Mouse Vps45 Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RNA Sequencing Reveals a Slow to Fast Muscle Fiber Type Transition After Olanzapine Infusion in Rats
RESEARCH ARTICLE RNA Sequencing Reveals a Slow to Fast Muscle Fiber Type Transition after Olanzapine Infusion in Rats Christopher J. Lynch1*, Yuping Xu1, Andras Hajnal2, Anna C. Salzberg3, Yuka Imamura Kawasawa4,5,6 1 Department of Cellular and Molecular Physiology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 2 Department of Neural and Behavioral Sciences, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 3 Department a11111 of Public Health Sciences, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 4 Department of Pharmacology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 5 Department of Biochemistry and Molecular Biology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 6 The Institute for Personalized Medicine, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America * [email protected] OPEN ACCESS Citation: Lynch CJ, Xu Y, Hajnal A, Salzberg AC, Kawasawa YI (2015) RNA Sequencing Reveals a Abstract Slow to Fast Muscle Fiber Type Transition after Olanzapine Infusion in Rats. PLoS ONE 10(4): Second generation antipsychotics (SGAs), like olanzapine, exhibit acute metabolic side ef- e0123966. doi:10.1371/journal.pone.0123966 fects leading to metabolic inflexibility, hyperglycemia, adiposity and diabetes. Understand- Academic Editor: Guillermo López Lluch, ing how SGAs affect the skeletal muscle transcriptome could elucidate approaches for Universidad Pablo de Olavide, Centro Andaluz de mitigating these side effects. Male Sprague-Dawley rats were infused intravenously with ve- Biología del Desarrollo-CSIC, SPAIN hicle or olanzapine for 24h using a dose leading to a mild hyperglycemia. -

Mutations in the VPS45 Gene, a SEC1 Homologue, Result in Vacuolar Protein Sorting Defects and Accumulation of Membrane Vesicles
Journal of Cell Science 107, 3449-3459 (1994) 3449 Printed in Great Britain © The Company of Biologists Limited 1994 Mutations in the VPS45 gene, a SEC1 homologue, result in vacuolar protein sorting defects and accumulation of membrane vesicles Christopher R. Cowles, Scott D. Emr* and Bruce F. Horazdovsky† Division of Cellular and Molecular Medicine & Howard Hughes Medical Institute, University of California, San Diego, School of Medicine, La Jolla, California 92093-0668, USA *Author for correspondence †Present address: Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas 75235-9038, USA SUMMARY Genetic analyses of vacuolar protein sorting in Saccha- kDa protein in labeled yeast cell extracts. Subcellular frac- romyces cerevisiae have uncovered a large number of tionation studies demonstrate that the majority of Vps45p mutants (vps) that missort and secrete vacuolar hydrolases. is associated with a high-speed membrane pellet fraction A small subset of vps mutants exhibit a temperature-con- that includes Golgi, transport vesicles and, potentially, ditional growth phenotype and show a severe defect in the endosomal membranes. Significantly, this fraction lacks localization of soluble vacuolar proteins, yet maintain a ER, vacuole and plasma membranes. Overexpression of near-normal vacuole structure. Here, we report on the Vps45p saturates the sites with which Vps45p associates. A cloning and characterization of the gene affected in one of vps45 null mutant accumulates vesicles, many of which these mutants, VPS45, which has been found to encode a were found to be present in large clusters. This accumula- member of a protein family that includes the yeast proteins tion of potential transport vesicles indicates that Vps45p Sec1p, Sly1p and Vps33p, as well as n-Sec1, UNC18 and may facilitate the targeting and/or fusion of these vesicles Rop from other eukaryotic organisms. -

