Eukaryotic-Like Protein Kinases in the Prokaryotes and the Myxobacterial Kinome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomic Signatures of Predatory Bacteria
The ISME Journal (2013) 7, 756–769 & 2013 International Society for Microbial Ecology All rights reserved 1751-7362/13 www.nature.com/ismej ORIGINAL ARTICLE By their genes ye shall know them: genomic signatures of predatory bacteria Zohar Pasternak1, Shmuel Pietrokovski2, Or Rotem1, Uri Gophna3, Mor N Lurie-Weinberger3 and Edouard Jurkevitch1 1Department of Plant Pathology and Microbiology, The Hebrew University of Jerusalem, Rehovot, Israel; 2Department of Molecular Genetics, Weizmann Institute of Science, Rehovot, Israel and 3Department of Molecular Microbiology and Biotechnology, George S. Wise Faculty of Life Sciences, Tel Aviv University, Tel Aviv, Israel Predatory bacteria are taxonomically disparate, exhibit diverse predatory strategies and are widely distributed in varied environments. To date, their predatory phenotypes cannot be discerned in genome sequence data thereby limiting our understanding of bacterial predation, and of its impact in nature. Here, we define the ‘predatome,’ that is, sets of protein families that reflect the phenotypes of predatory bacteria. The proteomes of all sequenced 11 predatory bacteria, including two de novo sequenced genomes, and 19 non-predatory bacteria from across the phylogenetic and ecological landscapes were compared. Protein families discriminating between the two groups were identified and quantified, demonstrating that differences in the proteomes of predatory and non-predatory bacteria are large and significant. This analysis allows predictions to be made, as we show by confirming from genome data an over-looked bacterial predator. The predatome exhibits deficiencies in riboflavin and amino acids biosynthesis, suggesting that predators obtain them from their prey. In contrast, these genomes are highly enriched in adhesins, proteases and particular metabolic proteins, used for binding to, processing and consuming prey, respectively. -

First Genomic Insights Into Members of a Candidate Bacterial Phylum Responsible for Wastewater Bulking
First genomic insights into members of a candidate bacterial phylum responsible for wastewater bulking Yuji Sekiguchi1, Akiko Ohashi1, Donovan H. Parks2, Toshihiro Yamauchi3, Gene W. Tyson2,4 and Philip Hugenholtz2,5 1 Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Tsukuba, Ibaraki, Japan 2 Australian Centre for Ecogenomics, School of Chemistry and Molecular Biosciences, The University of Queensland, St. Lucia, Queensland, Australia 3 Administrative Management Department, Kubota Kasui Corporation, Minato-ku, Tokyo, Japan 4 Advanced Water Management Centre, The University of Queensland, St. Lucia, Queensland, Australia 5 Institute for Molecular Bioscience, The University of Queensland, St. Lucia, Queensland, Australia ABSTRACT Filamentous cells belonging to the candidate bacterial phylum KSB3 were previously identified as the causative agent of fatal filament overgrowth (bulking) in a high-rate industrial anaerobic wastewater treatment bioreactor. Here, we obtained near complete genomes from two KSB3 populations in the bioreactor, including the dominant bulking filament, using diVerential coverage binning of metagenomic data. Fluorescence in situ hybridization with 16S rRNA-targeted probes specific for the two populations confirmed that both are filamentous organisms. Genome-based metabolic reconstruction and microscopic observation of the KSB3 filaments in the presence of sugar gradients indicate that both filament types are Gram-negative, strictly anaerobic fermenters capable of -

