Placozoa and Cnidaria Are Sister Taxa
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Number of Living Species in Australia and the World
Numbers of Living Species in Australia and the World 2nd edition Arthur D. Chapman Australian Biodiversity Information Services australia’s nature Toowoomba, Australia there is more still to be discovered… Report for the Australian Biological Resources Study Canberra, Australia September 2009 CONTENTS Foreword 1 Insecta (insects) 23 Plants 43 Viruses 59 Arachnida Magnoliophyta (flowering plants) 43 Protoctista (mainly Introduction 2 (spiders, scorpions, etc) 26 Gymnosperms (Coniferophyta, Protozoa—others included Executive Summary 6 Pycnogonida (sea spiders) 28 Cycadophyta, Gnetophyta under fungi, algae, Myriapoda and Ginkgophyta) 45 Chromista, etc) 60 Detailed discussion by Group 12 (millipedes, centipedes) 29 Ferns and Allies 46 Chordates 13 Acknowledgements 63 Crustacea (crabs, lobsters, etc) 31 Bryophyta Mammalia (mammals) 13 Onychophora (velvet worms) 32 (mosses, liverworts, hornworts) 47 References 66 Aves (birds) 14 Hexapoda (proturans, springtails) 33 Plant Algae (including green Reptilia (reptiles) 15 Mollusca (molluscs, shellfish) 34 algae, red algae, glaucophytes) 49 Amphibia (frogs, etc) 16 Annelida (segmented worms) 35 Fungi 51 Pisces (fishes including Nematoda Fungi (excluding taxa Chondrichthyes and (nematodes, roundworms) 36 treated under Chromista Osteichthyes) 17 and Protoctista) 51 Acanthocephala Agnatha (hagfish, (thorny-headed worms) 37 Lichen-forming fungi 53 lampreys, slime eels) 18 Platyhelminthes (flat worms) 38 Others 54 Cephalochordata (lancelets) 19 Cnidaria (jellyfish, Prokaryota (Bacteria Tunicata or Urochordata sea anenomes, corals) 39 [Monera] of previous report) 54 (sea squirts, doliolids, salps) 20 Porifera (sponges) 40 Cyanophyta (Cyanobacteria) 55 Invertebrates 21 Other Invertebrates 41 Chromista (including some Hemichordata (hemichordates) 21 species previously included Echinodermata (starfish, under either algae or fungi) 56 sea cucumbers, etc) 22 FOREWORD In Australia and around the world, biodiversity is under huge Harnessing core science and knowledge bases, like and growing pressure. -

Simple Animals Sponges and Placozoa the Twig of the Tree That Is the Animals the Tour Begins
Animal Kingdom Simple animals Sponges and placozoa Tom Hartman Asymmetry www.tuatara9.co.uk Animal form and function 1 Module 111121 2 The twig of the tree that is the animals Animals All other animals Animals Sponges Choanoflagellates Choanoflagellates Fungi Fungi 3 4 All other animals Animals (bilateralia) Radiata Sponges Choanoflagellates Fungi The tour begins 5 6 1 Animal Kingdom The Phylum Quirky phyla • In the standard Linnean system (and taxonomic systems • There are 39 animal phyla (+/- 10!) based on it), a Phylum is the taxonomic category between Kingdom and Class. • The Micrognathozoa were • A phylum is a major ranking of organisms, defined discovered in 2000 (1 species)in according to the most basic body-parts shared by that springs in Greenland. group. But we must include the creatures and their • Xenoturbellida removed from the common ancestor. molluscs and moved to the – Chordata (animals with a notochord - vertebrates and others), – Arthropoda (animals with a jointed exoskeleton) dueterostomes when DNA – Mollusca (animals with a shell-secreting mantle), evidence was discounted due to – Angiosperma (flowering plants), and so on. what it had eaten! – A number of traditional Phyla - e.g. Protozoa, possibly Arthropoda - • Cycliophora discovered on the are probably invalid (polyphyletic). lips of lobsters in 1995. 7 8 The Class Our journey • In the Linnean system (and taxonomic systems Kingdom Group Phylum based on it), a Class is the taxonomic category Porifera between Phylum and Order. Placozoa • A class is a major group of organisms, e.g. Cnidaria Mammalia, Gastropoda, Insecta, etc that Parazoa Ctenophora contains a large number of different Radiata Platyhelminthes sublineages, but have shared characteristics in Animalia Protostome Rotifera common e.g. -

