The Zebrafish Presomitic Mesoderm Elongates Through Compression-Extension
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

3 Embryology and Development
BIOL 6505 − INTRODUCTION TO FETAL MEDICINE 3. EMBRYOLOGY AND DEVELOPMENT Arlet G. Kurkchubasche, M.D. INTRODUCTION Embryology – the field of study that pertains to the developing organism/human Basic embryology –usually taught in the chronologic sequence of events. These events are the basis for understanding the congenital anomalies that we encounter in the fetus, and help explain the relationships to other organ system concerns. Below is a synopsis of some of the critical steps in embryogenesis from the anatomic rather than molecular basis. These concepts will be more intuitive and evident in conjunction with diagrams and animated sequences. This text is a synopsis of material provided in Langman’s Medical Embryology, 9th ed. First week – ovulation to fertilization to implantation Fertilization restores 1) the diploid number of chromosomes, 2) determines the chromosomal sex and 3) initiates cleavage. Cleavage of the fertilized ovum results in mitotic divisions generating blastomeres that form a 16-cell morula. The dense morula develops a central cavity and now forms the blastocyst, which restructures into 2 components. The inner cell mass forms the embryoblast and outer cell mass the trophoblast. Consequences for fetal management: Variances in cleavage, i.e. splitting of the zygote at various stages/locations - leads to monozygotic twinning with various relationships of the fetal membranes. Cleavage at later weeks will lead to conjoined twinning. Second week: the week of twos – marked by bilaminar germ disc formation. Commences with blastocyst partially embedded in endometrial stroma Trophoblast forms – 1) cytotrophoblast – mitotic cells that coalesce to form 2) syncytiotrophoblast – erodes into maternal tissues, forms lacunae which are critical to development of the uteroplacental circulation. -

From Bipotent Neuromesodermal Progenitors to Neural-Mesodermal Interactions During Embryonic Development
International Journal of Molecular Sciences Review From Bipotent Neuromesodermal Progenitors to Neural-Mesodermal Interactions during Embryonic Development Nitza Kahane and Chaya Kalcheim * Department of Medical Neurobiology, Institute of Medical Research Israel-Canada (IMRIC) and the Edmond and Lily Safra Center for Brain Sciences (ELSC), Hebrew University of Jerusalem-Hadassah Medical School, P.O. Box 12272, Jerusalem 9112102, Israel; [email protected] * Correspondence: [email protected] Abstract: To ensure the formation of a properly patterned embryo, multiple processes must operate harmoniously at sequential phases of development. This is implemented by mutual interactions between cells and tissues that together regulate the segregation and specification of cells, their growth and morphogenesis. The formation of the spinal cord and paraxial mesoderm derivatives exquisitely illustrate these processes. Following early gastrulation, while the vertebrate body elongates, a pop- ulation of bipotent neuromesodermal progenitors resident in the posterior region of the embryo generate both neural and mesodermal lineages. At later stages, the somitic mesoderm regulates aspects of neural patterning and differentiation of both central and peripheral neural progenitors. Reciprocally, neural precursors influence the paraxial mesoderm to regulate somite-derived myogen- esis and additional processes by distinct mechanisms. Central to this crosstalk is the activity of the axial notochord, which, via sonic hedgehog signaling, plays pivotal roles in neural, skeletal muscle and cartilage ontogeny. Here, we discuss the cellular and molecular basis underlying this complex Citation: Kahane, N.; Kalcheim, C. developmental plan, with a focus on the logic of sonic hedgehog activities in the coordination of the From Bipotent Neuromesodermal Progenitors to Neural-Mesodermal neural-mesodermal axis. -

The Genetic Basis of Mammalian Neurulation
REVIEWS THE GENETIC BASIS OF MAMMALIAN NEURULATION Andrew J. Copp*, Nicholas D. E. Greene* and Jennifer N. Murdoch‡ More than 80 mutant mouse genes disrupt neurulation and allow an in-depth analysis of the underlying developmental mechanisms. Although many of the genetic mutants have been studied in only rudimentary detail, several molecular pathways can already be identified as crucial for normal neurulation. These include the planar cell-polarity pathway, which is required for the initiation of neural tube closure, and the sonic hedgehog signalling pathway that regulates neural plate bending. Mutant mice also offer an opportunity to unravel the mechanisms by which folic acid prevents neural tube defects, and to develop new therapies for folate-resistant defects. 6 ECTODERM Neurulation is a fundamental event of embryogenesis distinct locations in the brain and spinal cord .By The outer of the three that culminates in the formation of the neural tube, contrast, the mechanisms that underlie the forma- embryonic (germ) layers that which is the precursor of the brain and spinal cord. A tion, elevation and fusion of the neural folds have gives rise to the entire central region of specialized dorsal ECTODERM, the neural plate, remained elusive. nervous system, plus other organs and embryonic develops bilateral neural folds at its junction with sur- An opportunity has now arisen for an incisive analy- structures. face (non-neural) ectoderm. These folds elevate, come sis of neurulation mechanisms using the growing battery into contact (appose) in the midline and fuse to create of genetically targeted and other mutant mouse strains NEURAL CREST the neural tube, which, thereafter, becomes covered by in which NTDs form part of the mutant phenotype7.At A migratory cell population that future epidermal ectoderm. -