RAB11-Mediated Trafficking and Human Cancers: an Updated Review
biology Review RAB11-Mediated Trafficking and Human Cancers: An Updated Review Elsi Ferro 1,2, Carla Bosia 1,2 and Carlo C. Campa 1,2,* 1 Department of Applied Science and Technology, Politecnico di Torino, 24 Corso Duca degli Abruzzi, 10129 Turin, Italy; [email protected] (E.F.); [email protected] (C.B.) 2 Italian Institute for Genomic Medicine, c/o IRCCS, Str. Prov. le 142, km 3.95, 10060 Candiolo, Italy * Correspondence: [email protected] Simple Summary: The small GTPase RAB11 is a master regulator of both vesicular trafficking and membrane dynamic defining the surface proteome of cellular membranes. As a consequence, the alteration of RAB11 activity induces changes in both the sensory and the transduction apparatuses of cancer cells leading to tumor progression and invasion. Here, we show that this strictly depends on RAB110s ability to control the sorting of signaling receptors from endosomes. Therefore, RAB11 is a potential therapeutic target over which to develop future therapies aimed at dampening the acquisition of aggressive traits by cancer cells. Abstract: Many disorders block and subvert basic cellular processes in order to boost their pro- gression. One protein family that is prone to be altered in human cancers is the small GTPase RAB11 family, the master regulator of vesicular trafficking. RAB11 isoforms function as membrane organizers connecting the transport of cargoes towards the plasma membrane with the assembly of autophagic precursors and the generation of cellular protrusions. These processes dramatically impact normal cell physiology and their alteration significantly affects the survival, progression and metastatization as well as the accumulation of toxic materials of cancer cells. -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Fall 2010 Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Renuka Nayak University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Computational Biology Commons, and the Genomics Commons Recommended Citation Nayak, Renuka, "Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress" (2010). Publicly Accessible Penn Dissertations. 1559. https://repository.upenn.edu/edissertations/1559 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1559 For more information, please contact [email protected]. Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Abstract Genes interact in networks to orchestrate cellular processes. Here, we used coexpression networks based on natural variation in gene expression to study the functions and interactions of human genes. We asked how these networks change in response to stress. First, we studied human coexpression networks at baseline. We constructed networks by identifying correlations in expression levels of 8.9 million gene pairs in immortalized B cells from 295 individuals comprising three independent samples. The resulting networks allowed us to infer interactions between biological processes. We used the network to predict the functions of poorly-characterized human genes, and provided some experimental support. Examining genes implicated in disease, we found that IFIH1, a diabetes susceptibility gene, interacts with YES1, which affects glucose transport. Genes predisposing to the same diseases are clustered non-randomly in the network, suggesting that the network may be used to identify candidate genes that influence disease susceptibility. -

PDF, Also Known As Version of Record
King’s Research Portal DOI: 10.1038/s41467-018-03283-z Document Version Publisher's PDF, also known as Version of record Link to publication record in King's Research Portal Citation for published version (APA): Patel, N., Weekes, D., Drosopoulos, K., Gazinska, P., Noel, E., Rashid, M., Mirza, H., Quist, J., Brasó-Maristany, F., Mathew, S., Ferro, R., Pereira, A. M., Prince, C., Noor, F., Francesch-Domenech, E., Marlow, R., de Rinaldis, E., Grigoriadis, A., Linardopoulos, S., ... Tutt, A. N. J. (2018). Integrated genomics and functional validation identifies malignant cell specific dependencies in triple negative breast cancer. Nature Communications, 9(1), [1044]. https://doi.org/10.1038/s41467-018-03283-z Citing this paper Please note that where the full-text provided on King's Research Portal is the Author Accepted Manuscript or Post-Print version this may differ from the final Published version. If citing, it is advised that you check and use the publisher's definitive version for pagination, volume/issue, and date of publication details. And where the final published version is provided on the Research Portal, if citing you are again advised to check the publisher's website for any subsequent corrections. General rights Copyright and moral rights for the publications made accessible in the Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognize and abide by the legal requirements associated with these rights. •Users may download and print one copy of any publication from the Research Portal for the purpose of private study or research. -

