Description of a New Member of the Family Erysipelotrichaceae: Clostridium Fusiformis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The 2014 Golden Gate National Parks Bioblitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event
National Park Service U.S. Department of the Interior Natural Resource Stewardship and Science The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 ON THIS PAGE Photograph of BioBlitz participants conducting data entry into iNaturalist. Photograph courtesy of the National Park Service. ON THE COVER Photograph of BioBlitz participants collecting aquatic species data in the Presidio of San Francisco. Photograph courtesy of National Park Service. The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 Elizabeth Edson1, Michelle O’Herron1, Alison Forrestel2, Daniel George3 1Golden Gate Parks Conservancy Building 201 Fort Mason San Francisco, CA 94129 2National Park Service. Golden Gate National Recreation Area Fort Cronkhite, Bldg. 1061 Sausalito, CA 94965 3National Park Service. San Francisco Bay Area Network Inventory & Monitoring Program Manager Fort Cronkhite, Bldg. 1063 Sausalito, CA 94965 March 2016 U.S. Department of the Interior National Park Service Natural Resource Stewardship and Science Fort Collins, Colorado The National Park Service, Natural Resource Stewardship and Science office in Fort Collins, Colorado, publishes a range of reports that address natural resource topics. These reports are of interest and applicability to a broad audience in the National Park Service and others in natural resource management, including scientists, conservation and environmental constituencies, and the public. The Natural Resource Report Series is used to disseminate comprehensive information and analysis about natural resources and related topics concerning lands managed by the National Park Service. -

Supplementary Information for Acute Infection of Mice with Clostridium
IMM-2013-2862, Sadighi Akha et al-Supplementary Information 1 Supplementary Information for Acute infection of mice with Clostridium difficile leads to eIF2α phosphorylation and pro-survival signalling as part of the mucosal inflammatory response Amir A. Sadighi Akha1, Casey M. Theriot2, John R. Erb-Downward1, Andrew J. McDermott3, Nicole R. Falkowski1, Heather M. Tyra4, D. Thomas Rutkowski4, Vincent B. Young2,3 and Gary B. Huffnagle1,3* IMM-2013-2862, Sadighi Akha et al-Supplementary Information 2 Supplementary Text Materials and Methods for Figures S1 and S2 DNA isolation, amplicon library preparation and 454 pyrosequencing- The DNeasy blood and tissue kit (Qiagen) was used to extract genomic DNA from the caecal and colonic contents obtained from untreated and C. difficile-infected mice. The extraction was performed according to the manufacturer’s instructions except for the following modifications: adding a bead-beating step using UltraClean fecal DNA bead tubes (Mo Bio Laboratories, Carlsbad, CA); doubling the amount of ATL buffer and the proteinase K used in the protocol; and decreasing by half the amount of the AE buffer used to elute the DNA. Subsequently, the V3, V4 and V5 hyper-variable regions of the 16S ribosomal RNA gene in each of the samples were targeted for amplification with the 357F and 929R primer sets [1]. Amplicons were purified with the Agencourt AMPure XP PCR purification system (Beckman Coulter, Indianapolis, IN), and quantified with the Quant- iT PicoGreen dsDNA kit (Life Technologies) to obtain an equal pool for pyrosequencing. They were then sequenced on a Roche 454 GS Junior Titanium platform according to the manufacturer’s specifications. -

The Oral Microbiome of Healthy Japanese People at the Age of 90
applied sciences Article The Oral Microbiome of Healthy Japanese People at the Age of 90 Yoshiaki Nomura 1,* , Erika Kakuta 2, Noboru Kaneko 3, Kaname Nohno 3, Akihiro Yoshihara 4 and Nobuhiro Hanada 1 1 Department of Translational Research, Tsurumi University School of Dental Medicine, Kanagawa 230-8501, Japan; [email protected] 2 Department of Oral bacteriology, Tsurumi University School of Dental Medicine, Kanagawa 230-8501, Japan; [email protected] 3 Division of Preventive Dentistry, Faculty of Dentistry and Graduate School of Medical and Dental Science, Niigata University, Niigata 951-8514, Japan; [email protected] (N.K.); [email protected] (K.N.) 4 Division of Oral Science for Health Promotion, Faculty of Dentistry and Graduate School of Medical and Dental Science, Niigata University, Niigata 951-8514, Japan; [email protected] * Correspondence: [email protected]; Tel.: +81-45-580-8462 Received: 19 August 2020; Accepted: 15 September 2020; Published: 16 September 2020 Abstract: For a healthy oral cavity, maintaining a healthy microbiome is essential. However, data on healthy microbiomes are not sufficient. To determine the nature of the core microbiome, the oral-microbiome structure was analyzed using pyrosequencing data. Saliva samples were obtained from healthy 90-year-old participants who attended the 20-year follow-up Niigata cohort study. A total of 85 people participated in the health checkups. The study population consisted of 40 male and 45 female participants. Stimulated saliva samples were obtained by chewing paraffin wax for 5 min. The V3–V4 hypervariable regions of the 16S ribosomal RNA (rRNA) gene were amplified by PCR. -