Understanding Paraxial Mesoderm Development and Sclerotome Specification for Skeletal Repair Shoichiro Tani 1,2, Ung-Il Chung2,3, Shinsuke Ohba4 and Hironori Hojo2,3
Tani et al. Experimental & Molecular Medicine (2020) 52:1166–1177 https://doi.org/10.1038/s12276-020-0482-1 Experimental & Molecular Medicine REVIEW ARTICLE Open Access Understanding paraxial mesoderm development and sclerotome specification for skeletal repair Shoichiro Tani 1,2, Ung-il Chung2,3, Shinsuke Ohba4 and Hironori Hojo2,3 Abstract Pluripotent stem cells (PSCs) are attractive regenerative therapy tools for skeletal tissues. However, a deep understanding of skeletal development is required in order to model this development with PSCs, and for the application of PSCs in clinical settings. Skeletal tissues originate from three types of cell populations: the paraxial mesoderm, lateral plate mesoderm, and neural crest. The paraxial mesoderm gives rise to the sclerotome mainly through somitogenesis. In this process, key developmental processes, including initiation of the segmentation clock, formation of the determination front, and the mesenchymal–epithelial transition, are sequentially coordinated. The sclerotome further forms vertebral columns and contributes to various other tissues, such as tendons, vessels (including the dorsal aorta), and even meninges. To understand the molecular mechanisms underlying these developmental processes, extensive studies have been conducted. These studies have demonstrated that a gradient of activities involving multiple signaling pathways specify the embryonic axis and induce cell-type-specific master transcription factors in a spatiotemporal manner. Moreover, applying the knowledge of mesoderm development, researchers have attempted to recapitulate the in vivo development processes in in vitro settings, using mouse and human PSCs. In this review, we summarize the state-of-the-art understanding of mesoderm development and in vitro modeling of mesoderm development using PSCs. We also discuss future perspectives on the use of PSCs to generate skeletal tissues for basic research and clinical applications. -
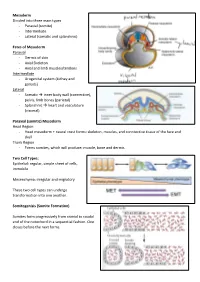
Mesoderm Divided Into Three Main Types - Paraxial (Somite) - Intermediate - Lateral (Somatic and Splanchnic)
Mesoderm Divided into three main types - Paraxial (somite) - Intermediate - Lateral (somatic and splanchnic) Fates of Mesoderm Paraxial - Dermis of skin - Axial Skeleton - Axial and limb muscles/tendons Intermediate - Urogenital system (kidney and gonads) Lateral - Somatic inner body wall (connective), pelvis, limb bones (parietal) - Splanchnic heart and vasculature (visceral) Paraxial (somitic) Mesoderm Head Region - Head mesoderm + neural crest forms: skeleton, muscles, and conntective tissue of the face and skull Trunk Region - Forms somites, which will produce: muscle, bone and dermis Two Cell Types: Epithelial: regular, simple sheet of cells, immobile Mesenchyma: irregular and migratory These two cell types can undergo transformation into one another. Somitogenisis (Somite Formation) Somites form progressively from cranial to caudal end of the notochord in a sequential fashion. One closes before the next forms. Somite Differentiation The somite splits into the epithelial dermamyotome (dermis/muscle) and the messenchymal sclerotome (skeletal). The somite is all paraxial mesoderm. Somite location determines the fate of its associates derma/myo/sclerotomes. Intermediate Mesoderm Urogenital system: - Kidneys - Gonads - Reproductive Duct Systems Runs alongside the paraxial mesoderm. Urogenital System Along mesonephric duct: - Pronephros, mesonephros, and metanephros - Pronephros fall away as gonad develops on ventral-medial side of mesonephros. - Metanephrogenic mesenchyme gives rise to kidney. The mesonephric duct will become the Wolffian duct forming at the nephric bud. The Mullerian duct forms via an invagination on the dorsal side of the nephric duct. The gonad will degenerate one of the two ducts depending on the hormones it produces. XX degenerates Wolffian duct – no testosterone, anti-Mullerian hormone (AMH) not produced, and Mullerian duct can develop in addition to female reproductive organs (ovaries, vagina) XY degenerates Mullerian duct – testosterone, AMH produced, Wolffian duct continues as male reproductive organs (testes, penis) develop. -

The Migration of Paraxial and Lateral Plate Mesoderm Cells Emerging from the Late Primitive Streak Is Controlled by Different Wnt Signals
The Migration of Paraxial and Lateral Plate Mesoderm Cells Emerging from the Late Primitive Streak is Controlled by Different Wnt Signals The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Sweetman, Dylan, Laura Wagstaff, Oliver Cooper, Cornelis Weijer, and Andrea Münsterberg. 2008. The migration of paraxial and lateral plate mesoderm cells emerging from the late primitive streak is controlled by different Wnt signals. BMC Developmental Biology 8: 63. Published Version doi:10.1186/1471-213X-8-63 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:4774184 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA BMC Developmental Biology BioMed Central Research article Open Access The migration of paraxial and lateral plate mesoderm cells emerging from the late primitive streak is controlled by different Wnt signals Dylan Sweetman1, Laura Wagstaff2, Oliver Cooper1,3, Cornelis Weijer4 and Andrea Münsterberg*1 Address: 1School of Biological Sciences, University of East Anglia, Norwich Research Park, Norwich, NR4 7TJ, UK, 2Biomedical Research Centre, University of East Anglia, Norwich Research Park, Norwich, NR4 7TJ, UK, 3Centre for Neuroregeneration Research, Harvard Medical School, McLean Hospital, Belmont, MA 02478, USA and 4College of Life Sciences -

12. Development of Axial Skeleton and Extremities. Muscles and Skin
Z. Tonar, M. Králíčková: Outlines of lectures on embryology for 2 nd year students of General medicine and Dentistry License Creative Commons - http://creativecommons.org/licenses/by-nc-nd/3.0/ 12. Development of axial skeleton and extremities. Muscles and skin. Timeline − 19 days: somites emerge in the gastrula − 4 weeks: sclerotome cells migrate along the neural tube − 5 weeks: mesenchymal blastema of the axial skeleton − 6 weeks: mesenchymal blastema of limbs − 8 weeks: rotation of the limbs − 8 weeks: individual muscles differentiate − 10 weeks: primary ossification centres in diaphyses − 3 months: bones of the skull develop − 9 months: diaphyses ossified; secondary ossification centres emerge In general, bone tissue originates from: − the of the somitic paraxial mesoderm, namely from the ventromedial part, the sclerotome − the head non-segmented mesoderm − the somatopleuric lateral plate mesoderm skeleton of limbs − the neural crest, which differentiates into the head ectomesenchyme − mesenchyme o its cells migrate and differentiate into fibroblasts, the source of the desmogenous (intramembranous) ossification o its cells imgrate and differentiate into chondroblasts, the source of the chondrogenous ossification of the hyaline cartilage models Limbs − week 4: limb buds o somatopleuric mesenchyme differentiates into bones and connective tissues o myogenic cells, angioblasts and nerves grow in o surface ectoderm thickens into the apical ectodermal ridge − week 6: each limb is divided proximodistally into three components: o autopod -

The Dorsal Neural Tube Organizes the Dermamyotome and Induces Axial Myocytes in the Avian Embryo
Development 122, 231-241 (1996) 231 Printed in Great Britain © The Company of Biologists Limited 1996 DEV3253 The dorsal neural tube organizes the dermamyotome and induces axial myocytes in the avian embryo Martha S. Spence1, Joseph Yip2 and Carol A. Erickson1,* 1Section of Molecular and Cellular Biology, University of California, Davis, CA 95616, USA 2Department of Neurobiology, School of Medicine, University of Pittsburgh, Pittsburgh, PA 15261, USA *Author for correspondence (e-mail: [email protected]) SUMMARY Somites, like all axial structures, display dorsoventral reported previously to induce sclerotome. Thus, we have polarity. The dorsal portion of the somite forms the der- demonstrated that in the context of the embryonic envi- mamyotome, which gives rise to the dermis and axial mus- ronment, a dorsalizing signal from the dorsal neural tube culature, whereas the ventromedial somite disperses to can compete with the diffusible ventralizing signal from the generate the sclerotome, which later comprises the notochord. vertebrae and intervertebral discs. Although the neural In contrast to dorsal neural tube, pieces of ventral neural tube and notochord are known to regulate some aspects of tube, dorsal ectoderm or neural crest cells, all of which this dorsoventral pattern, the precise tissues that initially have been postulated to control dermamyotome formation specify the dermamyotome, and later the myotome from it, or to induce myogenesis, either fail to do so or provoke only have been controversial. Indeed, dorsal and ventral neural minimal inductive responses in any of our assays. However, tube, notochord, ectoderm and neural crest cells have all complicating the issue, we find consistent with previous been proposed to influence dermamyotome formation or to studies that following ablation of the entire neural tube, regulate myocyte differentiation. -