Network Mining Approach to Cancer Biomarker Discovery
NETWORK MINING APPROACH TO CANCER BIOMARKER DISCOVERY THESIS Presented in Partial Fulfillment of the Requirements for the Degree Master of Science in the Graduate School of The Ohio State University By Praneeth Uppalapati, B.E. Graduate Program in Computer Science and Engineering The Ohio State University 2010 Thesis Committee: Dr. Kun Huang, Advisor Dr. Raghu Machiraju Copyright by Praneeth Uppalapati 2010 ABSTRACT With the rapid development of high throughput gene expression profiling technology, molecule profiling has become a powerful tool to characterize disease subtypes and discover gene signatures. Most existing gene signature discovery methods apply statistical methods to select genes whose expression values can differentiate different subject groups. However, a drawback of these approaches is that the selected genes are not functionally related and hence cannot reveal biological mechanism behind the difference in the patient groups. Gene co-expression network analysis can be used to mine functionally related sets of genes that can be marked as potential biomarkers through survival analysis. We present an efficient heuristic algorithm EigenCut that exploits the properties of gene co- expression networks to mine functionally related and dense modules of genes. We apply this method to brain tumor (Glioblastoma Multiforme) study to obtain functionally related clusters. If functional groups of genes with predictive power on patient prognosis can be identified, insights on the mechanisms related to metastasis in GBM can be obtained and better therapeutical plan can be developed. We predicted potential biomarkers by dividing the patients into two groups based on their expression profiles over the genes in the clusters and comparing their survival outcome through survival analysis. -
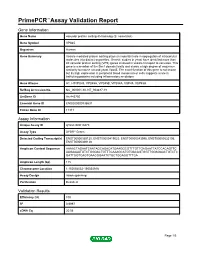
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name vacuolar protein sorting 45 homolog (S. cerevisiae) Gene Symbol VPS45 Organism Human Gene Summary Vesicle mediated protein sorting plays an important role in segregation of intracellular molecules into distinct organelles. Genetic studies in yeast have identified more than 40 vacuolar protein sorting (VPS) genes involved in vesicle transport to vacuoles. This gene is a member of the Sec1 domain family and shows a high degree of sequence similarity to mouse rat and yeast Vps45. The exact function of this gene is not known but its high expression in peripheral blood mononuclear cells suggests a role in trafficking proteins including inflammatory mediators. Gene Aliases H1, H1VPS45, VPS45A, VPS45B, VPS54A, VSP45, VSP45A RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.443750 Ensembl Gene ID ENSG00000136631 Entrez Gene ID 11311 Assay Information Unique Assay ID qHsaCID0010473 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000369130, ENST00000419023, ENST00000543996, ENST00000535106, ENST00000369128 Amplicon Context Sequence AAAACTAGAATCAATAGCAGACATGAAGGCGTTTGTTGAGAATTATCCACAGTTC AAGAAAATGTCTGGGACTGTTTCAAAGCATGTGACAGTGGTTGGAGAACTGTCTC GATTGGTCAGTGAACGGAATCTGCTGGAGGTTTCA Amplicon Length (bp) 115 Chromosome Location 1:150054052-150054916 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9997 cDNA Cq 20.56 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 81 gDNA Cq Specificity (%) 100 Information to assist -
A Generalized Gene Set Data Mining Approach
Journal of Clinical Medicine Article Comorbidities and Susceptibility to COVID-19: A Generalized Gene Set Data Mining Approach Micaela F. Beckman, Farah Bahrani Mougeot * and Jean-Luc C. Mougeot * Department of Oral Medicine, Carolinas Medical Center, Atrium Health, Charlotte, NC 28203, USA; [email protected] * Correspondence: [email protected] (F.B.M.); [email protected] (J.-L.C.M.) Abstract: The COVID-19 pandemic has led to over 2.26 million deaths for almost 104 million confirmed cases worldwide, as of 4 February 2021 (WHO). Risk factors include pre-existing conditions such as cancer, cardiovascular disease, diabetes, and obesity. Although several vaccines have been deployed, there are few alternative anti-viral treatments available in the case of reduced or non- existent vaccine protection. Adopting a long-term holistic approach to cope with the COVID-19 pandemic appears critical with the emergence of novel and more infectious SARS-CoV-2 variants. Our objective was to identify comorbidity-associated single nucleotide polymorphisms (SNPs), potentially conferring increased susceptibility to SARS-CoV-2 infection using a computational meta-analysis approach. SNP datasets were downloaded from a publicly available genome-wide association studies (GWAS) catalog for 141 of 258 candidate COVID-19 comorbidities. Gene-level SNP analysis was performed to identify significant pathways by using the program MAGMA. An SNP annotation program was used to analyze MAGMA-identified genes. Differential gene expression was determined for significant genes across 30 general tissue types using the Functional and Annotation Mapping of GWAS online tool GENE2FUNC. COVID-19 comorbidities (n = 22) from six disease categories Citation: Beckman, M.F.; Mougeot, were found to have significant associated pathways, validated by Q–Q plots (p < 0.05). -

1 Supplementary Material Figure S1. Volcano Plot of Differentially
Supplementary material Figure S1. Volcano Plot of differentially expressed genes between preterm infants fed own mother’s milk (OMM) or pasteurized donated human milk (DHM). Table S1. The 10 most representative biological processes filtered for enrichment p- value in preterm infants. Biological Processes p-value Quantity of DEG* Transcription, DNA-templated 3.62x10-24 189 Regulation of transcription, DNA-templated 5.34x10-22 188 Transport 3.75x10-17 140 Cell cycle 1.03x10-13 65 Gene expression 3.38x10-10 60 Multicellular organismal development 6.97x10-10 86 1 Protein transport 1.73x10-09 56 Cell division 2.75x10-09 39 Blood coagulation 3.38x10-09 46 DNA repair 8.34x10-09 39 Table S2. Differential genes in transcriptomic analysis of exfoliated epithelial intestinal cells between preterm infants fed own mother’s milk (OMM) and pasteurized donated human milk (DHM). Gene name Gene Symbol p-value Fold-Change (OMM vs. DHM) (OMM vs. DHM) Lactalbumin, alpha LALBA 0.0024 2.92 Casein kappa CSN3 0.0024 2.59 Casein beta CSN2 0.0093 2.13 Cytochrome c oxidase subunit I COX1 0.0263 2.07 Casein alpha s1 CSN1S1 0.0084 1.71 Espin ESPN 0.0008 1.58 MTND2 ND2 0.0138 1.57 Small ubiquitin-like modifier 3 SUMO3 0.0037 1.54 Eukaryotic translation elongation EEF1A1 0.0365 1.53 factor 1 alpha 1 Ribosomal protein L10 RPL10 0.0195 1.52 Keratin associated protein 2-4 KRTAP2-4 0.0019 1.46 Serine peptidase inhibitor, Kunitz SPINT1 0.0007 1.44 type 1 Zinc finger family member 788 ZNF788 0.0000 1.43 Mitochondrial ribosomal protein MRPL38 0.0020 1.41 L38 Diacylglycerol -

