Gene Expression-Based Recurrence Prediction of Hepatitis B Virus-Related Human Hepatocellular Carcinoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Effect of Distinct Regulator of G-Protein Signaling 10 Isoforms On
EFFECT OF DISTINCT REGULATOR OF G-PROTEIN SIGNALING 10 ISOFORMS ON CYTOKINE PRODUCTION. by BENJAMIN JACKWOOD (Under the Direction of Shelley Hooks) ABSTRACT G-protein coupled receptors (GPCRs) mediate a wide variety of cellular functions related to cell proliferation and survival. Regulators of G-protein Signaling (RGS) proteins that are important negative regulators of both G-proteins and GPCR products. The focus of this thesis involves two human protein variants of RGS10 and their effects on cytokine levels. RGS proteins are GTPase Accelerating Proteins (GAPs) which can facilitate an increased rate of GTP hydrolysis to drive inactivation of GPCR signaling. Based on their ability to regulate GPCRs, RGS proteins are implicated in multiple disease states including cancer and neuro-inflammation. The aim of this study was to define the similarities or differences among RGS10 protein isoforms, and help understand their non-canonical function. Particularly, differences in primary sequence of RGS10 protein variants and their ability to mediate inflammatory cytokines in human embryonic kidney (HEK) cells was investigated. INDEX WORDS: GPCR, RGS10, ISOFORM, TNF-, INFLAMMATION, VARIANTS EFFECT OF DISTINCT REGULATOR OF G-PROTEIN SIGNALING 10 ISOFORMS ON CYTOKINE PRODUCTION. by BENJAMIN JACKWOOD BS, University of North Georgia, 2015 A Thesis Submitted to the Pharmaceutical and Biomedical Sciences department of The University of Georgia in Partial Fulfillment of the Requirements for the Degree. MASTER OF SCIENCE ATHENS, GEORGIA 2017 © 2017 Benjamin Jackwood All Rights Reserved EFFECT OF DISTINCT REGULATOR OF G-PROTEIN RGS10 ISOFORMS ON CYTOKINE PRODUCTION. by BENJAMIN JACKWOOD Major Professor: Shelley B. Hooks Committee: Phillip Greenspan Jason Zastre Electronic Version Approved: Suzanne Barbour Dean of the Graduate School The University of Georgia May 2017 ACKNOWLEDGEMENTS Thanks to my family, friends, and helpful lab mates for all of their support during my time working and studying at the University of Georgia. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

GENE TRANSCRIPTION PROFILE of the DETACHED RETINA (AN AOS THESIS) by David N
GENE TRANSCRIPTION PROFILE OF THE DETACHED RETINA (AN AOS THESIS) BY David N. Zacks MD PhD ABSTRACT Purpose: Separation of the neurosensory retina from the retinal pigment epithelium (RPE) yields many morphologic and functional consequences, including death of the photoreceptor cells, Müller cell hypertrophy, and inner retinal rewiring. Many of these changes are due to the separation-induced activation of specific genes. In this work, we define the gene transcription profile within the retina as a function of time after detachment. We also define the early activation of kinases that might be responsible for the detachment- induced changes in gene transcription. Methods: Separation of the retina from the RPE was induced in Brown-Norway rats by the injection of 1% hyaluronic acid into the subretinal space. Retinas were harvested at 1, 7, and 28 days after separation. Gene transcription profiles for each time point were determined using the Affymetrix Rat 230A gene microarray chip. Transcription levels in detached retinas were compared to those of nondetached retinas with the BRB-ArrayTools Version 3.6.0 using a random variance analysis of variance (ANOVA) model. Confirmation of the significant transcriptional changes for a subset of the genes was performed using microfluidic quantitative real- time polymerase chain reaction (qRT-PCR) assays. Kinase activation was explored using Western blot analysis to look for early phosphorylation of any of the 3 main families of mitogen-activated protein kinases (MAPK): the p38 family, the Janus kinase family, and the p42/p44 family. Results: Retinas separated from the RPE showed extensive alterations in their gene transcription profile. -