Kinome Profiling of Clinical Cancer Specimens
Published OnlineFirst March 23, 2010; DOI: 10.1158/0008-5472.CAN-09-3989 Review Cancer Research Kinome Profiling of Clinical Cancer Specimens Kaushal Parikh and Maikel P. Peppelenbosch Abstract Over the past years novel technologies have emerged to enable the determination of the transcriptome and proteome of clinical samples. These data sets will prove to be of significant value to our elucidation of the mechanisms that govern pathophysiology and may provide biological markers for future guidance in person- alized medicine. However, an equally important goal is to define those proteins that participate in signaling pathways during the disease manifestation itself or those pathways that are made active during successful clinical treatment of the disease: the main challenge now is the generation of large-scale data sets that will allow us to define kinome profiles with predictive properties on the outcome-of-disease and to obtain insight into tissue-specific analysis of kinase activity. This review describes the current techniques available to gen- erate kinome profiles of clinical tissue samples and discusses the future strategies necessary to achieve new insights into disease mechanisms and treatment targets. Cancer Res; 70(7); 2575–8. ©2010 AACR. Background the current state of the cell or tissue as characterized by pro- teomic and metabolic measurements (3). The past five years have seen an exponential increase in In this review we describe and evaluate the various tech- technological development to explore the genome and the niques that have recently become available to study the ki- proteome to understand the molecular basis of disease. How- nomes of clinical tissue samples. -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
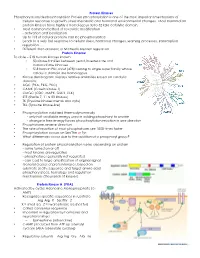
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

The Myxocoumarins a and B from Stigmatella Aurantiaca Strain MYX-030
The myxocoumarins A and B from Stigmatella aurantiaca strain MYX-030 Tobias A. M. Gulder*1, Snežana Neff2, Traugott Schüz2, Tammo Winkler2, René Gees2 and Bettina Böhlendorf*2 Full Research Paper Open Access Address: Beilstein J. Org. Chem. 2013, 9, 2579–2585. 1Kekulé Institute of Organic Chemistry and Biochemistry, University of doi:10.3762/bjoc.9.293 Bonn, Gerhard-Domagk-Straβe 1, 53121 Bonn, Germany and 2Syngenta Crop Protection AG, CH-4002 Basel, Switzerland Received: 21 August 2013 Accepted: 01 November 2013 Email: Published: 20 November 2013 Tobias A. M. Gulder* - [email protected]; Bettina Böhlendorf* - [email protected] This article is part of the Thematic Series "Natural products in synthesis and biosynthesis" and is dedicated to Prof. Dr. Gerhard Höfle on the * Corresponding author occasion of his 74th birthday. Keywords: Guest Editor: J. S. Dickschat antifungal activity; myxobacteria; natural products; Stigmatella aurantiaca; structure elucidation © 2013 Gulder et al; licensee Beilstein-Institut. License and terms: see end of document. Abstract The myxobacterial strain Stigmatella aurantiaca MYX-030 was selected as promising source for the discovery of new biologically active natural products by our screening methodology. The isolation, structure elucidation and initial biological evaluation of the myxocoumarins derived from this strain are described in this work. These compounds comprise an unusual structural framework and exhibit remarkable antifungal properties. Introduction Despite declining interest of most big R&D-driven chemical epothilones A (1), Figure 1), microtubule-stabilizing macrolac- companies in recent years, natural products continue to serve as tones that are clinically used in cancer therapy [16-20]. one of the most important sources of new bioactive chemical entities, both in the pharmaceutical [1-4] as well as in the agro- But also from an agrochemical point of view, myxobacteria chemical industry [5-8]. -

Evolution and Design Governing Signal Precision and Amplification
Evolution and Design Governing Signal Precision and Amplification in a Bacterial Chemosensory Pathway Mathilde Guzzo, Rym Agrebi, Leon Espinosa, Gregory Baronian, Virginie Molle, Emilia M. F. Mauriello, Céline Brochier-Armanet, Tam Mignot To cite this version: Mathilde Guzzo, Rym Agrebi, Leon Espinosa, Gregory Baronian, Virginie Molle, et al.. Evolution and Design Governing Signal Precision and Amplification in a Bacterial Chemosensory Pathway. PLoS Genetics, Public Library of Science, 2015, 11 (8), pp.e1005460. 10.1371/journal.pgen.1005460. hal- 01452074 HAL Id: hal-01452074 https://hal.archives-ouvertes.fr/hal-01452074 Submitted on 27 Sep 2018 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License RESEARCH ARTICLE Evolution and Design Governing Signal Precision and Amplification in a Bacterial Chemosensory Pathway Mathilde Guzzo1☯, Rym Agrebi1☯¤, Leon Espinosa1, Grégory Baronian2, Virginie Molle2, Emilia M. F. Mauriello1, Céline Brochier-Armanet3, Tâm Mignot1* 1 Laboratoire de Chimie Bactérienne, Institut de Microbiologie de la Méditerranée, CNRS Aix-Marseille University UMR 7283, Marseille, France, 2 Laboratoire de Dynamique des Interactions Membranaires Normales et Pathologiques, CNRS Universités de Montpellier II et I, UMR 5235, case 107, Montpellier, France, 3 Université de Lyon, Université Lyon 1, CNRS, UMR5558, Laboratoire de Biométrie et Biologie Evolutive, Villeurbanne, France ☯ These authors contributed equally to this work. -

