Brevibacterium Linens Relevant to Cheese Ripening: a Review1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Growth and Adaptation of Microorganisms on the Cheese Surface Christophe Monnet, Sophie Landaud-Liautaud, Pascal Bonnarme, Dominique Swennen
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Archive Ouverte en Sciences de l'Information et de la Communication Growth and adaptation of microorganisms on the cheese surface Christophe Monnet, Sophie Landaud-Liautaud, Pascal Bonnarme, Dominique Swennen To cite this version: Christophe Monnet, Sophie Landaud-Liautaud, Pascal Bonnarme, Dominique Swennen. Growth and adaptation of microorganisms on the cheese surface. FEMS Microbiology Letters, Wiley-Blackwell, 2015, 362 (1), pp.1-9. 10.1093/femsle/fnu025. hal-01535275 HAL Id: hal-01535275 https://hal.archives-ouvertes.fr/hal-01535275 Submitted on 28 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. 1 Growth and adaptation of microorganisms on the cheese surface 2 3 4 Christophe Monnet1,2, Sophie Landaud2,1, Pascal Bonnarme1,2 & Dominique Swennen3,4 5 6 1 INRA, UMR782 Génie et Microbiologie des Procédés Alimentaires, 78370 Thiverval- 7 Grignon, France 8 2 AgroParisTech, UMR782 Génie et Microbiologie des Procédés Alimentaires, 78370 9 Thiverval-Grignon, France 10 3 INRA, UMR1319 Micalis, 78370 Thiverval-Grignon, France 11 4 AgroParisTech, UMR1319 Micalis, 78370 Thiverval-Grignon, France 12 13 * Corresponding author. -

Corynebacterium Sp.|NML98-0116
1 Limnochorda_pilosa~GCF_001544015.1@NZ_AP014924=Bacteria-Firmicutes-Limnochordia-Limnochordales-Limnochordaceae-Limnochorda-Limnochorda_pilosa 0,9635 Ammonifex_degensii|KC4~GCF_000024605.1@NC_013385=Bacteria-Firmicutes-Clostridia-Thermoanaerobacterales-Thermoanaerobacteraceae-Ammonifex-Ammonifex_degensii 0,985 Symbiobacterium_thermophilum|IAM14863~GCF_000009905.1@NC_006177=Bacteria-Firmicutes-Clostridia-Clostridiales-Symbiobacteriaceae-Symbiobacterium-Symbiobacterium_thermophilum Varibaculum_timonense~GCF_900169515.1@NZ_LT827020=Bacteria-Actinobacteria-Actinobacteria-Actinomycetales-Actinomycetaceae-Varibaculum-Varibaculum_timonense 1 Rubrobacter_aplysinae~GCF_001029505.1@NZ_LEKH01000003=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_aplysinae 0,975 Rubrobacter_xylanophilus|DSM9941~GCF_000014185.1@NC_008148=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_xylanophilus 1 Rubrobacter_radiotolerans~GCF_000661895.1@NZ_CP007514=Bacteria-Actinobacteria-Rubrobacteria-Rubrobacterales-Rubrobacteraceae-Rubrobacter-Rubrobacter_radiotolerans Actinobacteria_bacterium_rbg_16_64_13~GCA_001768675.1@MELN01000053=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_rbg_16_64_13 1 Actinobacteria_bacterium_13_2_20cm_68_14~GCA_001914705.1@MNDB01000040=Bacteria-Actinobacteria-unknown_class-unknown_order-unknown_family-unknown_genus-Actinobacteria_bacterium_13_2_20cm_68_14 1 0,9803 Thermoleophilum_album~GCF_900108055.1@NZ_FNWJ01000001=Bacteria-Actinobacteria-Thermoleophilia-Thermoleophilales-Thermoleophilaceae-Thermoleophilum-Thermoleophilum_album -