Fatty Acid Diets: Regulation of Gut Microbiota Composition and Obesity and Its Related Metabolic Dysbiosis
International Journal of Molecular Sciences Review Fatty Acid Diets: Regulation of Gut Microbiota Composition and Obesity and Its Related Metabolic Dysbiosis David Johane Machate 1, Priscila Silva Figueiredo 2 , Gabriela Marcelino 2 , Rita de Cássia Avellaneda Guimarães 2,*, Priscila Aiko Hiane 2 , Danielle Bogo 2, Verônica Assalin Zorgetto Pinheiro 2, Lincoln Carlos Silva de Oliveira 3 and Arnildo Pott 1 1 Graduate Program in Biotechnology and Biodiversity in the Central-West Region of Brazil, Federal University of Mato Grosso do Sul, Campo Grande 79079-900, Brazil; [email protected] (D.J.M.); [email protected] (A.P.) 2 Graduate Program in Health and Development in the Central-West Region of Brazil, Federal University of Mato Grosso do Sul, Campo Grande 79079-900, Brazil; pri.fi[email protected] (P.S.F.); [email protected] (G.M.); [email protected] (P.A.H.); [email protected] (D.B.); [email protected] (V.A.Z.P.) 3 Chemistry Institute, Federal University of Mato Grosso do Sul, Campo Grande 79079-900, Brazil; [email protected] * Correspondence: [email protected]; Tel.: +55-67-3345-7416 Received: 9 March 2020; Accepted: 27 March 2020; Published: 8 June 2020 Abstract: Long-term high-fat dietary intake plays a crucial role in the composition of gut microbiota in animal models and human subjects, which affect directly short-chain fatty acid (SCFA) production and host health. This review aims to highlight the interplay of fatty acid (FA) intake and gut microbiota composition and its interaction with hosts in health promotion and obesity prevention and its related metabolic dysbiosis. -

Gut Microbiota Differences Between Psoriatic Arthritis and Undifferentiated Arthritis Patients
Gut microbiota differences between psoriatic arthritis and undifferentiated arthritis patients Chung-Yuan Hsu Chang Gung Memorial Hospital Kaohsiung Branch Ho-Chang Kuo Chang Gung Memorial Hospital Kaohsiung Branch YU-JIH Su ( [email protected] ) Chang Gung Memorial Hospital Kaohsiung Branch https://orcid.org/0000-0002-7274-3458 Research article Keywords: Psoriatic arthritis, undifferentiated arthritis, enthesitis, dactylitis, microbiota Posted Date: February 28th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-15400/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/15 Abstract Introduction Psoriatic arthritis (PSA) is a form of immune-mediated inammatory arthritis. Studying the gut microbiota of PSA patients may offer new insights into the pathophysiology of this inammatory joint disease. We designed a prospective study to examine gut microbiome from patients with PSA, primarily with enthesitis and dactylitis, and compared the data with undifferentiated arthritis patients (NO PSA), without enthesitis or dactylitis. Methods We enrolled nine PSA patients and 10 NO PSA patients in this study. The fecal samples were investigated by using 16S rRNA amplicon sequencing, followed by bioinformatics and statistical analyses. Results None of the available objective clinical laboratory data could differentiate PSA from the NO PSA subgroup. The microbiota result shows that Family: XIII_AD3011 is signicantly higher in NO PSA patients than PSA patients’ stool samples (p=0.039). Megasphaera elsdenii in the PSA was 10000 times higher than in the NO PSA group. Conclusion Our results demonstrated high intra-group homogeneous and high inter-group heterogeneous microbiota. The clinical symptoms of either enthesitis or dactylitis link to the specic microbiota in the current study. -