Identification of Protein O-Glcnacylation Sites Using Electron Transfer Dissociation Mass Spectrometry on Native Peptides
Identification of protein O-GlcNAcylation sites using electron transfer dissociation mass spectrometry on native peptides Robert J. Chalkleya, Agnes Thalhammerb, Ralf Schoepfera,b, and A. L. Burlingamea,1 aDepartment of Pharmaceutical Chemistry, University of California, 600 16th Street, Genentech Hall, Suite N472, San Francisco, CA 94158; and bLaboratory for Molecular Pharmacology, and Department of Neuroscience, Physiology, and Pharmacology, University College London, Gower Street, London WC1E 6BT, United Kingdom Edited by James A. Wells, University of California, San Francisco, CA, and approved April 15, 2009 (received for review January 12, 2009) Protein O-GlcNAcylation occurs in all animals and plants and is and synaptic plasticity have been found O-GlcNAcylated (8). The implicated in modulation of a wide range of cytosolic and nuclear importance of O-GlcNAcylation is highlighted by the brain-specific protein functions, including gene silencing, nutrient and stress sens- OGT knockout mouse model that displays developmental defects ing, phosphorylation signaling, and diseases such as diabetes and that result in neonatal death (9). Most interestingly, in analogy to Alzheimer’s. The limiting factor impeding rapid progress in decipher- phosphorylation, synaptic activity leads to a dynamic modulation of ing the biological functions of protein O-GlcNAcylation has been the O-GlcNAc levels (10) and is also involved in long-term potentiation inability to easily identify exact residues of modification. We describe and memory (11). a robust, high-sensitivity strategy able to assign O-GlcNAcylation sites Phosphorylation is the most heavily studied regulatory PTM, and of native modified peptides using electron transfer dissociation mass proteomic approaches for global enrichment, detection, and char- spectrometry. -

Inactivation of Stigmatella Aurantiaca Csga Gene Impares Rippling Formation- Genetika, Vol 47, No
UDC 575 DOI: 10.2298/GENSR1501119M Original scientific paper INACTIVATION OF Stigmatella aurantiaca csg A GENE IMPARES RIPPLING FORMATION Ana MILOSEVIC-DJERIC 1, Susanne MÜLLER 2, and Hans URLICH SCHAIRER 3 1 Department of Gynaecology and Obstetrics, Hospital Centre Uzice, Uzice, Serbia 2 Univeristy of Iowa, Iowa City, USA 3Zentrum fur Molekulare Biologie, ZMBH, Heidelberg, Germany Milosevic-Djeric A., S. Müller, and H. Urlich Schairer (2015): Inactivation of Stigmatella aurantiaca csgA gene impares rippling formation- Genetika, Vol 47, No. 1, 119-130. Stigmatella aurantiaca fruiting body development depends on cell-cell interactions. One type of the signaling molecule stigmolone isolated from S. aurantiaca cells acts to help cells to stay together in the aggregation phase. Another gene product involved in intercellular signaling in S. aurantiaca is the csg A homolog of Myxococcus xanthus . In close relative M. xanthus C signal the product of the csg A gene is required for rippling, aggregation and sporulation. Isolation of homologous gene in S. aurantiaca implicates a probable role of CsgA in intercellular communication. Inactivation of the gene by insertion mutagenesis caused alterations in S. aurantiaca fruiting. The motility behavior of the cells during development was changed as well as their ability to stay more closely together in the early stages of development. Inactivation of the csg A gene completely abolished rippling of the cells. This indicates the crucial role of the CsgA protein in regulating this rhythmic behavior. Key words: cell-cell signaling, development, fruiting body formation, myxobacteria, Stigmatella aurantiaca INTRODUCTION Stigmatella aurantiaca belongs to the myxobacteria famous for their ability to form multicellular structures fruiting bodies in response to the starvation conditions ( SHIMKETS et al ., 2006; ZHANG et al., 2012; CLAESSEN et al ., 2014). -