Biological and Chemical Diversity of Bacteria Associated with a Marine Flatworm
Article Biological and Chemical Diversity of Bacteria Associated with a Marine Flatworm Hui-Na Lin 1,2,†, Kai-Ling Wang 1,3,†, Ze-Hong Wu 4,5, Ren-Mao Tian 6, Guo-Zhu Liu 7 and Ying Xu 1,* 1 Shenzhen Key Laboratory of Marine Bioresource & Eco-Environmental Science, Shenzhen Engineering Laboratory for Marine Algal Biotechnology, College of Life Sciences and Oceanography, Shenzhen University, Shenzhen 518060, China; [email protected] (H.-N.L.); [email protected] (K.-L.W.) 2 School of Life Sciences, Xiamen University, Xiamen 361102, China 3 Key Laboratory of Marine Drugs, Ministry of Education of China, School of Medicine and Pharmacy, Ocean University of China, Qingdao 266003, China 4 The Eighth Affiliated Hospital, Sun Yat-sen University, Shenzhen 518033, China; [email protected] 5 Integrated Chinese and Western Medicine Postdoctoral Research Station, Jinan University, Guangzhou 510632, China 6 Division of Life Science, The Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong SAR, China; [email protected] 7 HEC Research and Development Center, HEC Pharm Group, Dongguan 523871, China; [email protected] * Correspondence: [email protected]; Tel.: +86-755-26958849; Fax: +86-755-26534274 † These authors contributed equally to this work. Received: 20 July 2017; Accepted: 29 August 2017; Published: 1 September 2017 Abstract: The aim of this research is to explore the biological and chemical diversity of bacteria associated with a marine flatworm Paraplanocera sp., and to discover the bioactive metabolites from culturable strains. A total of 141 strains of bacteria including 45 strains of actinomycetes and 96 strains of other bacteria were isolated, identified and fermented on a small scale. -

Studies on Antibacterial Activity and Diversity of Cultivable Actinobacteria Isolated from Mangrove Soil in Futian and Maoweihai of China
Hindawi Evidence-Based Complementary and Alternative Medicine Volume 2019, Article ID 3476567, 11 pages https://doi.org/10.1155/2019/3476567 Research Article Studies on Antibacterial Activity and Diversity of Cultivable Actinobacteria Isolated from Mangrove Soil in Futian and Maoweihai of China Feina Li,1 Shaowei Liu,1 Qinpei Lu,1 Hongyun Zheng,1,2 Ilya A. Osterman,3,4 Dmitry A. Lukyanov,3 Petr V. Sergiev,3,4 Olga A. Dontsova,3,4,5 Shuangshuang Liu,6 Jingjing Ye,1,2 Dalin Huang ,2 and Chenghang Sun 1 1 Institute of Medicinal Biotechnology, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing 100050, China 2College of Basic Medical Sciences, Guilin Medical University, Guilin 541004, China 3Center of Life Sciences, Skolkovo Institute of Science and Technology, Moscow 143025, Russia 4Department of Chemistry, A.N. Belozersky Institute of Physico-Chemical Biology, Lomonosov Moscow State University, Moscow 119992, Russia 5Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Te Russian Academy of Sciences, Moscow 117997, Russia 6China Pharmaceutical University, Nanjing 210009, China Correspondence should be addressed to Dalin Huang; [email protected] and Chenghang Sun; [email protected] Received 29 March 2019; Accepted 21 May 2019; Published 9 June 2019 Guest Editor: Jayanta Kumar Patra Copyright © 2019 Feina Li et al. Tis is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Mangrove is a rich and underexploited ecosystem with great microbial diversity for discovery of novel and chemically diverse antimicrobial compounds. Te goal of the study was to explore the pharmaceutical actinobacterial resources from mangrove soil and gain insight into the diversity and novelty of cultivable actinobacteria. -

