Impact of IQ on the Diagnostic Yield of Chromosomal Microarray in A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
Investigation of Modifier Genes Within Copy Number Variations in Rett Syndrome
See discussions, stats, and author profiles for this publication at: http://www.researchgate.net/publication/51147767 Investigation of modifier genes within copy number variations in Rett syndrome ARTICLE in JOURNAL OF HUMAN GENETICS · MAY 2011 Impact Factor: 2.53 · DOI: 10.1038/jhg.2011.50 · Source: PubMed CITATIONS DOWNLOADS VIEWS 6 89 134 15 AUTHORS, INCLUDING: Dag H Yasui Maria Antonietta Mencarelli University of California, Davis Azienda Ospedaliera Universitaria Senese 30 PUBLICATIONS 1,674 CITATIONS 58 PUBLICATIONS 962 CITATIONS SEE PROFILE SEE PROFILE Francesca Mari Janine M Lasalle Università degli Studi di Siena University of California, Davis 81 PUBLICATIONS 1,658 CITATIONS 98 PUBLICATIONS 3,525 CITATIONS SEE PROFILE SEE PROFILE Available from: Janine M Lasalle Retrieved on: 22 July 2015 Europe PMC Funders Group Author Manuscript J Hum Genet. Author manuscript; available in PMC 2012 January 01. Published in final edited form as: J Hum Genet. 2011 July ; 56(7): 508–515. doi:10.1038/jhg.2011.50. Europe PMC Funders Author Manuscripts Investigation of modifier genes within copy number variations in Rett syndrome Rosangela Artuso1,*, Filomena Tiziana Papa1,*, Elisa Grillo1, Mafalda Mucciolo1, Dag H. Yasui2, Keith W. Dunaway2, Vittoria Disciglio1, Maria Antonietta Mencarelli1, Marzia Pollazzon1, Michele Zappella3, Giuseppe Hayek4, Francesca Mari1, Alessandra Renieri1, Janine M. LaSalle2, and Francesca Ariani1 1 Medical Genetics Section, Biotechnology Department, University of Siena, Italy 2 Medical Microbiology and Immunology, Genome Center, School of Medicine, University of California, Davis, CA, USA 3 Child Neuropsychiatry, Versilia Hospital, Viareggio, Italy 4 Infantile Neuropsychiatry, Siena General Hospital, Italy Abstract MECP2 mutations are responsible for two different phenotypes in females, classical Rett syndrome and the milder Zappella variant (Z-RTT). -

Recurrent Inversion Toggling and Great Ape Genome Evolution
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Nat Genet Manuscript Author . Author manuscript; Manuscript Author available in PMC 2020 December 15. Published in final edited form as: Nat Genet. 2020 August ; 52(8): 849–858. doi:10.1038/s41588-020-0646-x. Recurrent inversion toggling and great ape genome evolution David Porubsky1,2,9, Ashley D. Sanders3,9, Wolfram Höps3, PingHsun Hsieh1, Arvis Sulovari1, Ruiyang Li1, Ludovica Mercuri4, Melanie Sorensen1, Shwetha C. Murali1,5, David Gordon1,5, Stuart Cantsilieris1,6, Alex A. Pollen7, Mario Ventura4, Francesca Antonacci4, Tobias Marschall8, Jan O. Korbel3, Evan E. Eichler1,5,* 1Department of Genome Sciences, University of Washington School of Medicine, Seattle, Washington, USA. 2Max Planck Institute for Informatics, Saarland Informatics Campus E1.4, Saarbrücken, Germany. 3European Molecular Biology Laboratory (EMBL), Genome Biology Unit, Heidelberg, Germany. 4Dipartimento di Biologia, Università degli Studi di Bari “Aldo Moro”, Bari, Italy. 5Howard Hughes Medical Institute, University of Washington, Seattle, Washington, USA. 6Centre for Eye Research Australia, Department of Surgery (Ophthalmology), University of Melbourne, Royal Victorian Eye and Ear Hospital, East Melbourne, Victoria, Australia. 7Department of Neurology, University of California, San Francisco (UCSF), San Francisco, California, USA. 8Institute for Medical Biometry and Bioinformatics, Medical Faculty, Heinrich Heine University Düsseldorf, Germany. 9These authors contributed equally to this work. Abstract Inversions play an important role in disease and evolution but are difficult to characterize because their breakpoints map to large repeats. We increased by six-fold the number (n = 1,069) of previously reported great ape inversions using Strand-seq and long-read sequencing. We find that the X chromosome is most enriched (2.5-fold) for inversions based on its size and duplication content. -

10Q11.22Q11.23 Deletions and Microdeletions