GSK3 and Its Interactions with the PI3K/AKT/Mtor Signalling Network
Heriot-Watt University Research Gateway GSK3 and its interactions with the PI3K/AKT/mTOR signalling network Citation for published version: Hermida, MA, Kumar, JD & Leslie, NR 2017, 'GSK3 and its interactions with the PI3K/AKT/mTOR signalling network', Advances in Biological Regulation, vol. 65, pp. 5-15. https://doi.org/10.1016/j.jbior.2017.06.003 Digital Object Identifier (DOI): 10.1016/j.jbior.2017.06.003 Link: Link to publication record in Heriot-Watt Research Portal Document Version: Peer reviewed version Published In: Advances in Biological Regulation Publisher Rights Statement: © 2017 Elsevier B.V. General rights Copyright for the publications made accessible via Heriot-Watt Research Portal is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy Heriot-Watt University has made every reasonable effort to ensure that the content in Heriot-Watt Research Portal complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 27. Sep. 2021 Accepted Manuscript GSK3 and its interactions with the PI3K/AKT/mTOR signalling network Miguel A. Hermida, J. Dinesh Kumar, Nick R. Leslie PII: S2212-4926(17)30124-0 DOI: 10.1016/j.jbior.2017.06.003 Reference: JBIOR 180 To appear in: Advances in Biological Regulation Received Date: 13 June 2017 Accepted Date: 23 June 2017 Please cite this article as: Hermida MA, Dinesh Kumar J, Leslie NR, GSK3 and its interactions with the PI3K/AKT/mTOR signalling network, Advances in Biological Regulation (2017), doi: 10.1016/ j.jbior.2017.06.003. -

Novel Myxobacteria As a Potential Source of Natural Products and Description of Inter-Species Nature of C-Signal
Novel myxobacteria as a potential source of natural products and description of inter-species nature of C-signal Dissertation zur Erlangung des Grades des Doktors der Naturwissenschaften der Naturwissenschaftlich-Technischen Fakultät III Chemie, Pharmazie, Bio- und Werkstoffwissenschaften der Universität des Saarlandes von Ram Prasad Awal (M. Sc. in Medical Microbiology) Saarbrücken 2016 Tag des Kolloquiums: ......19.12.2016....................................... Dekan: ......Prof. Dr. Guido Kickelbick.............. Berichterstatter: ......Prof. Dr. Rolf Müller...................... ......Prof. Dr. Manfred J. Schmitt........... ............................................................... Vositz: ......Prof. Dr. Uli Kazmaier..................... Akad. Mitarbeiter: ......Dr. Jessica Stolzenberger.................. iii Diese Arbeit entstand unter der Anleitung von Prof. Dr. Rolf Müller in der Fachrichtung 8.2, Pharmazeutische Biotechnologie der Naturwissenschaftlich-Technischen Fakultät III der Universität des Saarlandes von Oktober 2012 bis September 2016. iv Acknowledgement Above all, I would like to express my special appreciation and thanks to my advisor Professor Dr. Rolf Müller. It has been an honor to be his Ph.D. student and work in his esteemed laboratory. I appreciate for his supervision, inspiration and for allowing me to grow as a research scientist. Your guidance on both research as well as on my career have been invaluable. I would also like to thank Professor Dr. Manfred J. Schmitt for his scientific support and suggestions to my research work. I am thankful for my funding sources that made my Ph.D. work possible. I was funded by Deutscher Akademischer Austauschdienst (DAAD) for 3 and half years and later on by Helmholtz-Institute. Many thanks to my co-advisors: Dr. Carsten Volz, who supported and guided me through the challenging research and Dr. Ronald Garcia for introducing me to the wonderful world of myxobacteria. -