Brevibacterium Sandarakinum Sp. Nov., Isolated from a Wall of an Indoor Environment
This is an author manuscript that has been accepted for publication in International Journal of Systematic and Evolutionary Microbiology, copyright Society for General Microbiology, but has not been copy-edited, formatted or proofed. Cite this article as appearing in International Journal of Systematic and Evolutionary Microbiology. This version of the manuscript may not be duplicated or reproduced, other than for personal use or within the rule of ‘Fair Use of Copyrighted Materials’ (section 17, Title 17, US Code), without permission from the copyright owner, Society for General Microbiology. The View metadata, citation and similar papers at core.ac.uk brought to you by CORE Society for General Microbiology disclaims any responsibility or liability for errors or omissions in this version of the manuscript or in any version derived from it by any other parties. The final copy-edited, published article, which is the version of record, can be found at http://ijs.sgmjournals.org,provided by Giessener Elektronische and is freely Bibliothek available without a subscription 24 months after publication. First published in: Int J Syst Evol Microbiol, 2009. 60(4) 909-913. doi:10.1099/ijs.0.014100-0 Brevibacterium sandarakinum sp. nov., isolated from a wall of an indoor environment Peter Ka¨mpfer,1 Jenny Scha¨fer,1 Nicole Lodders1 and Hans-Ju¨rgen Busse2 Correspondence 1Institut fu¨r Angewandte Mikrobiologie, Justus-Liebig-Universita¨t Giessen, D-35392 Giessen, Peter Ka¨mpfer Germany [email protected] 2Institut fu¨r Bakteriologie, Mykologie und Hygiene, Veterina¨rmedizinische Universita¨t, A-1210 Wien, giessen.de Austria A Gram-stain-positive, rod-shaped, non-endospore-forming, orange-pigmented (coloured) actinobacterium (01-Je-003T) was isolated from the wall of an indoor environment primarily colonized with moulds. -

Exploring the Diversity and Antimicrobial Potential of Marine Actinobacteria from the Comau Fjord in Northern Patagonia, Chile
ORIGINAL RESEARCH published: 19 July 2016 doi: 10.3389/fmicb.2016.01135 Exploring the Diversity and Antimicrobial Potential of Marine Actinobacteria from the Comau Fjord in Northern Patagonia, Chile Agustina Undabarrena 1, Fabrizio Beltrametti 2, Fernanda P. Claverías 1, Myriam González 1, Edward R. B. Moore 3, 4, Michael Seeger 1 and Beatriz Cámara 1* 1 Laboratorio de Microbiología Molecular y Biotecnología Ambiental, Departamento de Química & Centro de Biotecnología Daniel Alkalay Lowitt, Universidad Técnica Federico Santa María, Valparaíso, Chile, 2 Actygea S.r.l., Gerenzano, Italy, 3 Culture Collection University of Gothenburg (CCUG), Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden, 4 Department of Infectious Diseases, Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden Edited by: Bioprospecting natural products in marine bacteria from fjord environments are attractive Learn-Han Lee, due to their unique geographical features. Although, Actinobacteria are well known Monash University Malaysia Campus, Malaysia for producing a myriad of bioactive compounds, investigations regarding fjord-derived Reviewed by: marine Actinobacteria are scarce. In this study, the diversity and biotechnological Atte Von Wright, potential of Actinobacteria isolated from marine sediments within the Comau University of Eastern Finland, Finland fjord, in Northern Chilean Patagonia, were assessed by culture-based approaches. Polpass Arul Jose, Central Salt and Marine Chemicals The 16S rRNA gene sequences revealed that members phylogenetically related Research Institute, India to the Micrococcaceae, Dermabacteraceae, Brevibacteriaceae, Corynebacteriaceae, *Correspondence: Microbacteriaceae, Dietziaceae, Nocardiaceae, and Streptomycetaceae families were Beatriz Cámara [email protected] present at the Comau fjord. A high diversity of cultivable Actinobacteria (10 genera) was retrieved by using only five different isolation media. -

