (Molecular Hematology Group of the International Centre for Genetic Engineering and Biotechnology (ICGEB) in Trieste, Italy). Abstract Young, E. 2017. Recurrent Genetic Mutations in Lymphoid Malignancies. Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1312. 82 pp. Uppsala: Acta Universitatis Upsaliensis. ISBN 978-91-554-9850-4. In recent years, the genetic landscape of B-cell derived lymphoid malignancies, including chronic lymphocytic leukemia (CLL), has been rapidly unraveled, identifying recurrent genetic mutations with potential clinical impact. Interestingly, ~30% of all CLL patients can be assigned to more homogeneous subsets based on the expression of a similar or “stereotyped” B-cell receptor (BcR). Considering that biased distribution of genetic mutations was recently indicated in specific stereotyped subsets, in paper I, we screened 565 subset cases, preferentially assigned to clinically aggressive subsets, and confirm the SF3B1 mutational bias in subset #2 (45%), but also report on similarly marked enrichment in subset #3 (46%). In contrast, NOTCH1 mutations were predominantly detected in subsets #1, #8, #59 and #99 (22-34%). This data further highlights a subset-biased acquisition of genetic mutations in the pathogenesis of at least certain subsets.