Inform Network Support Rare Chromosome Disorder Support Group The Stables, Station Road West, Oxted, Surrey RH8 9EE, United Kingdom Tel: +44(0)1883 723356 [email protected] I www.rarechromo.org Join Unique for family links, information and support. Unique is a charity without government funding, existing entirely on donations and grants. If you can, please make a donation via our website at http://www.rarechromo.org/donate Please help us to help you! 10q11.22q11.23 Facebook groups www.facebook.com/groups/chromosome10disorder/ www.facebook.com/groups/152331614838414/ Deletions and Unique mentions other organisations’ message boards and websites to help families looking for information. This does not imply that we endorse their content or have any responsibility for it. Microdeletions This information guide is not a substitute for personal medical advice. Families should consult a medically qualified clinician in all matters relating to genetic diagnosis, management and health. Information on genetic changes is a very fast-moving field and while the information in this guide is believed to be the best available at the time of publication, some facts may later change. Unique does its best to keep abreast of changing information and to review its published guides as needed. This booklet was compiled by Unique (AP) and reviewed by Dr Corrina Powell, Specialist Registrar in Clinical Genetics, and Rosa Spencer- Tansley, Trainee Genetic Counsellor, Leicester Royal Infirmary Clinical Genetics Department ,UK. Version 1 2019 (AP) Copyright © Unique 2020 Rare Chromosome Disorder Support Group Charity Number 1110661 Registered in England and Wales Company Number 5460413 rarechromo.org 20 Genetic information Genetic information genetic Detailed 10q11.22q11.23 microdeletions Chromosome Position Important Microdeletion A 10q11.2q11.23 microdeletion is a rare genetic condition caused by the loss of a 10 Mb genes examples small piece of genetic material from one of the body’s 46 chromosomes – chromosome 10. -

Phenotype Informatics
Freie Universit¨atBerlin Department of Mathematics and Computer Science Phenotype informatics: Network approaches towards understanding the diseasome Sebastian Kohler¨ Submitted on: 12th September 2012 Dissertation zur Erlangung des Grades eines Doktors der Naturwissenschaften (Dr. rer. nat.) am Fachbereich Mathematik und Informatik der Freien Universitat¨ Berlin ii 1. Gutachter Prof. Dr. Martin Vingron 2. Gutachter: Prof. Dr. Peter N. Robinson 3. Gutachter: Christopher J. Mungall, Ph.D. Tag der Disputation: 16.05.2013 Preface This thesis presents research work on novel computational approaches to investigate and characterise the association between genes and pheno- typic abnormalities. It demonstrates methods for organisation, integra- tion, and mining of phenotype data in the field of genetics, with special application to human genetics. Here I will describe the parts of this the- sis that have been published in peer-reviewed journals. Often in modern science different people from different institutions contribute to research projects. The same is true for this thesis, and thus I will itemise who was responsible for specific sub-projects. In chapter 2, a new method for associating genes to phenotypes by means of protein-protein-interaction networks is described. I present a strategy to organise disease data and show how this can be used to link diseases to the corresponding genes. I show that global network distance measure in interaction networks of proteins is well suited for investigat- ing genotype-phenotype associations. This work has been published in 2008 in the American Journal of Human Genetics. My contribution here was to plan the project, implement the software, and finally test and evaluate the method on human genetics data; the implementation part was done in close collaboration with Sebastian Bauer. -
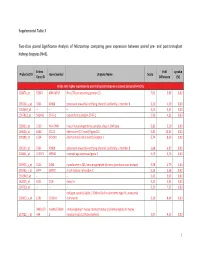
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Determining the Causes of Recessive Retinal Dystrophy
Determining the causes of recessive retinal dystrophy By Mohammed El-Sayed Mohammed El-Asrag BSc (hons), MSc (hons) Genetics and Molecular Biology Submitted in accordance with the requirements for the degree of Doctor of Philosophy The University of Leeds School of Medicine September 2016 The candidate confirms that the work submitted is his own, except where work which has formed part of jointly- authored publications has been included. The contribution of the candidate and the other authors to this work has been explicitly indicated overleaf. The candidate confirms that appropriate credit has been given within the thesis where reference has been made to the work of others. This copy has been supplied on the understanding that it is copyright material and that no quotation from the thesis may be published without proper acknowledgement. The right of Mohammed El-Sayed Mohammed El-Asrag to be identified as author of this work has been asserted by his in accordance with the Copyright, Designs and Patents Act 1988. © 2016 The University of Leeds and Mohammed El-Sayed Mohammed El-Asrag Jointly authored publications statement Chapter 3 (first results chapter) of this thesis is entirely the work of the author and appears in: Watson CM*, El-Asrag ME*, Parry DA, Morgan JE, Logan CV, Carr IM, Sheridan E, Charlton R, Johnson CA, Taylor G, Toomes C, McKibbin M, Inglehearn CF and Ali M (2014). Mutation screening of retinal dystrophy patients by targeted capture from tagged pooled DNAs and next generation sequencing. PLoS One 9(8): e104281. *Equal first- authors. Shevach E, Ali M, Mizrahi-Meissonnier L, McKibbin M, El-Asrag ME, Watson CM, Inglehearn CF, Ben-Yosef T, Blumenfeld A, Jalas C, Banin E and Sharon D (2015). -

10Q11.22Q11.23 Deletions and Microdeletions
10q11.22q11.23 Deletions and Microdeletions rarechromo.org Genetic information Genetic 10q11.22q11.23 microdeletions A 10q11.2q11.23 microdeletion is a rare genetic condition caused by the loss of a small piece of genetic material from one of the body’s 46 chromosomes – chromosome 10. For typical development, chromosomes should contain the expected amount of genetic material. Like many other chromosome disorders, having a missing piece of chromosome 10 may affect a child’s health, development and intellectual abilities. The symptoms observed in people with a 10q11.22q10.23 microdeletion are variable and depend on a number of factors including what and how much genetic material is missing. Background on chromosomes Our bodies are made up of many different types of cells, almost all of which contain the same chromosomes. Each chromosome consists of DNA that codes for our genes. Genes can be thought of as individual instructions that tell the body how to develop, grow and function. Chromosomes come in pairs with one member of each chromosome pair being inherited from each parent. Most of our cells have 23 pairs of chromosomes (a total of 46) as shown in this image. Eggs and sperm, however, have 23 unpaired chromosomes. When an egg and sperm join together at conception, the chromosomes pair up to make a total of 46. Chromosome pairs are numbered 1 to 22 Chromosomes pairs 1-22, X and Y (male) and the 23rd pair comprises the sex Chromosome pair 10 is circled in red chromosomes that determine biological sex. They can be viewed under a microscope and arranged roughly according to size as the image shows. -

Estudo Genético Da Síndrome De Silver-Russell
ADRIANO BONALDI Estudo genético da síndrome de Silver-Russell Dissertação apresentada ao Departamento de Genética e Biologia Evolutiva do Instituto de Biociências da Universidade de São Paulo, para a obtenção de Título de Mestre em Ciências, na Área de Biologia/Genética. São Paulo 2011 ADRIANO BONALDI Estudo genético da síndrome de Silver-Russell Dissertação apresentada ao Departamento de Genética e Biologia Evolutiva do Instituto de Biociências da Universidade de São Paulo, para a obtenção de Título de Mestre em Ciências, na Área de Biologia/Genética. São Paulo 2011 Orientadora: Dra. Angela M. Vianna Morgante BONALDI, ADRIANO Estudo genético da síndrome de Silver-Russell Dissertação (Mestrado) - Instituto de Biociências da Universidade de São Paulo, Departamento de Genética e Biologia Evolutiva 