Gene Ortholog Metabolic Functional Gene Definition LDA
Table S1: Gene Metabolic Functional Gene Definition LDA effect p-value p-value Normospermia Semen with Ortholog size (FDR) without leucocytes Leukocytes K02025 multiple sugar transport system permease 2.261725843 0.012724 0.012752175 0.064354389 0.100693286 protein K00557 tRNA (uracil-5-)-methyltransferase 1.874490605 0.012724 0.012758059 0.015985129 0.001204857 K16087 hemoglobin/transferrin/lactoferrin 2.070152298 0.01991 0.020019188 0.025014425 0.001708239 receptor protein K02119 V/A-type H+/Na+-transporting ATPase 1.710055455 0.024682 0.024843508 0.002815807 0.012874332 subunit C K10806 acyl-CoA thioesterase YciA 1.841834075 0.030413 0.030660244 0.017565963 0.003870828 K07783 MFS transporter, OPA family, sugar 1.722774099 0.030413 0.030702979 0.011095006 0.000731635 phosphate sensor protein UhpC K02122 V/A-type H+/Na+-transporting ATPase 1.697757071 0.030413 0.030712492 0.00294449 0.012716599 subunit F K01995 branched-chain amino acid transport 2.209931713 0.037253 0.037677881 0.09816811 0.130399195 system ATP-binding protein K03808 paraquat-inducible protein A 1.69974082 0.037253 0.037698353 0.010599677 0.000781914 K03576 LysR family transcriptional regulator, 1.87590023 0.037253 0.03770128 0.019489044 0.004660097 regulator for metE and metH K03604 LacI family transcriptional regulator, purine 1.949109778 0.045361 0.046168073 0.027013054 0.009424649 nucleotide synthesis repressor K02806 PTS system, nitrogen regulatory IIA 1.980305469 0.045361 0.046186102 0.026697121 0.007783834 component K03583 exodeoxyribonuclease V gamma subunit -

UNIVERSITY of CALIFORNIA RIVERSIDE Gustatory Receptors In
UNIVERSITY OF CALIFORNIA RIVERSIDE Gustatory Receptors in Mosquito Olfaction and Host-Seeking Behavior A Dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Entomology by Genevieve Mitchell Tauxe March 2015 Dissertation Committee: Dr. Anandasankar Ray, Chairperson Dr. Michael E. Adams Dr. Ring T. Cardé Dr. Jocelyn G. Millar Copyright by Genevieve Mitchell Tauxe 2015 The Dissertation of Genevieve Mitchell Tauxe is approved: Committee Chairperson University of California, Riverside Acknowledgements First, I thank the anonymous odor donors who participated in the experiments described here. They variously washed their feet in the bathroom sink, walked around for hours with beads in their socks (which, no, is not terribly comfortable), and continually showed up no matter how many times we kept asking for more socks. Thank you for your generosity, and for your stinky feet. Now go find yourself on page 73. I thank my advisor, Anand Ray, for all of his help and support over my graduate career. He has taught me so much about how to do research, and also how to compose manuscripts, give presentations, and talk about science. I also thank my committee members, Ring Cardé, Jocelyn Millar, and Mike Adams, for offering so much helpful advice and support over the years, both during official committee meetings and on many other occasions. I thank all the Ray lab and Dahanukar lab members for making the laboratory a fun and productive place to work. In particular, Paulina Ngo performed most of the behavior tests reported in Chapter 4, Tom Guda helped with many behavioral assays, and Greg Pask helped with cloning. -

Genomic and Transcriptomic Insights Into Calcium Carbonate Biomineralization by Marine Actinobacterium Brevibacterium Linens BS258
fmicb-08-00602 April 4, 2017 Time: 15:2 # 1 ORIGINAL RESEARCH published: 06 April 2017 doi: 10.3389/fmicb.2017.00602 Genomic and Transcriptomic Insights into Calcium Carbonate Biomineralization by Marine Actinobacterium Brevibacterium linens BS258 Yuying Zhu1,2,3†, Ning Ma1,2,3†, Weihua Jin4, Shimei Wu5 and Chaomin Sun1,2* 1 Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, Qingdao, China, 2 Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao, China, 3 College of Earth Science, University of Chinese Academy of Sciences, Beijing, China, 4 College of Biotechnology and Bioengineering, Zhejiang University of Technology, Hangzhou, China, 5 College of Life Sciences, Qingdao University, Qingdao, China Calcium carbonate (CaCO3) biomineralization has been investigated due to its wide range of scientific and technological implications, however, the molecular mechanisms of this important geomicrobiological process are largely unknown. Here, a urease- Edited by: positive marine actinobacterium Brevibacterium linens BS258 was demonstrated to Qiang Wang, Institute of Hydrobiology (CAS), China effectively form CaCO3 precipitates. Surprisingly, this bacterium could also dissolve 2C Reviewed by: the formed CaCO3 with the increase of the Ca concentration. To disclose the Xiaoke Hu, mechanisms of biomineralization, the genome of B. linens BS258 was further completely Yantai Institute of Coastal Zone Research (CAS), China sequenced. Interestingly, the expression of three carbonic anhydrases was significantly C Bruno A. Martinez, up-regulated along with the increase of Ca2 concentration and the extent of calcite Lawrence Berkeley National dissolution. Moreover, transcriptome analyses revealed that increasing concentration Laboratory, USA of Ca2C induced KEGG pathways including quorum sensing (QS) in B. -

