The Intein of the Thermoplasma A-Atpase a Subunit: Structure, Evolution and Expression in E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression of Cyclomaltodextrinase Gene from Bacillus Halodurans C-125 and Characterization of Its Multisubstrate Specificity
Food Sci. Biotechnol. Vol. 18, No. 3, pp. 776 ~ 781 (2009) ⓒ The Korean Society of Food Science and Technology Expression of Cyclomaltodextrinase Gene from Bacillus halodurans C-125 and Characterization of Its Multisubstrate Specificity Hye-Jeong Kang, Chang-Ku Jeong, Myoung-Uoon Jang, Seung-Ho Choi, Min-Hong Kim1, Jun-Bae Ahn2, Sang-Hwa Lee3, Sook-Ja Jo3, and Tae-Jip Kim* Department of Food Science and Technology, Chungbuk National University, Cheongju, Chungbuk 361-763, Korea 1MH2 Biochemical Co., Ltd., Eumseong, Chungbuk 369-841, Korea 2Department of Food Service Industry, Seowon University, Cheongju, Chungbuk 361-741, Korea 3Department of Food and Nutrition, Seowon University, Cheongju, Chungbuk 361-741, Korea Abstract A putative cyclomaltodextrinase (BHCD) gene was found from the genome of Bacillus halodurans C-125, which encodes 578 amino acids with a predicted molecular mass of 67,279 Da. It shares 42-59% of amino acid sequence identity with common cyclomaltodextrinase (CDase)-family enzymes. The corresponding gene was cloned by polymerase chain reaction (PCR) and the dimeric enzyme with C-terminal 6-histidines was successfully overproduced and purified from recombinant Escherichia coli. BHCD showed the highest activity against β-CD at pH 7.0 and 50oC. Due to its versatile hydrolysis and transglycosylation activities, BHCD has been confirmed as a member of CDases. However, BHCD can be distinguished from other typical CDases on the basis of its novel multisubstrate specificity. While typical CDases have over 10 times higher activity on β-CD than starch or pullulan, the CD-hydrolyzing activity of BHCD is only 2.3 times higher than pullulan. -

Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma Divulgatum
microorganisms Article Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma divulgatum Rafael Bargiela 1 , Karin Lanthaler 1,2, Colin M. Potter 1,2 , Manuel Ferrer 3 , Alexander F. Yakunin 1,2, Bela Paizs 1,2, Peter N. Golyshin 1,2 and Olga V. Golyshina 1,2,* 1 School of Natural Sciences, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK; [email protected] (R.B.); [email protected] (K.L.); [email protected] (C.M.P.); [email protected] (A.F.Y.); [email protected] (B.P.); [email protected] (P.N.G.) 2 Centre for Environmental Biotechnology, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK 3 Systems Biotechnology Group, Department of Applied Biocatalysis, CSIC—Institute of Catalysis, Marie Curie 2, 28049 Madrid, Spain; [email protected] * Correspondence: [email protected]; Tel.: +44-1248-388607; Fax: +44-1248-382569 Received: 27 April 2020; Accepted: 15 May 2020; Published: 19 May 2020 Abstract: The archaeon Cuniculiplasma divulgatum is ubiquitous in acidic environments with low-to-moderate temperatures. However, molecular mechanisms underlying its ability to thrive at lower temperatures remain unexplored. Using mass spectrometry (MS)-based proteomics, we analysed the effect of short-term (3 h) exposure to cold. The C. divulgatum genome encodes 2016 protein-coding genes, from which 819 proteins were identified in the cells grown under optimal conditions. In line with the peptidolytic lifestyle of C. divulgatum, its intracellular proteome revealed the abundance of proteases, ABC transporters and cytochrome C oxidase. From 747 quantifiable polypeptides, the levels of 582 proteins showed no change after the cold shock, whereas 104 proteins were upregulated suggesting that they might be contributing to cold adaptation. -

A Korarchaeal Genome Reveals Insights Into the Evolution of the Archaea
A korarchaeal genome reveals insights into the evolution of the Archaea James G. Elkinsa,b, Mircea Podarc, David E. Grahamd, Kira S. Makarovae, Yuri Wolfe, Lennart Randauf, Brian P. Hedlundg, Ce´ line Brochier-Armaneth, Victor Kunini, Iain Andersoni, Alla Lapidusi, Eugene Goltsmani, Kerrie Barryi, Eugene V. Koonine, Phil Hugenholtzi, Nikos Kyrpidesi, Gerhard Wannerj, Paul Richardsoni, Martin Kellerc, and Karl O. Stettera,k,l aLehrstuhl fu¨r Mikrobiologie und Archaeenzentrum, Universita¨t Regensburg, D-93053 Regensburg, Germany; cBiosciences Division, Oak Ridge National Laboratory, Oak Ridge, TN 37831; dDepartment of Chemistry and Biochemistry, University of Texas, Austin, TX 78712; eNational Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894; fDepartment of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520; gSchool of Life Sciences, University of Nevada, Las Vegas, NV 89154; hLaboratoire de Chimie Bacte´rienne, Unite´ Propre de Recherche 9043, Centre National de la Recherche Scientifique, Universite´de Provence Aix-Marseille I, 13331 Marseille Cedex 3, France; iU.S. Department of Energy Joint Genome Institute, Walnut Creek, CA 94598; jInstitute of Botany, Ludwig Maximilians University of Munich, D-80638 Munich, Germany; and kInstitute of Geophysics and Planetary Physics, University of California, Los Angeles, CA 90095 Communicated by Carl R. Woese, University of Illinois at Urbana–Champaign, Urbana, IL, April 2, 2008 (received for review January 7, 2008) The candidate division Korarchaeota comprises a group of uncul- and sediment samples from Obsidian Pool as an inoculum. The tivated microorganisms that, by their small subunit rRNA phylog- cultivation system supported the stable growth of a mixed commu- eny, may have diverged early from the major archaeal phyla nity of hyperthermophilic bacteria and archaea including an or- Crenarchaeota and Euryarchaeota. -

Archaeal Adaptation to Higher Temperatures Revealed by Genomic Sequence of Thermoplasma Volcanium
Archaeal adaptation to higher temperatures revealed by genomic sequence of Thermoplasma volcanium Tsuyoshi Kawashima*†, Naoki Amano*†‡, Hideaki Koike*†, Shin-ichi Makino†, Sadaharu Higuchi†, Yoshie Kawashima-Ohya†, Koji Watanabe§, Masaaki Yamazaki§, Keiichi Kanehori¶, Takeshi Kawamotoʈ, Tatsuo Nunoshiba**, Yoshihiro Yamamoto††, Hironori Aramaki‡‡, Kozo Makino§§, and Masashi Suzuki†¶¶ †National Institute of Bioscience and Human Technology, Core Research for Evolutional Science and Technology Centre of Structural Biology, 1-1 Higashi, Tsukuba 305-0046, Japan; ‡Doctoral Program in Medical Sciences, University of Tsukuba, 1-1-1 Tennohdai, Tsukuba 305-0006, Japan; §Bioscience Research Laboratory, Fujiya, 228 Soya, Hadano 257-0031, Japan; ¶DNA Analysis Department, Techno Research Laboratory, Hitachi Science Systems, 1-280 Higashi-Koigakubo, Kokubunji 185-8601, Japan; ʈDepartment of Biochemistry, Hiroshima University, School of Dentistry, 1-2-3 Kasumi, Minami-ku, Hiroshima 734-8553, Japan; **Department of Molecular and Cellular Biology, Biological Institute, Graduate School of Science, Tohoku University, Sendai 980-8578, Japan; ††Department of Genetics, Hyogo College of Medicine, Nishinomiya 663-8501, Japan; ‡‡Department of Molecular Biology, Daiichi College of Pharmaceutical Science, 22-1 Tamagawa-cho, Minami-ku, Fukuoka 815-8511, Japan; and §§Department of Molecular Microbiology, The Research Institute of Microbial Diseases, Osaka University, 3-1 Yamadaoka, Suita 565-0871, Japan Edited by Aaron Klug, Royal Society of London, London, United Kingdom, and approved October 16, 2000 (received for review August 18, 2000) The complete genomic sequence of the archaeon Thermoplasma contigs. The remaining gaps were bridged by DNA fragments volcanium, possessing optimum growth temperature (OGT) of constructed using the PCR. The average repetition in sequencing 60°C, is reported. By systematically comparing this genomic se- the same base positions was 13-fold. -

The Main (Glyco) Phospholipid (MPL) of Thermoplasma Acidophilum
International Journal of Molecular Sciences Review The Main (Glyco) Phospholipid (MPL) of Thermoplasma acidophilum Hans-Joachim Freisleben 1,2 1 Goethe-Universität, Gustav-Embden-Zentrum, Laboratory of Microbiological Chemistry, Theodor-Stern-Kai 7, D-60590 Frankfurt am Main, Germany; [email protected] 2 Universitas Indonesia, Medical Research Unit, Faculty of Medicine, Jalan Salemba Raya 6, Jakarta 10430, Indonesia Received: 19 September 2019; Accepted: 18 October 2019; Published: 21 October 2019 Abstract: The main phospholipid (MPL) of Thermoplasma acidophilum DSM 1728 was isolated, purified and physico-chemically characterized by differential scanning calorimetry (DSC)/differential thermal analysis (DTA) for its thermotropic behavior, alone and in mixtures with other lipids, cholesterol, hydrophobic peptides and pore-forming ionophores. Model membranes from MPL were investigated; black lipid membrane, Langmuir-Blodgett monolayer, and liposomes. Laboratory results were compared to computer simulation. MPL forms stable and resistant liposomes with highly proton-impermeable membrane and mixes at certain degree with common bilayer-forming lipids. Monomeric bacteriorhodopsin and ATP synthase from Micrococcus luteus were co-reconstituted and light-driven ATP synthesis measured. This review reports about almost four decades of research on Thermoplasma membrane and its MPL as well as transfer of this research to Thermoplasma species recently isolated from Indonesian volcanoes. Keywords: Thermoplasma acidophilum; Thermoplasma volcanium; -

Different Proteins Mediate Step-Wise Chromosome Architectures in 2 Thermoplasma Acidophilum and Pyrobaculum Calidifontis
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Different Proteins Mediate Step-wise Chromosome Architectures in 2 Thermoplasma acidophilum and Pyrobaculum calidifontis 3 4 5 Hugo Maruyama1†*, Eloise I. Prieto2†, Takayuki Nambu1, Chiho Mashimo1, Kosuke 6 Kashiwagi3, Toshinori Okinaga1, Haruyuki Atomi4, Kunio Takeyasu5 7 8 1 Department of Bacteriology, Osaka Dental University, Hirakata, Japan 9 2 National Institute of Molecular Biology and Biotechnology, University of the Philippines 10 Diliman, Quezon City, Philippines 11 3 Department of Fixed Prosthodontics, Osaka Dental University, Hirakata, Japan 12 4 Department of Synthetic Chemistry and Biological Chemistry, Graduate School of Engineering, 13 Kyoto University, Kyoto, Japan 14 5 Graduate School of Biostudies, Kyoto University, Kyoto, Japan 15 † These authors have contributed equally to this work 16 17 * Correspondence: 18 Hugo Maruyama 19 [email protected]; [email protected] 20 21 Keywords: archaea, higher-order chromosome structure, nucleoid, chromatin, HTa, histone, 22 transcriptional regulator, horizontal gene transfer 23 Running Title: Step-wise chromosome architecture in Archaea 24 Manuscript length: 6955 words 25 Number of Figures: 7 26 Number of Tables: 3 bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

(Agus Kurnia)Ok
HAYATI Journal of Biosciences September 2012 Available online at: Vol. 19 No. 3, p 150-154 http://journal.ipb.ac.id/index.php/hayati EISSN: 2086-4094 DOI: 10.4308/hjb.19.3.150 SHORT COMMUNICATION Archaeal Life on Tangkuban Perahu- Sampling and Culture Growth in Indonesian Laboratories SRI HANDAYANI1, IMAN SANTOSO1, HANS-JOACHIM FREISLEBEN2∗, HARALD HUBER3, ANDI1, FERY ARDIANSYAH1, CENMI MULYANTO1, ZESSINDA LUTHFA1, ROSARI SALEH1, SERUNI KUSUMA UDYANINGSIH FREISLEBEN1, SEPTELIA INAWATI WANANDI2, MICHAEL THOMM3 1Faculty of Mathematics and Natural Sciences, Universitas Indonesia, Jakarta-Depok, Jakarta 10430, Indonesia 2Faculty of Medicine, Universitas Indonesia, Jalan Salemba Raya No. 6, Jakarta 10430, Indonesia 3Department of Microbiology, Archaea Centre, University of Regensburg, Germany Received May 9, 2012/Accepted September 21, 2012 The aim of the expedition to Tangkuban Perahu, West Java was to obtain archaeal samples from the solfatara fields located in Domas crater. This was one of the places, where scientists from the University of Regensburg Germany had formerly isolated Indonesian archaea, especially Thermoplasma and Sulfolobus species but not fully characterized. We collected five samples from mud holes with temperatures from 57 to 88 oC and pH of 1.5-2. A portion of each sample was grown at the University of Regensburg in modified Allen’s medium at 80 oC. From four out of five samples enrichment cultures were obtained, autotrophically on elemental sulphur and heterotrophically on sulfur and yeast extract; electron micrographs are presented. In the laboratories of Universitas Indonesia the isolates were cultured at 55-60 oC in order to grow tetraetherlipid synthesizing archaea, both Thermoplasmatales and Sulfolobales. Here, we succeeded to culture the same type of archaeal cells, which had been cultured in Regensburg, probably a Sulfolobus species and in Freundt’s medium, Thermoplasma species. -
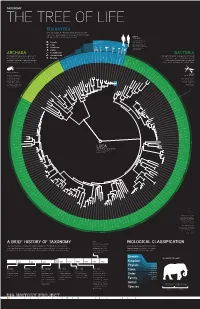
A Brief History of Taxonomy Biological Classification
TAXONOMY THE TREE OF LIFE EUKARYOTA This domain includes all of the plants, animals, and fungi, and some single-celled organisms. Eukaryotes are distinguished by their complex cells, which contain a membrane-enclosed nucleus. Humans Homo sapiens The creatures most familiar to us, Our species, primates in the animals, are members of the Animalia kingdom of the Animalia same kingdom. Eukaryota, is thought to have Fungi Mosquito Red first evolved in Africa about Pufferfish Junglefowl Roundworm Mouse 200,000 years ago. Genetically, Amoebozoa Chimpanzee our closest living relative Plantae is the chimpanzee. Archaeplastida Schizosaccharomyces pombe ARCHAEA Saccharomyces cerevisiae BACTERIA Caenorhabditis briggsae Caenorhabditis elegans Eremothecium gossypii These single-celled prokaryotic organisms often Chromalveolata Dictyostelium discoideum These single-celled prokaryotic organisms were among Cyanidioschyzon merolae live in extreme environmental conditions. Once Excavata Arabidopsis thaliana the first life forms to appear on Earth. Often spherical, Plasmodium falciparum considered to be Bacteria, these microorganisms Cryptosporidium hominis rod-like, or spiral in shape, these microorganisms Thalassiosira pseudonana Oryza sativa Anopheles gambiae Drosophila melanogaster Takifugu rubripes Danio rerio are now recognized as a separate domain of life. Gallus gallus function without a membrane-enclosed cell nucleus. Rattus norvegicus Mus musculus Methanococcus jannaschii Leishmania major Homo sapiens Pan troglodytes Methanococcus maripaludi Thermoanaerobacter -

Biotechnology of Archaea- Costanzo Bertoldo and Garabed Antranikian
BIOTECHNOLOGY– Vol. IX – Biotechnology Of Archaea- Costanzo Bertoldo and Garabed Antranikian BIOTECHNOLOGY OF ARCHAEA Costanzo Bertoldo and Garabed Antranikian Technical University Hamburg-Harburg, Germany Keywords: Archaea, extremophiles, enzymes Contents 1. Introduction 2. Cultivation of Extremophilic Archaea 3. Molecular Basis of Heat Resistance 4. Screening Strategies for the Detection of Novel Enzymes from Archaea 5. Starch Processing Enzymes 6. Cellulose and Hemicellulose Hydrolyzing Enzymes 7. Chitin Degradation 8. Proteolytic Enzymes 9. Alcohol Dehydrogenases and Esterases 10. DNA Processing Enzymes 11. Archaeal Inteins 12. Conclusions Glossary Bibliography Biographical Sketches Summary Archaea are unique microorganisms that are adapted to survive in ecological niches such as high temperatures, extremes of pH, high salt concentrations and high pressure. They produce novel organic compounds and stable biocatalysts that function under extreme conditions comparable to those prevailing in various industrial processes. Some of the enzymes from Archaea have already been purified and their genes successfully cloned in mesophilic hosts. Enzymes such as amylases, pullulanases, cyclodextrin glycosyltransferases, cellulases, xylanases, chitinases, proteases, alcohol dehydrogenase,UNESCO esterases, and DNA-modifying – enzymesEOLSS are of potential use in various biotechnological processes including in the food, chemical and pharmaceutical industries. 1. Introduction SAMPLE CHAPTERS The industrial application of biocatalysts began in 1915 with the introduction of the first detergent enzyme by Dr. Röhm. Since that time enzymes have found wider application in various industrial processes and production (see Enzyme Production). The most important fields of enzyme application are nutrition, pharmaceuticals, diagnostics, detergents, textile and leather industries. There are more than 3000 enzymes known to date that catalyze different biochemical reactions among the estimated total of 7000; only 100 enzymes are being used industrially. -

Temperature and Elemental Sulfur Shape Microbial Communities in Two Extremely Acidic Aquatic Volcanic Environments
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.24.312660; this version posted September 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Temperature and elemental sulfur shape microbial communities in two extremely acidic aquatic volcanic environments Diego Rojas-Gätjens1, Alejandro Arce-Rodríguez2α, Fernando Puente-Sánchez3, Roberto Avendaño1, Eduardo Libby4, Raúl Mora-Amador5,6, Keilor Rojas-Jimenez7, Paola Fuentes-Schweizer4,8, Dietmar H. Pieper2 & Max Chavarría1,4,9* 1Centro Nacional de Innovaciones Biotecnológicas (CENIBiot), CeNAT-CONARE, 1174-1200, San José, Costa Rica 2Microbial Interactions and Processes Research Group, Helmholtz Centre for Infection Research, 38124, Braunschweig, Germany 3Systems Biology Program, Centro Nacional de Biotecnología (CNB-CSIC), C/Darwin 3, 28049 Madrid, Spain 4Escuela de Química, Universidad de Costa Rica, 11501-2060, San José, Costa Rica 5Escuela Centroamericana de Geología, Universidad de Costa Rica, 11501-2060, San José, Costa Rica 6Laboratorio de Ecología Urbana, Universidad Estatal a Distancia, 11501-2060, San José, Costa Rica 7Escuela de Biología, Universidad de Costa Rica, 11501-2060, San José, Costa Rica, 8Centro de Investigación en Electroquímica y Energía Química (CELEQ), Universidad de Costa Rica, 11501-2060, San José, Costa Rica 9Centro de Investigaciones en Productos Naturales -

Perspectives on the Origin of Microfilaments, Microtubules, the Rel- Evant Chaperonin System and Cytoskeletal Motors- a Commenta
Cell Research (2003); 13(4):219-227 http://www.cell-research.com REVIEW Perspectives on the origin of microfilaments, microtubules, the rel- evant chaperonin system and cytoskeletal motors- a commentary on the spiro- chaete origin of flagella JING YAN LI*, CHUAN FEN WU** Laboratory of Cellular and Molecular Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, Yunnan 650223, China ABSTRACT The origin of cytoskeleton and the origin of relevant intracellular transportation system are big problems for understanding the emergence of eukaryotic cells. The present article summarized relevant information of evidences and molecular traces on the origin of actin, tubulin, the chaperonin system for folding them, myosins, kinesins, axonemal dyneins and cytoplasmic dyneins. On this basis the authors proposed a series of works, which should be done in the future, and indicated the ways for reaching the targets. These targets are mainly: 1) the reconstruction of evolutionary path from MreB protein of archaeal ancestor of eukaryotic cells to typical actin; 2) the finding of the MreB or MreB-related proteins in crenarchaea and using them to examine J. A. Lake's hypothesis on the origin of eukaryote from "eocytes" (crenarchaea); 3) the examinations of the existence and distribution of cytoskeleton made of MreB-related protein within coccoid archaea, espe- cially in amoeboid archaeon Thermoplasm acidophilum; 4) using Thermoplasma as a model of archaeal an- cestor of eukaryotic cells; 5) the searching for the homolog of ancestral dynein in present-day living archaea. During the writing of this article, Margulis' famous spirochaete hypothesis on the origin of flagella and cilia was unexpectedly involved and analyzed from aspects of tubulins, dyneins and spirochaetes. -

Rare Horizontal Gene Transfer in a Unique Motility Structure Elie Desmond, Céline Brochier-Armanet, Simonetta Gribaldo
Phylogenomics of the archaeal flagellum: rare horizontal gene transfer in a unique motility structure Elie Desmond, Céline Brochier-Armanet, Simonetta Gribaldo To cite this version: Elie Desmond, Céline Brochier-Armanet, Simonetta Gribaldo. Phylogenomics of the archaeal flagel- lum: rare horizontal gene transfer in a unique motility structure. BMC Evolutionary Biology, BioMed Central, 2007, 7, pp.53-70. 10.1186/1471-2148-7-106. hal-00698404 HAL Id: hal-00698404 https://hal.archives-ouvertes.fr/hal-00698404 Submitted on 7 Apr 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License BMC Evolutionary Biology BioMed Central Research article Open Access Phylogenomics of the archaeal flagellum: rare horizontal gene transfer in a unique motility structure Elie Desmond1, Celine Brochier-Armanet2,3 and Simonetta Gribaldo*1 Address: 1Unite Biologie Moléculaire du Gène chez les Extremophiles, Institut Pasteur, 25 rue du Dr. Roux, 75724 Paris Cedex 15, France, 2Université de Provence Aix-Marseille