Suppression of the Gtpase-Activating Protein RGS10 Increases Rheb-GTP and Mtor Signaling in Ovarian Cancer Cells Molly K
Cancer Letters 369 (2015) 175–183 Contents lists available at ScienceDirect Cancer Letters journal homepage: www.elsevier.com/locate/canlet Original Articles Suppression of the GTPase-activating protein RGS10 increases Rheb-GTP and mTOR signaling in ovarian cancer cells Molly K. Altman, Ali A. Alshamrani, Wei Jia, Ha T. Nguyen, Jada M. Fambrough, Sterling K. Tran, Mihir B. Patel, Pooya Hoseinzadeh, Aaron M. Beedle, Mandi M. Murph * Department of Pharmaceutical and Biomedical Sciences, College of Pharmacy, The University of Georgia, 240 W. Green Street, Athens, GA 30602, USA ARTICLE INFO ABSTRACT Article history: The regulator of G protein signaling 10 (RGS10) protein is a GTPase activating protein that accelerates Received 26 June 2015 the hydrolysis of GTP and therefore canonically inactivates G proteins, ultimately terminating signaling. Received in revised form 14 August 2015 Rheb is a small GTPase protein that shuttles between its GDP- and GTP-bound forms to activate mTOR. Accepted 17 August 2015 Since RGS10 suppression augments ovarian cancer cell viability, we sought to elucidate the molecular mechanism. Following RGS10 suppression in serum-free conditions, phosphorylation of mTOR, the eu- Keywords: karyotic translation initiation factor 4E binding protein 1 (4E-BP1), p70S6K and S6 Ribosomal Protein Regulator of G protein Signaling 10 protein appear. Furthermore, suppressing RGS10 increases activated Rheb, suggesting RGS10 antagonizes mTOR (RGS10) Lysophosphatidic acid signaling via the small G-protein. The effects of RGS10 suppression are enhanced after stimulating cells mTOR with the growth factor, lysophosphatidic acid, and reduced with mTOR inhibitors, temsirolimus and INK- 4E-BP1 128. Suppression of RGS10 leads to an increase in cell proliferation, even in the presence of etoposide. -

Expression and Function of Regulator of G-Protein Signaling 10
EXPRESSION AND FUNCTION OF REGULATOR OF G-PROTEIN SIGNALING 10 (RGS10) IN OVARIAN CANCER AND MICROGLIA by MOURAD WAGDY AHMED ALI (Under the Direction of Shelley B. Hooks) ABSTRACT G-Protein coupled receptors (GPCRs) mediate a wide array of cellular functions, such as cell proliferation, migration, and survival. Regulators of G-protein signaling (RGS) proteins are a diverse family of proteins that regulate signaling pathways downstream of GPCRs by acting on G-proteins. The focus of this dissertation is on the regulation of G-protein pathways in cancer and inflammation by RGS proteins, particularly by RGS10. We focused on signaling initiated by two related receptor families, lysophosphatidic acid (LPA) and sphingosine-1-phosphate (S1P) receptors, which are implicated in ovarian cancer and neuroinflammation, respectively. Aberrant expression and mutations in RGS proteins have been implicated in diseases such as cancer and autoimmune disorders. The aim of this study was to define the function and the expression of RGS proteins, particularly RGS10, in ovarian cancer and microglia. LPA is the predominant growth factor in ovarian cancer, promoting proliferation, migration, and survival. RGS proteins negatively regulate LPA-mediated effects in ovarian cancer. We determined that RGS proteins, RGS10 and RGS17 regulate LPA- mediated survival in ovarian cancer cells. Specifically, our data demonstrate that RGS10 and RGS17 negatively regulate LPA-mediated AKT survival pathway in ovarian cancer cells. Further, we show that RGS10 and RGS17 are down-regulated in chemoresistant ovarian cancer cells, and our results show that RGS10 is epigenetically silenced in chemoresistant ovarian cancer cells via increased DNA methylation and decreased histone acetylation of the RGS10 promoter by DNA methyltransferase 1 (DNMT1), and histone deacetylase 1 (HDAC1), respectively. -