Modified Recipe to Inhibit GSK-3 for the Living Fungal Biomaterial Manufacture
bioRxiv preprint doi: https://doi.org/10.1101/496265; this version posted December 13, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Modified recipe to inhibit GSK-3 for the living fungal 2 biomaterial manufacture 1¶ 1¶ 1 1 1 3 Jinhui Chang , Po Lam Chan , Yichun Xie , Man Kit Cheung , Ka Lee Ma , Hoi Shan 1* 4 Kwan 1 5 School of Life Sciences, The Chinese University of Hong Kong, Shatin, New Territories, 6 Hong Kong 7 *Corresponding author: E-mail: [email protected] ¶ 8 These authors contributed equally to this work. 9 10 11 bioRxiv preprint doi: https://doi.org/10.1101/496265; this version posted December 13, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 12 Abstract 13 Living fungal mycelium with suppressed or abolished fruit-forming ability is a self-healing 14 substance particularly valuable biomaterial for further engineering and development in 15 applications such as monitoring/sensing environmental changes and secreting signals. The 16 ability to suppress fungal fruiting is also a useful tool for maintaining stability (e.g., shape, 17 form) of a mycelium-based biomaterial with ease and lower cost. 18 The objective of this present study is to provide a biochemical solution to regulate the fruiting 19 body formation to replace heat killing of mycelium during production. -

Polyamine Distribution Profiles Among Some Members Within Delta-And Epsilon-Subclasses of Proteobacteria
Microbiol. Cult. Coll. June. 2004. p. 3 ― 8 Vol. 20, No. 1 Polyamine Distribution Profiles among Some Members within Delta-and Epsilon-Subclasses of Proteobacteria Koei Hamana1)*, Tomoko Saito1), Mami Okada1), and Masaru Niitsu2) 1)Department of Laboratory Sciences, School of Health Sciences, Faculty of Medicine, Gunma University, 39- 15 Showa-machi 3-chome, Maebashi, Gunma 371-8514, Japan 2)Faculty of Pharmaceutical Sciences, Josai University, Keyakidai 1-chome-1, Sakado, Saitama 350-0295, Japan Cellular polyamines of 18 species(13 genera)belonging to the delta and epsilon subclasses of the class Proteobacteria were analyzed by HPLC and GC. In the delta subclass, the four marine myxobacteria(the order Myxococcales), Enhygromyxa salina, Haliangium ochroceum, Haliangium tepidum and Plesiocystis pacifica contained spermidine. Fe(III)-reducing two Geobacter species and two Pelobacter species belonging to the order Desulfuromonadales con- tained spermidine. Bdellovibrio bacteriovorus was absent in cellular polyamines. Bacteriovorax starrii contained putrescine and spermidine. Bacteriovorax stolpii contained spermidine and homo- spermidine. Spermidine was the major polyamine in the sulfate-reducing delta proteobacteria belonging to the genera Desulfovibrio, Desulfacinum, Desulfobulbus, Desulfococcus and Desulfurella, and some species of them contained cadaverine. Within the epsilon subclass, three Sulfurospirillum species ubiquitously contained spermidine and one of the three contained sper- midine and cadaverine. Thiomicrospora denitrificans contained cadaverine and spermidine as the major polyamine. These data show that cellular polyamine profiles can be used as a chemotaxonomic marker within delta and epsilon subclasses. Key words: polyamine, spermidine, homospermidine, Proteobacteria The class Proteobacteria is a major taxon of the 18, 26). Fe(Ⅲ)-reducing members belonging to the gen- domain Bacteria and is phylogenetically divided into the era Pelobacter, Geobacter, Desulfuromonas and alpha, beta, gamma, delta and epsilon subclasses.