Rind Austrian Hard Cheese Surfaces and Its Production Environment
Animal Science Publications Animal Science 2-2018 Autochthonous facility-specific microbiota dominates washed- rind Austrian hard cheese surfaces and its production environment Narciso M. Quijada University of Veterinary Medicine Vienna Evelyne Mann University of Veterinary Medicine Vienna Martin Wagner University of Veterinary Medicine Vienna David Rodríguez-Lázaro Universidad de Burgos Marta Hernández Instituto Tecnológico Agrario de Castilla y León See next page for additional authors Follow this and additional works at: https://lib.dr.iastate.edu/ans_pubs Part of the Bacteriology Commons, Environmental Microbiology and Microbial Ecology Commons, Food Processing Commons, and the Genetics and Genomics Commons The complete bibliographic information for this item can be found at https://lib.dr.iastate.edu/ ans_pubs/525. For information on how to cite this item, please visit http://lib.dr.iastate.edu/ howtocite.html. This Article is brought to you for free and open access by the Animal Science at Iowa State University Digital Repository. It has been accepted for inclusion in Animal Science Publications by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Autochthonous facility-specific microbiota dominates washed-rind Austrian hard cheese surfaces and its production environment Abstract Cheese ripening involves the succession of complex microbial communities that are responsible for the organoleptic properties of the final products. The food processing environment can act as a source of natural microbial inoculation, especially in traditionally manufactured products. Austrian Vorarlberger Bergkäse (VB) is an artisanal washed-rind hard cheese produced in the western part of Austria without the addition of external ripening cultures. -

Brevibacterium from Austrian Hard Cheese Harbor a Putative Histamine Catabolism Pathway and a Plasmid for Adaptation to the Cheese Environment
Animal Science Publications Animal Science 2019 Brevibacterium from Austrian hard cheese harbor a putative histamine catabolism pathway and a plasmid for adaptation to the cheese environment Justin M. Anast Iowa State University, [email protected] Monika Dzieciol University of Veterinary Medicine Vienna Dylan L. Schultz Iowa State University, [email protected] Martin Wagner University of Veterinary Medicine Vienna Evelyne Mann University of Veterinary Medicine Vienna See next page for additional authors Follow this and additional works at: https://lib.dr.iastate.edu/ans_pubs Part of the Agriculture Commons, Animal Sciences Commons, Food Microbiology Commons, and the Genetics and Genomics Commons The complete bibliographic information for this item can be found at https://lib.dr.iastate.edu/ ans_pubs/520. For information on how to cite this item, please visit http://lib.dr.iastate.edu/ howtocite.html. This Article is brought to you for free and open access by the Animal Science at Iowa State University Digital Repository. It has been accepted for inclusion in Animal Science Publications by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Brevibacterium from Austrian hard cheese harbor a putative histamine catabolism pathway and a plasmid for adaptation to the cheese environment Abstract The genus Brevibacterium harbors many members important for cheese ripening. We performed real-time quantitative PCR (qPCR) to determine the abundance of Brevibacterium on rinds of Vorarlberger Bergkäse, an Austrian artisanal washed-rind hard cheese, over 160 days of ripening. Our results show that Brevibacterium are abundant on Vorarlberger Bergkäse rinds throughout the ripening time. -
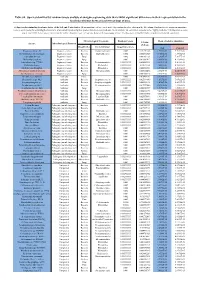
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp.