Investigation of Modifier Genes Within Copy Number Variations in Rett Syndrome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Complement Factor H Deficiency and Endocapillary Glomerulonephritis Due to Paternal Isodisomy and a Novel Factor H Mutation
Genes and Immunity (2011) 12, 90–99 & 2011 Macmillan Publishers Limited All rights reserved 1466-4879/11 www.nature.com/gene ORIGINAL ARTICLE Complement factor H deficiency and endocapillary glomerulonephritis due to paternal isodisomy and a novel factor H mutation L Schejbel1, IM Schmidt2, M Kirchhoff3, CB Andersen4, HV Marquart1, P Zipfel5 and P Garred1 1Department of Clinical Immunology, Laboratory of Molecular Medicine, Rigshospitalet, Copenhagen, Denmark; 2Department of Pediatrics, Rigshospitalet, Copenhagen, Denmark; 3Department of Clinical Genetics, Rigshospitalet, Copenhagen, Denmark; 4Department of Pathology, Rigshospitalet, Copenhagen, Denmark and 5Department of Infection Biology, Leibniz Institute for Natural Product Research and Infection Biology, Jena, Germany Complement factor H (CFH) is a regulator of the alternative complement activation pathway. Mutations in the CFH gene are associated with atypical hemolytic uremic syndrome, membranoproliferative glomerulonephritis type II and C3 glomerulonephritis. Here, we report a 6-month-old CFH-deficient child presenting with endocapillary glomerulonephritis rather than membranoproliferative glomerulonephritis (MPGN) or C3 glomerulonephritis. Sequence analyses showed homozygosity for a novel CFH missense mutation (Pro139Ser) associated with severely decreased CFH plasma concentration (o6%) but normal mRNA splicing and expression. The father was heterozygous carrier of the mutation, but the mother was a non-carrier. Thus, a large deletion in the maternal CFH locus or uniparental isodisomy was suspected. Polymorphic markers across chromosome 1 showed homozygosity for the paternal allele in all markers and a lack of the maternal allele in six informative markers. This combined with a comparative genomic hybridization assay demonstrated paternal isodisomy. Uniparental isodisomy increases the risk of homozygous variations in other genes on the affected chromosome. -

Recurrent Inversion Toggling and Great Ape Genome Evolution
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Nat Genet Manuscript Author . Author manuscript; Manuscript Author available in PMC 2020 December 15. Published in final edited form as: Nat Genet. 2020 August ; 52(8): 849–858. doi:10.1038/s41588-020-0646-x. Recurrent inversion toggling and great ape genome evolution David Porubsky1,2,9, Ashley D. Sanders3,9, Wolfram Höps3, PingHsun Hsieh1, Arvis Sulovari1, Ruiyang Li1, Ludovica Mercuri4, Melanie Sorensen1, Shwetha C. Murali1,5, David Gordon1,5, Stuart Cantsilieris1,6, Alex A. Pollen7, Mario Ventura4, Francesca Antonacci4, Tobias Marschall8, Jan O. Korbel3, Evan E. Eichler1,5,* 1Department of Genome Sciences, University of Washington School of Medicine, Seattle, Washington, USA. 2Max Planck Institute for Informatics, Saarland Informatics Campus E1.4, Saarbrücken, Germany. 3European Molecular Biology Laboratory (EMBL), Genome Biology Unit, Heidelberg, Germany. 4Dipartimento di Biologia, Università degli Studi di Bari “Aldo Moro”, Bari, Italy. 5Howard Hughes Medical Institute, University of Washington, Seattle, Washington, USA. 6Centre for Eye Research Australia, Department of Surgery (Ophthalmology), University of Melbourne, Royal Victorian Eye and Ear Hospital, East Melbourne, Victoria, Australia. 7Department of Neurology, University of California, San Francisco (UCSF), San Francisco, California, USA. 8Institute for Medical Biometry and Bioinformatics, Medical Faculty, Heinrich Heine University Düsseldorf, Germany. 9These authors contributed equally to this work. Abstract Inversions play an important role in disease and evolution but are difficult to characterize because their breakpoints map to large repeats. We increased by six-fold the number (n = 1,069) of previously reported great ape inversions using Strand-seq and long-read sequencing. We find that the X chromosome is most enriched (2.5-fold) for inversions based on its size and duplication content. -

A Hybrid CFHR3-1 Gene Causes Familial C3 Glomerulopathy
BRIEF COMMUNICATION www.jasn.org A Hybrid CFHR3-1 Gene Causes Familial C3 Glomerulopathy †‡ Talat H. Malik,* Peter J. Lavin, Elena Goicoechea de Jorge,* Katherine A. Vernon,* | Kirsten L. Rose,* Mitali P. Patel,* Marcel de Leeuw,§ John J. Neary, Peter J. Conlon,¶ † Michelle P. Winn, and Matthew C. Pickering* *Centre for Complement and Inflammation Research, Imperial College, London, United Kingdom; †Department of Medicine, Duke University Medical Center, Durham, North Carolina; ‡Trinity Health Kidney Centre, Tallaght Hospital, Trinity College, Dublin, Ireland; §Beckman Coulter Genomics, Grenoble, France; |Department of Renal Medicine, Royal Infirmary Edinburgh, Edinburgh, United Kingdom; and ¶Department of Nephrology, Beaumont Hospital, Dublin, Ireland ABSTRACT Controlled activation of the complement system, a key component of innate immunity, that CFHR1 and CFHR3 impair comple- enables destruction of pathogens with minimal damage to host tissue. Complement ment processing within the kidney. This factor H (CFH), which inhibits complement activation, and five CFH-related proteins hypothesis would predict that an increase (CFHR1–5) compose a family of structurally related molecules. Combined deletion of in CFHR1 and CFHR3 copy number would CFHR3 and CFHR1 is common and confers a protective effect in IgA nephropathy. Here, enhance susceptibility to complement- we report an autosomal dominant complement-mediated GN associated with abnormal mediated kidney injury. Here, we report increases in copy number across the CFHR3 and CFHR1 loci. In addition to normal a novel CFHR3–1 hybrid gene located on copies of these genes, affected individuals carry a unique hybrid CFHR3–1 gene. an allele that also contained intact copies In addition to identifying an association between these genetic observations and of the CFHR1 and CFHR3 genes. -

The Impact of Complement Genes on the Risk of Late-Onset Alzheimer's
G C A T T A C G G C A T genes Article The Impact of Complement Genes on the Risk of Late-Onset Alzheimer’s Disease Sarah M. Carpanini 1,2,† , Janet C. Harwood 3,† , Emily Baker 1, Megan Torvell 1,2, The GERAD1 Consortium ‡, Rebecca Sims 3 , Julie Williams 1 and B. Paul Morgan 1,2,* 1 UK Dementia Research Institute at Cardiff University, School of Medicine, Cardiff, CF24 4HQ, UK; [email protected] (S.M.C.); [email protected] (E.B.); [email protected] (M.T.); [email protected] (J.W.) 2 Division of Infection and Immunity, School of Medicine, Systems Immunity Research Institute, Cardiff University, Cardiff, CF14 4XN, UK 3 Division of Psychological Medicine and Clinical Neurosciences, School of Medicine, Cardiff University, Cardiff, CF24 4HQ, UK; [email protected] (J.C.H.); [email protected] (R.S.) * Correspondence: [email protected] † These authors contributed equally to this work. ‡ Data used in the preparation of this article were obtained from the Genetic and Environmental Risk for Alzheimer’s disease (GERAD1) Consortium. As such, the investigators within the GERAD1 consortia contributed to the design and implementation of GERAD1 and/or provided data but did not participate in analysis or writing of this report. A full list of GERAD1 investigators and their affiliations is included in Supplementary File S1. Abstract: Late-onset Alzheimer’s disease (LOAD), the most common cause of dementia, and a huge global health challenge, is a neurodegenerative disease of uncertain aetiology. To deliver Citation: Carpanini, S.M.; Harwood, effective diagnostics and therapeutics, understanding the molecular basis of the disease is essential. -

Cell-Deposited Matrix Improves Retinal Pigment Epithelium Survival on Aged Submacular Human Bruch’S Membrane
Retinal Cell Biology Cell-Deposited Matrix Improves Retinal Pigment Epithelium Survival on Aged Submacular Human Bruch’s Membrane Ilene K. Sugino,1 Vamsi K. Gullapalli,1 Qian Sun,1 Jianqiu Wang,1 Celia F. Nunes,1 Noounanong Cheewatrakoolpong,1 Adam C. Johnson,1 Benjamin C. Degner,1 Jianyuan Hua,1 Tong Liu,2 Wei Chen,2 Hong Li,2 and Marco A. Zarbin1 PURPOSE. To determine whether resurfacing submacular human most, as cell survival is the worst on submacular Bruch’s Bruch’s membrane with a cell-deposited extracellular matrix membrane in these eyes. (Invest Ophthalmol Vis Sci. 2011;52: (ECM) improves retinal pigment epithelial (RPE) survival. 1345–1358) DOI:10.1167/iovs.10-6112 METHODS. Bovine corneal endothelial (BCE) cells were seeded onto the inner collagenous layer of submacular Bruch’s mem- brane explants of human donor eyes to allow ECM deposition. here is no fully effective therapy for the late complications of age-related macular degeneration (AMD), the leading Control explants from fellow eyes were cultured in medium T cause of blindness in the United States. The prevalence of only. The deposited ECM was exposed by removing BCE. Fetal AMD-associated choroidal new vessels (CNVs) and/or geo- RPE cells were then cultured on these explants for 1, 14, or 21 graphic atrophy (GA) in the U.S. population 40 years and older days. The explants were analyzed quantitatively by light micros- is estimated to be 1.47%, with 1.75 million citizens having copy and scanning electron microscopy. Surviving RPE cells from advanced AMD, approximately 100,000 of whom are African explants cultured for 21 days were harvested to compare bestro- American.1 The prevalence of AMD increases dramatically with phin and RPE65 mRNA expression. -

327341205.Pdf
CORE Metadata, citation and similar papers at core.ac.uk Provided by Newcastle University E-Prints Challis RC, Araujo GSR, Wong EKS, Anderson HE, Awan A, Dorman AM, Waldron M, Wilson V, Brocklebank V, Strain L, Morgan BP, Harris CL, Marchbank KJ, Goodship THJ, Kavanagh D. A De Novo Deletion in the Regulators of Complement Activation Cluster Producing a Hybrid Complement Factor H/Complement Factor H–Related 3 Gene in Atypical Hemolytic Uremic Syndrome. Journal of the American Society of Nephrology 2015, 27(6), 1617-1624. Copyright: This is the authors’ accepted manuscript of an article that has been published in its final definitive form by the American Society of Nephrology, 2015. DOI link to article: http://dx.doi.org/10.1681/ASN.2015010100 Date deposited: 05/10/2015 Embargo release date: 01 June 2017 Newcastle University ePrints - eprint.ncl.ac.uk CFH/CFHR3 hybrid gene in aHUS A De Novo Deletion in the Regulators of Complement Activation Cluster Producing a Hybrid Complement Factor H/Complement Factor H–Related 3 Gene in Atypical Hemolytic Uremic Syndrome Rachel C. Challis1*, Geisilaine S.R. Araujo1*, Edwin K.S. Wong1*, Holly E. Anderson1, Atif Awan2, Anthony M. Dorman3, Mary Waldron2, Valerie Wilson1, Vicky Brocklebank1, Lisa Strain1, B. Paul Morgan4, Claire L. Harris4, Kevin J. Marchbank5, Timothy H.J. Goodship1, David Kavanagh1 1Institute of Genetic Medicine, Newcastle University, Newcastle upon Tyne. 2Department of Nephrology, Children’s University Hospital, Dublin, Ireland. 3Beaumont Hospital and RCSI, Dublin, Ireland. 4 Institute of Infection and Immunity, Cardiff University School of Medicine, Heath Park, Cardiff, United Kingdom. 5Institute of Cellular Medicine, Newcastle University, Newcastle upon Tyne. -

10Q11.22Q11.23 Deletions and Microdeletions
Inform Network Support Rare Chromosome Disorder Support Group The Stables, Station Road West, Oxted, Surrey RH8 9EE, United Kingdom Tel: +44(0)1883 723356 [email protected] I www.rarechromo.org Join Unique for family links, information and support. Unique is a charity without government funding, existing entirely on donations and grants. If you can, please make a donation via our website at http://www.rarechromo.org/donate Please help us to help you! 10q11.22q11.23 Facebook groups www.facebook.com/groups/chromosome10disorder/ www.facebook.com/groups/152331614838414/ Deletions and Unique mentions other organisations’ message boards and websites to help families looking for information. This does not imply that we endorse their content or have any responsibility for it. Microdeletions This information guide is not a substitute for personal medical advice. Families should consult a medically qualified clinician in all matters relating to genetic diagnosis, management and health. Information on genetic changes is a very fast-moving field and while the information in this guide is believed to be the best available at the time of publication, some facts may later change. Unique does its best to keep abreast of changing information and to review its published guides as needed. This booklet was compiled by Unique (AP) and reviewed by Dr Corrina Powell, Specialist Registrar in Clinical Genetics, and Rosa Spencer- Tansley, Trainee Genetic Counsellor, Leicester Royal Infirmary Clinical Genetics Department ,UK. Version 1 2019 (AP) Copyright © Unique 2020 Rare Chromosome Disorder Support Group Charity Number 1110661 Registered in England and Wales Company Number 5460413 rarechromo.org 20 Genetic information Genetic information genetic Detailed 10q11.22q11.23 microdeletions Chromosome Position Important Microdeletion A 10q11.2q11.23 microdeletion is a rare genetic condition caused by the loss of a 10 Mb genes examples small piece of genetic material from one of the body’s 46 chromosomes – chromosome 10. -

Phenotype Informatics
Freie Universit¨atBerlin Department of Mathematics and Computer Science Phenotype informatics: Network approaches towards understanding the diseasome Sebastian Kohler¨ Submitted on: 12th September 2012 Dissertation zur Erlangung des Grades eines Doktors der Naturwissenschaften (Dr. rer. nat.) am Fachbereich Mathematik und Informatik der Freien Universitat¨ Berlin ii 1. Gutachter Prof. Dr. Martin Vingron 2. Gutachter: Prof. Dr. Peter N. Robinson 3. Gutachter: Christopher J. Mungall, Ph.D. Tag der Disputation: 16.05.2013 Preface This thesis presents research work on novel computational approaches to investigate and characterise the association between genes and pheno- typic abnormalities. It demonstrates methods for organisation, integra- tion, and mining of phenotype data in the field of genetics, with special application to human genetics. Here I will describe the parts of this the- sis that have been published in peer-reviewed journals. Often in modern science different people from different institutions contribute to research projects. The same is true for this thesis, and thus I will itemise who was responsible for specific sub-projects. In chapter 2, a new method for associating genes to phenotypes by means of protein-protein-interaction networks is described. I present a strategy to organise disease data and show how this can be used to link diseases to the corresponding genes. I show that global network distance measure in interaction networks of proteins is well suited for investigat- ing genotype-phenotype associations. This work has been published in 2008 in the American Journal of Human Genetics. My contribution here was to plan the project, implement the software, and finally test and evaluate the method on human genetics data; the implementation part was done in close collaboration with Sebastian Bauer. -