Susceptibility Genes for Age-Related Maculopathy on Chromosome 10Q26 Johanna Jakobsdottir,1 Yvette P
Am. J. Hum. Genet. 77:389–407, 2005 Susceptibility Genes for Age-Related Maculopathy on Chromosome 10q26 Johanna Jakobsdottir,1 Yvette P. Conley,2,3 Daniel E. Weeks,1,2 Tammy S. Mah,4 Robert E. Ferrell,2 and Michael B. Gorin2,4 Departments of 1Biostatistics and 2Human Genetics, Graduate School of Public Health, 3Department of Health Promotion and Development, School of Nursing, and 4UPMC Eye Center, Department of Ophthalmology, School of Medicine, University of Pittsburgh, Pittsburgh On the basis of genomewide linkage studies of families affected with age-related maculopathy (ARM), we previously identified a significant linkage peak on 10q26, which has been independently replicated by several groups. We performed a focused SNP genotyping study of our families and an additional control cohort. We identified a strong association signal overlying three genes, PLEKHA1, LOC387715, and PRSS11. All nonsynonymous SNPs in this critical region were genotyped, yielding a highly significant association (P ! .00001) between PLEKHA1/LOC387715 and ARM. Although it is difficult to determine statistically which of these two genes is most important, SNPs in PLEKHA1 are more likely to account for the linkage signal in this region than are SNPs in LOC387715; thus, this gene and its alleles are implicated as an important risk factor for ARM. We also found weaker evidence supporting the possible involvement of the GRK5/RGS10 locus in ARM. These associations appear to be independent of the association of ARM with the Y402H allele of complement factor H, which has previously been reported as a major susceptibility factor for ARM. The combination of our analyses strongly implicates PLEKHA1/LOC387715 as primarily responsible for the evidence of linkage of ARM to the 10q26 locus and as a major contributor to ARM susceptibility. -

Integrated Network Analysis Identifying Potential Novel Drug Candidates
www.nature.com/scientificreports OPEN Integrated network analysis identifying potential novel drug candidates and targets for Parkinson’s disease Pusheng Quan1, Kai Wang2, Shi Yan1, Shirong Wen1, Chengqun Wei3, Xinyu Zhang1, Jingwei Cao1 & Lifen Yao1* This study aimed to identify potential novel drug candidates and targets for Parkinson’s disease. First, 970 genes that have been reported to be related to PD were collected from fve databases, and functional enrichment analysis of these genes was conducted to investigate their potential mechanisms. Then, we collected drugs and related targets from DrugBank, narrowed the list by proximity scores and Inverted Gene Set Enrichment analysis of drug targets, and identifed potential drug candidates for PD treatment. Finally, we compared the expression distribution of the candidate drug-target genes between the PD group and the control group in the public dataset with the largest sample size (GSE99039) in Gene Expression Omnibus. Ten drugs with an FDR < 0.1 and their corresponding targets were identifed. Some target genes of the ten drugs signifcantly overlapped with PD-related genes or already known therapeutic targets for PD. Nine diferentially expressed drug-target genes with p < 0.05 were screened. This work will facilitate further research into the possible efcacy of new drugs for PD and will provide valuable clues for drug design. Parkinson’s disease (PD) is a pervasive, progressive, disabling neurodegenerative disorder with motor and non- motor features1. PD places a signifcant burden on society and the afected individuals, and approximately 6.1 million people worldwide had been diagnosed with PD in 20162. Dopamine depletion leading to hyperactivity of the corticostriatal glutamatergic pathway is thought to be primarily responsible for parkinsonian symptoms such as resting tremors, rigidity, dyskinesia and postural instability 3,4. -

Gαi2 Signaling Regulates Inflammasome Priming and Cytokine Production by Biasing Macrophage Phenotype Determination
Gαi2 Signaling Regulates Inflammasome Priming and Cytokine Production by Biasing Macrophage Phenotype Determination This information is current as Ali Vural, Neel R. Nabar, Il-Young Hwang, Silke Sohn, of September 26, 2021. Chung Park, Mikael C. I. Karlsson, Joe B. Blumer and John H. Kehrl J Immunol published online 25 January 2019 http://www.jimmunol.org/content/early/2019/01/24/jimmun ol.1801145 Downloaded from Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on September 26, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2019 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Published January 25, 2019, doi:10.4049/jimmunol.1801145 The Journal of Immunology Gai2 Signaling Regulates Inflammasome Priming and Cytokine Production by Biasing Macrophage Phenotype Determination Ali Vural,*,1,2 Neel R. Nabar,*,†,1 Il-Young Hwang,* Silke Sohn,† Chung Park,* Mikael C. I. Karlsson,† Joe B. Blumer,‡ and John H. Kehrl* Macrophages exist as innate immune subsets that exhibit phenotypic heterogeneity and functional plasticity. -
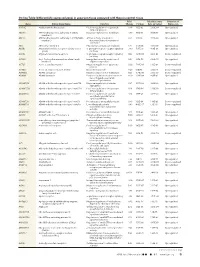
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

The High NRF2 Expression Confers Chemotherapy Resistance Partly Through Up-Regulated DUSP1 in Myelodysplastic Syndromes
Myelodysplastic Syndromes SUPPLEMENTARY APPENDIX The high NRF2 expression confers chemotherapy resistance partly through up-regulated DUSP1 in myelodysplastic syndromes Peipei Lin, 1,2,3,4 * Yanling Ren, 1,2,3 * Xiaomei Yan, 4 Yingwan Luo, 1,2,3 Hua Zhang, 1,2,3 Meenu Kesarwani, 4 Jiachen Bu, 4 Di Zhan, 4 Yile Zhou, 1,2,4 Yuting Tang, 4 Shuanghong Zhu, 1,2,3 Weilai Xu, 1,2,3 Xinping Zhou, 1,2,3 Chen Mei, 1,2,3 Liya Ma, 1,2,3 Li Ye, 1,2,3 Chao Hu, 1,2 Mohammad Azam, 4 Wei Ding, 5 Jie Jin, 1,2 Gang Huang 4# and Hongyan Tong 1,2,3# 1Department of Hematology, the First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou, China; 2Institute of Hema - tology, Zhejiang University, Hangzhou, China; 3Myelodysplastic Syndromes Diagnosis and Therapy Center, the First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou, China; 4Divisions of Pathology and Experimental Hematology and Cancer Biology, Cincinnati Children's Hospital Medical Center, OH, USA and 5Department of Pathology, the First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou, China *PL and YR contributed equally to this work. #HYT and GH contributed equally to this study as joint senior authors. ©2019 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2018.197749 Received: May 15, 2018. Accepted: September 26, 2018. Pre-published: September 27, 2018. Correspondence: HONGYAN TONG - [email protected] GANG HUANG - [email protected] Title: The high NRF2 expression confers chemotherapy resistance -

Novel and Highly Recurrent Chromosomal Alterations in Se´Zary Syndrome
Research Article Novel and Highly Recurrent Chromosomal Alterations in Se´zary Syndrome Maarten H. Vermeer,1 Remco van Doorn,1 Remco Dijkman,1 Xin Mao,3 Sean Whittaker,3 Pieter C. van Voorst Vader,4 Marie-Jeanne P. Gerritsen,5 Marie-Louise Geerts,6 Sylke Gellrich,7 Ola So¨derberg,8 Karl-Johan Leuchowius,8 Ulf Landegren,8 Jacoba J. Out-Luiting,1 Jeroen Knijnenburg,2 Marije IJszenga,2 Karoly Szuhai,2 Rein Willemze,1 and Cornelis P. Tensen1 Departments of 1Dermatology and 2Molecular Cell Biology, Leiden University Medical Center, Leiden, the Netherlands; 3Department of Dermatology, St Thomas’ Hospital, King’s College, London, United Kingdom; 4Department of Dermatology, University Medical Center Groningen, Groningen, the Netherlands; 5Department of Dermatology, Radboud University Nijmegen Medical Center, Nijmegen, the Netherlands; 6Department of Dermatology, Gent University Hospital, Gent, Belgium; 7Department of Dermatology, Charite, Berlin, Germany; and 8Department of Genetics and Pathology, Rudbeck Laboratory, University of Uppsala, Uppsala, Sweden Abstract Introduction This study was designed to identify highly recurrent genetic Se´zary syndrome (Sz) is an aggressive type of cutaneous T-cell alterations typical of Se´zary syndrome (Sz), an aggressive lymphoma/leukemia of skin-homing, CD4+ memory T cells and is cutaneous T-cell lymphoma/leukemia, possibly revealing characterized by erythroderma, generalized lymphadenopathy, and pathogenetic mechanisms and novel therapeutic targets. the presence of neoplastic T cells (Se´zary cells) in the skin, lymph High-resolution array-based comparative genomic hybridiza- nodes, and peripheral blood (1). Sz has a poor prognosis, with a tion was done on malignant T cells from 20 patients. disease-specific 5-year survival of f24% (1).