A Yeast Lysosomal Biogenesis Map Uncovers the Cargo Spectrum of Lysosomal Protein Targeting Pathways
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.24.453616; this version posted July 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A yeast lysosomal biogenesis map uncovers the cargo spectrum of lysosomal protein targeting pathways Sebastian Eising1, Bianca M. Esch1, Mike Wälte2, Prado Vargas Duarte3, Stefan Walter4, Christian Ungermann3,4, Maria Bohnert2,5, Florian Fröhlich1,3# 1 Osnabrück University Department of Biology/Chemistry Molecular Membrane Biology Group Barbarastraße 13 49076 Osnabrück, Germany 2 University of Münster Institute of Cell Dynamics and Imaging, Von-Esmarch-Straße 56 48149 Münster, Germany 3 Osnabrück University Department of Biology/Chemistry Biochemistry section Barbarastrasse 13 49076 Osnabrück, Germany 4 Osnabrück University Center of Cellular Nanoanalytics Osnabrück (CellNanOs) Barbarastrasse 11 49076 Osnabrück, Germany 5 University of Münster, Cells in Motion Interfaculty Centre (CiM), Waldeyerstr. 15 48149 Münster, Germany # Corresponding author: Email: [email protected] (F. Fröhlich) Phone: +49-541-969-2794 (F.F.) Keywords: Lysosomal biogenesis, vacuole, CPY pathway, CVT pathway, AP-3 trafficking bioRxiv preprint doi: https://doi.org/10.1101/2021.07.24.453616; this version posted July 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The lysosome is the major catabolic organelle and a key metabolic signaling center of the cell. Mutations in lysosomal proteins can have catastrophic effects, causing neurodegeneration, cancer, and age-related diseases. The vacuole is the lysosomal analog of Saccharomyces cerevisiae that harbors many conserved proteins. -

Identification and Characterisation of the Underlying Defects in Patients with Inherited Platelet Bleeding Disorders
Identification and Characterisation of the Underlying Defects in Patients with Inherited Platelet Bleeding Disorders By: Maryam Ahmed Aldossary A thesis submitted in partial fulfilment of the requirements for the degree of Doctor of Philosophy The University of Sheffield Faculty of Medicine, Dentistry and Health Department of Infection, Immunity & Cardiovascular Disease Submission Date January 2019 Abstract The underlying genetic defects remain unknown in about 50% of inherited platelet bleeding disorders (IPDs). This study investigated the use of whole exome sequencing (WES) to identify candidate gene defects in 34 index cases enrolled in the UK Genotyping and Phenotyping of Platelets study with a history of bleeding, whose platelets demonstrated defects in agonist-induced dense granule secretion or Gi- signalling. WES analysis identified a median of 98 candidate disease-causing variants per index case highlighting the complexity of IPDs. Sixteen variants were in genes that had previously been associated with IPDs, two of which were selected for further characterisation. Two novel FLI1 alterations, predicting p.Arg340Cys/His substitutions in the DNA binding domain of FLI1 were shown to reduce transcriptional activity and nuclear accumulation of FLI1, suggesting that these variants interfere with the regulation of essential megakaryocyte genes by FLI1 and may explain the bleeding tendency in affected patients. Expression of a novel truncated p.Arg430* variant of ETV6 revealed it to be stably expressed, possessing normal repressor activity in HEK 293T cells and a slight reduction in repressor activity in megakaryocytic cells. Further studies are required to confirm the pathogenicity of this variant. To identify novel genes involved in platelet granule biogenesis and secretion, gene expression was examined in megakaryocytic cells before and after knockdown of FLI1, defects in which are associated with platelet granule abnormalities.