Bovine Host Genome Acts on Speci C Metabolism, Communication and Genetic Processes of Rumen Microbes Host-Genomically Linked To
Bovine host genome acts on specic metabolism, communication and genetic processes of rumen microbes host-genomically linked to methane emissions Marina Martínez-Álvaro SRUC https://orcid.org/0000-0003-2295-5839 Marc Auffret SRUC Carol-Anne Duthie SRUC Richard Dewhurst SRUC Matthew Cleveland Genus plc Mick Watson Roslin Institute https://orcid.org/0000-0003-4211-0358 Rainer Roehe ( [email protected] ) SRUC https://orcid.org/0000-0002-4880-3756 Article Keywords: bovine host genome, rumen, CH4 Posted Date: May 17th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-290150/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License 1 Bovine host genome acts on specific metabolism, communication and 2 genetic processes of rumen microbes host-genomically linked to methane 3 emissions 4 Marina Martínez-Álvaro1, Marc D. Auffret1, Carol-Anne Duthie1, Richard J. Dewhurst1, 5 Matthew A. Cleveland2, Mick Watson3 and Rainer Roehe*1 6 1Scotland’s Rural College, Edinburgh, UK 7 2Genus plc, DeForest, WI, USA 8 3The Roslin Institute and the Royal (Dick) School of Veterinary Studies, University of 9 Edinburgh, UK 10 11 *Corresponding author. Email: [email protected] 12 13 14 Introductory paragraph 15 Whereas recent studies in different species showed that the host genome shapes the microbial 16 community profile, our new research strategy revealed substantial host genomic control of 17 comprehensive functional microbial processes in the rumen of bovines by utilising microbial 18 gene profiles from whole metagenomic sequencing. Of 1,107/225/1,141 rumen microbial 19 genera/metagenome assembled uncultured genomes (RUGs)/genes identified, 203/16/352 20 were significantly (P<2.02 x10-5) heritable (0.13 to 0.61), revealing substantial variation in 21 host genomic control. -

A Metagenomic Study of the Oral and Gut Microbiome in Crohn’S Disease Shijia Hu1* , Eileen Png2, Michelle Gowans3, David E
Hu et al. Gut Pathog (2021) 13:13 https://doi.org/10.1186/s13099-021-00409-5 Gut Pathogens RESEARCH Open Access Ectopic gut colonization: a metagenomic study of the oral and gut microbiome in Crohn’s disease Shijia Hu1* , Eileen Png2, Michelle Gowans3, David E. H. Ong3, Paola Florez de Sessions2, Jie Song2 and Niranjan Nagarajan2,4 Abstract Background: This study aims to characterize, the gut and oral microbiome in Asian subjects with Crohn’s disease (CD) using whole genome shotgun sequencing, thereby allowing for strain-level comparison. Methods: A case–control study with age, sex and ethnicity matched healthy controls was conducted. CD subjects were limited to well-controlled patients without oral manifestations. Fecal and saliva samples were collected for char- acterization of gut and oral microbiome respectively. Microbial DNA were extracted, libraries prepared and sequenced reads profled. Taxonomic diversity, taxonomic association, strain typing and microbial gene pathway analyses were conducted. Results: The study recruited 25 subjects with CD and 25 healthy controls. The oral microbe Streptococcus salivarius was found to be enriched and of concordant strains in the gut and oral microbiome of Crohn’s disease subjects. This was more likely in CD subjects with higher Crohn’s Disease Activity Index (184.3 2.9 vs 67.1 82.5, p 0.012) and active disease status (Diarrhoea/abdominal pain/blood-in-stool/fever and fatigue)± (p 0.016).± Gut species= found to be signifcantly depleted in CD compared to control (Relative abundance: Median[Range])= include: Faecalibacterium prausnitzii (0.03[0.00–4.56] vs 13.69[5.32–18.71], p 0.010), Roseburia inulinivorans (0.00[0.00–0.03] vs 0.21[0.01–0.53], p 0.010) and Alistipes senegalensis (0.00[0.00–0.00]= vs 0.00[0.00–0.02], p 0.029). -

WO 2018/064165 A2 (.Pdf)
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2018/064165 A2 05 April 2018 (05.04.2018) W !P O PCT (51) International Patent Classification: Published: A61K 35/74 (20 15.0 1) C12N 1/21 (2006 .01) — without international search report and to be republished (21) International Application Number: upon receipt of that report (Rule 48.2(g)) PCT/US2017/053717 — with sequence listing part of description (Rule 5.2(a)) (22) International Filing Date: 27 September 2017 (27.09.2017) (25) Filing Language: English (26) Publication Langi English (30) Priority Data: 62/400,372 27 September 2016 (27.09.2016) US 62/508,885 19 May 2017 (19.05.2017) US 62/557,566 12 September 2017 (12.09.2017) US (71) Applicant: BOARD OF REGENTS, THE UNIVERSI¬ TY OF TEXAS SYSTEM [US/US]; 210 West 7th St., Austin, TX 78701 (US). (72) Inventors: WARGO, Jennifer; 1814 Bissonnet St., Hous ton, TX 77005 (US). GOPALAKRISHNAN, Vanch- eswaran; 7900 Cambridge, Apt. 10-lb, Houston, TX 77054 (US). (74) Agent: BYRD, Marshall, P.; Parker Highlander PLLC, 1120 S. Capital Of Texas Highway, Bldg. One, Suite 200, Austin, TX 78746 (US). (81) Designated States (unless otherwise indicated, for every kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. -