Routine Use of Clinical Exome-Based Next-Generation Sequencing for Evaluation of Patients with Thrombotic Microangiopathies
Modern Pathology (2017) 30, 1739–1747 © 2017 USCAP, Inc All rights reserved 0893-3952/17 $32.00 1739 Routine use of clinical exome-based next-generation sequencing for evaluation of patients with thrombotic microangiopathies Joseph P Gaut1,2, Sanjay Jain1,2, John D Pfeifer1, Katinka A Vigh-Conrad1, Meagan Corliss1, Mukesh K Sharma1, Jonathan W Heusel1,3 and Catherine E Cottrell1,3,4 1Department of Pathology and Immunology, Division of Anatomic and Molecular Pathology, Washington University School of Medicine, St Louis, MO, USA; 2Department of Medicine, Washington University School of Medicine, Renal Division, St Louis, MO, USA; 3Department of Genetics, Washington University School of Medicine, St Louis, MO, USA and 4Institute for Genomic Medicine, Nationwide Children’s Hospital, St Louis, MO, USA Next-generation sequencing is increasingly used for clinical evaluation of patients presenting with thrombotic microangiopathies because it allows for simultaneous interrogation of multiple complement and coagulation pathway genes known to be associated with disease. However, the diagnostic yield is undefined in routine clinical practice. Historic studies relied on case–control cohorts, did not apply current guidelines for variant pathogenicity assessment, and used targeted gene enrichment combined with next-generation sequencing. A clinically enhanced exome, targeting ~ 54 Mb, was sequenced for 73 patients. Variant analysis and interpretation were performed on genes with biological relevance in thrombotic microangiopathy (C3, CD46, CFB, CFH, CFI, DGKE, and THBD). CFHR3-CFHR1 deletion status was also assessed using multiplex ligation-dependent probe amplification. Variants were classified using American College of Medical Genetics and Genomics guidelines. We identified 5 unique novel and 14 unique rare variants in 25% (18/73) of patients, including a total of 5 pathogenic, 4 likely pathogenic, and 15 variants of uncertain clinical significance. -
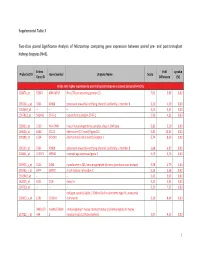
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Determining the Causes of Recessive Retinal Dystrophy
Determining the causes of recessive retinal dystrophy By Mohammed El-Sayed Mohammed El-Asrag BSc (hons), MSc (hons) Genetics and Molecular Biology Submitted in accordance with the requirements for the degree of Doctor of Philosophy The University of Leeds School of Medicine September 2016 The candidate confirms that the work submitted is his own, except where work which has formed part of jointly- authored publications has been included. The contribution of the candidate and the other authors to this work has been explicitly indicated overleaf. The candidate confirms that appropriate credit has been given within the thesis where reference has been made to the work of others. This copy has been supplied on the understanding that it is copyright material and that no quotation from the thesis may be published without proper acknowledgement. The right of Mohammed El-Sayed Mohammed El-Asrag to be identified as author of this work has been asserted by his in accordance with the Copyright, Designs and Patents Act 1988. © 2016 The University of Leeds and Mohammed El-Sayed Mohammed El-Asrag Jointly authored publications statement Chapter 3 (first results chapter) of this thesis is entirely the work of the author and appears in: Watson CM*, El-Asrag ME*, Parry DA, Morgan JE, Logan CV, Carr IM, Sheridan E, Charlton R, Johnson CA, Taylor G, Toomes C, McKibbin M, Inglehearn CF and Ali M (2014). Mutation screening of retinal dystrophy patients by targeted capture from tagged pooled DNAs and next generation sequencing. PLoS One 9(8): e104281. *Equal first- authors. Shevach E, Ali M, Mizrahi-Meissonnier L, McKibbin M, El-Asrag ME, Watson CM, Inglehearn CF, Ben-Yosef T, Blumenfeld A, Jalas C, Banin E and Sharon D (2015). -

10Q11.22Q11.23 Deletions and Microdeletions
10q11.22q11.23 Deletions and Microdeletions rarechromo.org Genetic information Genetic 10q11.22q11.23 microdeletions A 10q11.2q11.23 microdeletion is a rare genetic condition caused by the loss of a small piece of genetic material from one of the body’s 46 chromosomes – chromosome 10. For typical development, chromosomes should contain the expected amount of genetic material. Like many other chromosome disorders, having a missing piece of chromosome 10 may affect a child’s health, development and intellectual abilities. The symptoms observed in people with a 10q11.22q10.23 microdeletion are variable and depend on a number of factors including what and how much genetic material is missing. Background on chromosomes Our bodies are made up of many different types of cells, almost all of which contain the same chromosomes. Each chromosome consists of DNA that codes for our genes. Genes can be thought of as individual instructions that tell the body how to develop, grow and function. Chromosomes come in pairs with one member of each chromosome pair being inherited from each parent. Most of our cells have 23 pairs of chromosomes (a total of 46) as shown in this image. Eggs and sperm, however, have 23 unpaired chromosomes. When an egg and sperm join together at conception, the chromosomes pair up to make a total of 46. Chromosome pairs are numbered 1 to 22 Chromosomes pairs 1-22, X and Y (male) and the 23rd pair comprises the sex Chromosome pair 10 is circled in red chromosomes that determine biological sex. They can be viewed under a microscope and arranged roughly according to size as the image shows.