1. Síndrome de Silver-Russell 2. Imprinting genômico 3. Microrrearranjos Cromossômicos Universidade de São Paulo. Instituto de Biociências. Departamento de Genética de Biologia Evolutiva Comissão Julgadora _________________________ _________________________ _________________________ Orientadora Este trabalho foi realizado com os auxílios financeiros da CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) e da FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo) concedidos à orientadora (FAPESP-CEPID 98/14254-2) e ao aluno (CAPES-DS e FAPESP 2009/03341-8). Aos meus pais, Pedro e Lourdes À minha irmã, Fernanda Aos meus amigos e colegas Agradecimentos Agradeço: Ao Departamento de Genética e Biologia Evolutiva do Instituto de Biociências da Universidade de São Paulo, pela possibilidade de realização deste trabalho. À Dra. Angela M. Vianna Morgante, pela orientação neste projeto, por todos os ensinamentos, pela amizade e confiança depositada em mim. À Dra. -

Copy Number Determination of the Gene for the Human Pancreatic
Shebanits et al. BMC Biotechnology (2019) 19:31 https://doi.org/10.1186/s12896-019-0523-9 RESEARCH ARTICLE Open Access Copy number determination of the gene for the human pancreatic polypeptide receptor NPY4R using read depth analysis and droplet digital PCR Kateryna Shebanits1 , Torsten Günther2, Anna C. V. Johansson3, Khurram Maqbool4, Lars Feuk4, Mattias Jakobsson2,5 and Dan Larhammar1* Abstract Background: Copy number variation (CNV) plays an important role in human genetic diversity and has been associated with multiple complex disorders. Here we investigate a CNV on chromosome 10q11.22 that spans NPY4R, the gene for the appetite-regulating pancreatic polypeptide receptor Y4. This genomic region has been challenging to map due to multiple repeated elements and its precise organization has not yet been resolved. Previous studies using microarrays were interpreted to show that the most common copy number was 2 per genome. Results: We have investigated 18 individuals from the 1000 Genomes project using the well-established method of read depth analysis and the new droplet digital PCR (ddPCR) method. We find that the most common copy number for NPY4R is 4. The estimated number of copies ranged from three to seven based on read depth analyses with Control- FREEC and CNVnator, and from four to seven based on ddPCR. We suggest that the difference between our results and those published previously can be explained by methodological differences such as reference gene choice, data normalization and method reliability. Three high-quality archaic human genomes (two Neanderthal and one Denisova) display four copies of the NPY4R gene indicating that a duplication occurred prior to the human-Neanderthal/Denisova split. -

The Evolution and Population Diversity of Human-Specific Segmental Duplications
The evolution and population diversity of human-specific segmental duplications Megan Y. Dennis1,2, Lana Harshman2, Bradley J. Nelson2, Osnat Penn2, Stuart Cantsilieris2, John Huddleston2,3, Francesca Antonacci4, Kelsi Penewit2, Laura Denman2, Archana Raja2,3, Carl Baker2, Kenneth Mark2, Maika Malig2, Nicolette Janke2, Claudia Espinoza2, Holly A. Stessman2, Xander Nuttle2, Kendra Hoekzema2, Tina A. Graves5, Richard K. Wilson5, Evan E. Eichler2,3† 1Genome Center, MIND Institute, and Department of Biochemistry & Molecular Medicine, University of California, Davis, CA 95616, USA 2Department of Genome Sciences, University of Washington School of Medicine, Seattle, WA 98195, USA 3Howard Hughes Medical Institute, University of Washington, Seattle, WA 98195, USA 4Dipartimento di Biologia, Università degli Studi di Bari “Aldo Moro”, Bari 70125, Italy 5McDonnell Genome Institute at Washington University, Washington University School of Medicine, St. Louis, MO 63108, USA †Corresponding author: Evan E. Eichler, Ph.D. University of Washington School of Medicine Howard Hughes Medical Institute Box 355065 Foege S413C, 3720 15th Ave NE Seattle, WA 98195 E-mail: [email protected] 1 ABSTRACT Segmental duplications contribute significantly to the evolution, adaptation and disease- associated instability of the human genome. The largest and most identical duplications suffer from the poorest characterization, often corresponding to genome gaps and misassembly. Here we focus on creating a framework to understand the evolution, copy number variation and -