Early Embryonic Development Till Gastrulation (Humans)
Gargi College Subject: Comparative Anatomy and Developmental Biology Class: Life Sciences 2 SEM Teacher: Dr Swati Bajaj Date: 17/3/2020 Time: 2:00 pm to 3:00 pm EARLY EMBRYONIC DEVELOPMENT TILL GASTRULATION (HUMANS) CLEAVAGE: Cleavage in mammalian eggs are among the slowest in the animal kingdom, taking place some 12-24 hours apart. The first cleavage occurs along the journey of the embryo from oviduct toward the uterus. Several features distinguish mammalian cleavage: 1. Rotational cleavage: the first cleavage is normal meridional division; however, in the second cleavage, one of the two blastomeres divides meridionally and the other divides equatorially. 2. Mammalian blastomeres do not all divide at the same time. Thus the embryo frequently contains odd numbers of cells. 3. The mammalian genome is activated during early cleavage and zygotically transcribed proteins are necessary for cleavage and development. (In humans, the zygotic genes are activated around 8 cell stage) 4. Compaction: Until the eight-cell stage, they form a loosely arranged clump. Following the third cleavage, cell adhesion proteins such as E-cadherin become expressed, and the blastomeres huddle together and form a compact ball of cells. Blatocyst: The descendents of the large group of external cells of Morula become trophoblast (trophoblast produce no embryonic structure but rather form tissues of chorion, extraembryonic membrane and portion of placenta) whereas the small group internal cells give rise to Inner Cell mass (ICM), (which will give rise to embryo proper). During the process of cavitation, the trophoblast cells secrete fluid into the Morula to create blastocoel. As the blastocoel expands, the inner cell mass become positioned on one side of the ring of trophoblast cells, resulting in the distinctive mammalian blastocyst. -

Formation of Germ Layers (Second & Third Week of Development)
8.12.2014 Formation of Germ Layers (Second & Third week of Development) Dr. Archana Rani Associate Professor Department of Anatomy KGMU UP, Lucknow Day 8 • Blastocyst is partially embedded in the endometrial stroma. • Trophoblast differentiates into 2 layers: (i) Cytotrophoblast (ii) Syncytiotrophoblast • Cytotrophoblast shows mitotic division. Day 8 • Cells of inner cell mass (embryoblast) also differentiate into 2 layers: (i) Hypoblast layer (ii) Epiblast layer • Formation of amniotic cavity and embryonic disc. Day 9 • The blastocyst is more deeply embedded in the endometrium. • The penetration defect in the surface epithelium is closed by a fibrin coagulum. Day 9 • Large no. of vacuoles appear in syncytiotrophoblast which fuse to form lacunae which contains embryotroph. Day 9 • Hypoblast forms the roof of the exocoelomic cavity (primary yolk sac). • Heuser’s (exocoelomic membrane) • Extraembryonic mesoderm Day 11 & 12 • Formation of lacunar networks • Extraembryonic coelom (chorionic cavity) • Extraembryonic somatic mesoderm • Extraembryonic splanchnic mesoderm • Chorion Day 13 • Implantation bleeding • Villous structure of trophoblast. • Formation of Primary villi • Secondary (definitive) yolk sac • Chorionic plate (extraembronic mesoderm with cytotrophoblast) Third week of Development • Gastrulation (formation of all 3 germ layers) • Formation of primitive streak • Formation of notochord • Differentiation of 3 germ layers from Bilaminar to Trilaminar germ disc Formation of Primitive Streak (PS) • First sign of gastrulation • On 15th day • Primitive node • Primitive pit • Formation of mesenchyme on 16th day • Formation of embryonic endoderm • Intraembryonic mesoderm • Ectoderm • Epiblast is the source of all 3 germ layers Fate of Primitive Streak • Continues to form mesodermal cells upto early part of 4th week • Normally, the PS degenerates & diminishes in size. -