Bulleidia Extructa Gen. Nov., Sp. Nov., Isolated from the Oral Cavity
International Journal of Systematic and Evolutionary Microbiology (2000), 50, 979–983 Printed in Great Britain Bulleidia extructa gen. nov., sp. nov., isolated from the oral cavity Julia Downes,1 Bente Olsvik,†2 Sarah J. Hiom,1 David A. Spratt,1 Sarah L. Cheeseman,1 Ingar Olsen,2 Andrew J. Weightman3 and William G. Wade1 Author for correspondence: William G. Wade. Tel: j44 207 955 2849. Fax: j44 207 955 2847. e-mail: william.wade!kcl.ac.uk 1 Oral Microbiology Unit, Five strains of anaerobic non-sporing Gram-positive bacilli isolated from Division of Oral Medicine, advanced periodontitis (four strains) and a dentoalveolar abscess (one strain) Pathology, Microbiology and Immunology, Guy’s, that did not correspond to existing species were subjected to phenotypic and King’s and St Thomas’ genetic characterization. Following 16S rDNA sequence analysis, they were Dental Institute, found to constitute a novel branch of the low GMC Gram-positive division of King’s College London, Guy’s Hospital, London the phylogenetic tree related to Erysipelothrix rhusiopathiae and Holdemania SE1 9RT, UK filiformis. A new genus Bulleidia, and the species Bulleidia extructa, are 2 Institutt for oral biologi, proposed. Growth of B. extructa in broth media was poor but was enhanced by University of Oslo, the addition of fructose, glucose or maltose together with Tween 80. Glucose N-0316 Oslo, Norway and maltose were fermented and arginine was hydrolysed. Acetate, lactate 3 Cardiff School of and trace amounts of succinate were the end products of glucose Biosciences, Cardiff fermentation. The GMC content of the DNA of the type strain is 38 mol%. -

Perilla Frutescens Leaf Alters the Rumen Microbial Community of Lactating Dairy Cows
microorganisms Article Perilla frutescens Leaf Alters the Rumen Microbial Community of Lactating Dairy Cows Zhiqiang Sun, Zhu Yu and Bing Wang * College of Grass Science and Technology, China Agricultural University, Beijing 100193, China; [email protected] (Z.S.); [email protected] (Z.Y.) * Correspondence: [email protected] Received: 25 September 2019; Accepted: 12 November 2019; Published: 13 November 2019 Abstract: Perilla frutescens (L.) Britt., an annual herbaceous plant, has antibacterial, anti-inflammation, and antioxidant properties. To understand the effects of P. frutescens leaf on the ruminal microbial ecology of cattle, Illumina MiSeq 16S rRNA sequencing technology was used. Fourteen cows were used in a randomized complete block design trial. Two diets were fed to these cattle: a control diet (CON); and CON supplemented with 300 g/d P. frutescens leaf (PFL) per cow. Ruminal fluid was sampled at the end of the experiment for microbial DNA extraction. Overall, our findings revealed that supplementation with PFL could increase ruminal fluid pH value. The ruminal bacterial community of cattle was dominated by Bacteroidetes, Firmicutes, and Proteobacteria. The addition of PFL had a positive effect on Firmicutes, Actinobacteria, and Spirochaetes, but had no effect on Bacteroidetes and Proteobacteria compared with the CON. The supplementation with PFL significantly increased the abundance of Marvinbryantia, Acetitomaculum, Ruminococcus gauvreauii, Eubacterium coprostanoligenes, Selenomonas_1, Pseudoscardovia, norank_f__Muribaculaceae, and Sharpea, and decreased the abundance of Treponema_2 compared to CON. Eubacterium coprostanoligenes, and norank_f__Muribaculaceae were positively correlated with ruminal pH value. It was found that norank_f__Muribaculaceae and Acetitomaculum were positively correlated with milk yield, indicating that these different genera are PFL associated bacteria. -