Clinical, Immunological and Genetic Characteristic of Patients With
ORIGINAL ARTICLE Clinical, immunological and genetic characteristic of patients with clinical phenotype associated to LRBA-deficiency in Colombia Características clínicas, inmunológicas y genéticas de pacientes con fenotipo clínico asociado a la deficiencia de LRBA en Colombia Catalina Martínez-Jaramillo1 , Sebastian Gutierrez-Hincapie1 , Julio César Orrego Arango2 , Gloria María Vásquez-Duque3 , Ruth María Erazo-Garnica3 , Jose Luis Franco1 and Claudia Milena Trujillo-Vargas1 1 Universidad de Antioquia, Facultad de Medicina, Grupo de Inmunodeficiencias Primarias, Medellin, Colombia, 2 Universidad de Antioquia, Grupo de Inmunología Celular e Inmunogenética, Medellin, Colombia, 3 Universidad de Antioquia, Programa de Posgrado, Reumatología Pediátrica, Medellin, Colombia *[email protected] Abstract Background: LPS-responsive beige -like anchor protein (LRBA) deficiency is a primary immunodeficiency OPEN ACCESS disease caused by loss of LRBA protein expression, due to biallelic mutations in LRBA Citation: Martínez-Jaramillo C, gene. LRBA deficiency patients exhibit a clinically heterogeneous syndrome. The Gutierrez-Hincapie S, Orrego main clinical complication of LRBA deficiency is immune dysregulation. Furthermore, AJC, Vásquez-Duque GM, Erazo- hypogammaglobulinemia is found in more than half of patients with LRBA-deficiency. To Garnica RM, Franco JL et al. date, no patients with this condition have been reported in Colombia Clinical, immunological and genetic characteristic of patients with clinical Objective: phenotype associated to LRBA- deficiency in Colombia.Colomb Med To evaluate the expression of the LRBA protein in patients from Colombia with clinical (Cali). 2019; 50(3): 176-91 http:// dx.org/1025100/cm.v50i3.3969 phenotype associated to LRBA-deficiency. Received: 25 Feb 2019 Methods: Revised: 28 Aug 2019 In the present study the LRBA-expression in patients from Colombia with clinical Accepted: 16 Sep 2019 phenotype associated to LRBA-deficiency was evaluated. -

Abstracts from the 53Rd European Society of Human Genetics (ESHG) Conference: E-Posters
European Journal of Human Genetics (2020) 28:798–1016 https://doi.org/10.1038/s41431-020-00741-5 ABSTRACTS COLLECTION Abstracts from the 53rd European Society of Human Genetics (ESHG) Conference: e-Posters © European Society of Human Genetics 2020. Modified from the conference website and published with permission 2020 Volume 28 | Supplement 1 Virtual Conference June 6-9, 2020 Sponsorship: Publication of this supplement was sponsored by the European Society of Human Genetics. All content was reviewed and approved by the ESHG Scientific Programme Committee, which held full responsibility for the abstract selections. 1234567890();,: 1234567890();,: Disclosure Information: In order to help readers form their own judgments of potential bias in published abstracts, authors are asked to declare any competing financial interests. Contributions of up to EUR 10 000.- (Ten thousand Euros, or equivalent value in kind) per year per company are considered “Modest”. Contributions above EUR 10 000.- per year are considered “Significant”. Presenting authors are indicated with bold typeface in the contributor lists. e-Posters improve the prevention and treatment of congenital malformations. E-P01 Reproductive Genetics/Prenatal Genetics Material and methods: The data from Medical Genetic Center Register was analyzing by the MedCalc 15.8 E-P01.07 program. Epidemiological monitoring of congenital malforma- Results: The incidence rate of congenital malformations tions in Yakutia from 2007 to 2018 over a 12-year period averaged 29.4 cases per 1000 newborns with a standard deviation of 2.9. It’s high A. I. Fedorov1, A. L. Sukhomyasova1,2, A. N. Sleptsov2,1,N. indicator In Russia (>20 ‰). At this period, the average R.