Junctional Neurulation: a Unique Developmental Program Shaping a Discrete Region of the Spinal Cord Highly Susceptible to Neural Tube Defects
13208 • The Journal of Neuroscience, September 24, 2014 • 34(39):13208–13221 Development/Plasticity/Repair Junctional Neurulation: A Unique Developmental Program Shaping a Discrete Region of the Spinal Cord Highly Susceptible to Neural Tube Defects Alwyn Dady,1,2 X Emmanuelle Havis,1,2 Virginie Escriou,3 Martin Catala,1,2,4 and Jean-Loup Duband1,2 1Universite´ Pierre et Marie Curie-Paris 6, Laboratoire de Biologie du De´veloppement, 75005 Paris, France, 2Centre National de la Recherche Scientifique, Laboratoire de Biologie du De´veloppement, 75005 Paris, France, 3Unite´ de Technologies Chimiques et Biologiques pour la Sante´, Centre National de la Recherche Scientifique Unite´ Mixte de Recherche 8258, Institut National de la Sante´ et de la Recherche Me´dicale U1022, Universite´ Paris Descartes, Faculte´ de Pharmacie, 75006 Paris, France, and 4Assistance Publique-Hopitaux de Paris, Federation of Neurology, Groupe Hospitalier Pitie´-Salpeˆtrie`re, 75013 Paris, France In higher vertebrates, the primordium of the nervous system, the neural tube, is shaped along the rostrocaudal axis through two consecutive, radically different processes referred to as primary and secondary neurulation. Failures in neurulation lead to severe anomalies of the nervous system, called neural tube defects (NTDs), which are among the most common congenital malformations in humans. Mechanisms causing NTDs in humans remain ill-defined. Of particular interest, the thoracolumbar region, which encompasses many NTD cases in the spine, corresponds to the junction between primary and secondary neurulations. Elucidating which developmen- tal processes operate during neurulation in this region is therefore pivotal to unraveling the etiology of NTDs. Here, using the chick embryo as a model, we show that, at the junction, the neural tube is elaborated by a unique developmental program involving concerted movements of elevation and folding combined with local cell ingression and accretion. -
Mesoderma Differenciálódása
Differentiation of mesoderm Dr. Bódi Ildikó 2019.12.04. 36. Germ cells, fertilization, cleavage (the division of cells in the early embryo) 37. Blastulation, implantation, decidua 38. Development of the embryo shield, ectoderm, endoderm and mesoderm 2 weeks The two-layer embryo shield: epibast and hypoblast Extra-embryonic Coelom Coelom = embryonic body cavity Implantation and gastrulation Gastrulation: The blastula continues to develop, eventually forming a structure called the gastrula. (three-plate embryo shield!!) The extraembryonic mesoderm appears Gastrulation THE DEVELOPMENT OF MESODERMA, DERIVATIVES OF GERM LAYERS (Weeks 3-8 of Development) The TRILAMINAR EMBRYO – 3rd week 1. Induction from the hypolblast cells 2. Proliferation of the epiblast cells 3. Caudal forms the primitive streak with primitive node and primitive groove Heart field (in deep) NEURAL ECTODERM Endoderm 4. The proliferated epiblast cells Epidermis migrate into the 2 layers Medial region of somites Lateral region of 5. Forming the mesoderm somites Intraembrional TRILAMINAR EMBRYO: mesoderm ectoderm, mesoderm, endoderm Stem cells Extraembrional mesoderm chorda dorsalis Parts of Mesoderm Paraxialis mesoderm - somita Intermedier mesoderm gononephrotom Parietalis mesoderm somatopleura splanchnopleura PARTS OF MESODERM AXIAL - green PARAXIAL-yellow INTERMEDIER - red LATERAL - blue •Paraxial mesoderm - somites - musculoskeletal structures •Intermediate mesoderm - urogenital (kidney and genital) •Lateral plate mesoderm - body wall, body cavities, cardiovascular and GIT structures •Paraxial mesoderm - somites - musculoskeletal structures •Intermediate mesoderm - urogenital (kidney and genital) •Lateral plate mesoderm - body wall, body cavities, cardiovascular and GIT structures Week 4Scanning electron micrograph of a cross-section of a human embryo at week 4 (stage 11). Note the mesoderm structures now present and their relative position and size within the embryo.