The Gut Microbiome of the Sea Urchin, Lytechinus Variegatus, from Its Natural Habitat Demonstrates Selective Attributes of Micro
FEMS Microbiology Ecology, 92, 2016, fiw146 doi: 10.1093/femsec/fiw146 Advance Access Publication Date: 1 July 2016 Research Article RESEARCH ARTICLE The gut microbiome of the sea urchin, Lytechinus variegatus, from its natural habitat demonstrates selective attributes of microbial taxa and predictive metabolic profiles Joseph A. Hakim1,†, Hyunmin Koo1,†, Ranjit Kumar2, Elliot J. Lefkowitz2,3, Casey D. Morrow4, Mickie L. Powell1, Stephen A. Watts1,∗ and Asim K. Bej1,∗ 1Department of Biology, University of Alabama at Birmingham, 1300 University Blvd, Birmingham, AL 35294, USA, 2Center for Clinical and Translational Sciences, University of Alabama at Birmingham, Birmingham, AL 35294, USA, 3Department of Microbiology, University of Alabama at Birmingham, Birmingham, AL 35294, USA and 4Department of Cell, Developmental and Integrative Biology, University of Alabama at Birmingham, 1918 University Blvd., Birmingham, AL 35294, USA ∗Corresponding authors: Department of Biology, University of Alabama at Birmingham, 1300 University Blvd, CH464, Birmingham, AL 35294-1170, USA. Tel: +1-(205)-934-8308; Fax: +1-(205)-975-6097; E-mail: [email protected]; [email protected] †These authors contributed equally to this work. One sentence summary: This study describes the distribution of microbiota, and their predicted functional attributes, in the gut ecosystem of sea urchin, Lytechinus variegatus, from its natural habitat of Gulf of Mexico. Editor: Julian Marchesi ABSTRACT In this paper, we describe the microbial composition and their predictive metabolic profile in the sea urchin Lytechinus variegatus gut ecosystem along with samples from its habitat by using NextGen amplicon sequencing and downstream bioinformatics analyses. The microbial communities of the gut tissue revealed a near-exclusive abundance of Campylobacteraceae, whereas the pharynx tissue consisted of Tenericutes, followed by Gamma-, Alpha- and Epsilonproteobacteria at approximately equal capacities. -
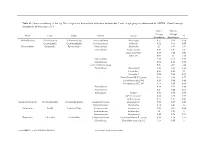
Genus Contributing to the Top 70% of Significant Dissimilarity of Bacteria Between Day 7 and 18 Age Groups As Determined by SIMPER
Table S1: Genus contributing to the top 70% of significant dissimilarity of bacteria between day 7 and 18 age groups as determined by SIMPER. Overall average dissimilarity between ages is 51%. Day 7 Day 18 Average Average Phyla Class Order Family Genus % Abundance Abundance Actinobacteria Actinobacteria Actinomycetales Actinomycetaceae Actinomyces 0.57 0.24 0.74 Coriobacteriia Coriobacteriales Coriobacteriaceae Collinsella 0.51 0.61 0.57 Bacteroidetes Bacteroidia Bacteroidales Bacteroidaceae Bacteroides 2.1 1.63 1.03 Marinifilaceae Butyricimonas 0.82 0.81 0.9 Sanguibacteroides 0.08 0.42 0.59 CAG-873 0.01 1.1 1.68 Marinifilaceae 0.38 0.78 0.91 Marinifilaceae 0.78 1.04 0.97 p-2534-18B5 gut group 0.06 0.7 1.04 Prevotellaceae Alloprevotella 0.46 0.83 0.89 Prevotella 2 0.95 1.11 1.3 Prevotella 7 0.22 0.42 0.56 Prevotellaceae NK3B31 group 0.62 0.68 0.77 Prevotellaceae UCG-003 0.26 0.64 0.89 Prevotellaceae UCG-004 0.11 0.48 0.68 Prevotellaceae 0.64 0.52 0.89 Prevotellaceae 0.27 0.42 0.55 Rikenellaceae Alistipes 0.39 0.62 0.77 dgA-11 gut group 0.04 0.38 0.56 RC9 gut group 0.79 1.17 0.95 Epsilonbacteraeota Campylobacteria Campylobacterales Campylobacteraceae Campylobacter 0.58 0.83 0.97 Helicobacteraceae Helicobacter 0.12 0.41 0.6 Firmicutes Bacilli Lactobacillales Enterococcaceae Enterococcus 0.36 0.31 0.64 Lactobacillaceae Lactobacillus 1.4 1.24 1 Streptococcaceae Streptococcus 0.82 0.58 0.53 Firmicutes Clostridia Clostridiales Christensenellaceae Christensenellaceae R-7 group 0.38 1.36 1.56 Clostridiaceae Clostridium sensu stricto 1 1.16 0.58