Comprehensive Phylogenomic Analyses Resolve Cnidarian Relationships and the Origins of Key Organismal Traits
Comprehensive phylogenomic analyses resolve cnidarian relationships and the origins of key organismal traits Ehsan Kayal1,2, Bastian Bentlage1,3, M. Sabrina Pankey5, Aki H. Ohdera4, Monica Medina4, David C. Plachetzki5*, Allen G. Collins1,6, Joseph F. Ryan7,8* Authors Institutions: 1. Department of Invertebrate Zoology, National Museum of Natural History, Smithsonian Institution 2. UPMC, CNRS, FR2424, ABiMS, Station Biologique, 29680 Roscoff, France 3. Marine Laboratory, university of Guam, UOG Station, Mangilao, GU 96923, USA 4. Department of Biology, Pennsylvania State University, University Park, PA, USA 5. Department of Molecular, Cellular and Biomedical Sciences, University of New Hampshire, Durham, NH, USA 6. National Systematics Laboratory, NOAA Fisheries, National Museum of Natural History, Smithsonian Institution 7. Whitney Laboratory for Marine Bioscience, University of Florida, St Augustine, FL, USA 8. Department of Biology, University of Florida, Gainesville, FL, USA PeerJ Preprints | https://doi.org/10.7287/peerj.preprints.3172v1 | CC BY 4.0 Open Access | rec: 21 Aug 2017, publ: 21 Aug 20171 Abstract Background: The phylogeny of Cnidaria has been a source of debate for decades, during which nearly all-possible relationships among the major lineages have been proposed. The ecological success of Cnidaria is predicated on several fascinating organismal innovations including symbiosis, colonial body plans and elaborate life histories, however, understanding the origins and subsequent diversification of these traits remains difficult due to persistent uncertainty surrounding the evolutionary relationships within Cnidaria. While recent phylogenomic studies have advanced our knowledge of the cnidarian tree of life, no analysis to date has included genome scale data for each major cnidarian lineage. Results: Here we describe a well-supported hypothesis for cnidarian phylogeny based on phylogenomic analyses of new and existing genome scale data that includes representatives of all cnidarian classes. -

University of Copenhagen, Zoological Museum, Review Universitetsparken 15, DK-2100 Copenhagen, Denmark CN, 0000-0001-6898-7655 Cite This Article: Nielsen C
Early animal evolution a morphologist's view Nielsen, Claus Published in: Royal Society Open Science DOI: 10.1098/rsos.190638 Publication date: 2019 Document version Publisher's PDF, also known as Version of record Document license: CC BY Citation for published version (APA): Nielsen, C. (2019). Early animal evolution: a morphologist's view. Royal Society Open Science, 6(7), 1-10. [190638]. https://doi.org/10.1098/rsos.190638 Download date: 30. sep.. 2021 Early animal evolution: a morphologist’s view royalsocietypublishing.org/journal/rsos Claus Nielsen The Natural History Museum of Denmark, University of Copenhagen, Zoological Museum, Review Universitetsparken 15, DK-2100 Copenhagen, Denmark CN, 0000-0001-6898-7655 Cite this article: Nielsen C. 2019 Early animal evolution: a morphologist’s view. R. Soc. open sci. Two hypotheses for the early radiation of the metazoans are vividly discussed in recent phylogenomic studies, the ‘Porifera- 6: 190638. first’ hypothesis, which places the poriferans as the sister group http://dx.doi.org/10.1098/rsos.190638 of all other metazoans, and the ‘Ctenophora-first’ hypothesis, which places the ctenophores as the sister group to all other metazoans. It has been suggested that an analysis of morphological characters (including specific molecules) could Received: 5 April 2019 throw additional light on the controversy, and this is the aim of Accepted: 4 July 2019 this paper. Both hypotheses imply independent evolution of nervous systems in Planulozoa and Ctenophora. The Porifera- first hypothesis implies no homoplasies or losses of major characters. The Ctenophora-first hypothesis shows no important synapomorphies of Porifera, Planulozoa and Placozoa. It implies Subject Category: either independent evolution, in Planulozoa and Ctenophora, of Biology (whole organism) a new digestive system with a gut with extracellular digestion, which enables feeding on larger organisms, or the subsequent Subject Areas: loss of this new gut in the Poriferans (and the re-evolution of the evolution collar complex). -

Xenacoelomorpha's Significance for Understanding Bilaterian Evolution
Available online at www.sciencedirect.com ScienceDirect Xenacoelomorpha’s significance for understanding bilaterian evolution Andreas Hejnol and Kevin Pang The Xenacoelomorpha, with its phylogenetic position as sister biology models are the fruitfly Drosophila melanogaster and group of the Nephrozoa (Protostomia + Deuterostomia), plays the nematode Caenorhabditis elegans, in which basic prin- a key-role in understanding the evolution of bilaterian cell types ciples of developmental processes have been studied in and organ systems. Current studies of the morphological and great detail. It might be because the field of evolutionary developmental diversity of this group allow us to trace the developmental biology — EvoDevo — has its origin in evolution of different organ systems within the group and to developmental biology and not evolutionary biology that reconstruct characters of the most recent common ancestor of species under investigation are often called ‘model spe- Xenacoelomorpha. The disparity of the clade shows that there cies’. Criteria for selected representative species are cannot be a single xenacoelomorph ‘model’ species and primarily the ease of access to collected material and strategic sampling is essential for understanding the evolution their ability to be cultivated in the lab [1]. In some cases, of major traits. With this strategy, fundamental insights into the a supposedly larger number of ancestral characters or a evolution of molecular mechanisms and their role in shaping dominant role in ecosystems have played an additional animal organ systems can be expected in the near future. role in selecting model species. These arguments were Address used to attract sufficient funding for genome sequencing Sars International Centre for Marine Molecular Biology, University of and developmental studies that are cost-intensive inves- Bergen, Thormøhlensgate 55, 5008 Bergen, Norway tigations. -

Sponges) and Phylum Cnidaria (Jellyfish, Sea Anemones and Corals
4/14/2014 Kingdom Animalia: Phylum Porifera (sponges) and Phylum Cnidaria (jellyfish, sea anemones and corals) 1 4/14/2014 Animals have different types of symmetry AsymmetricalÆ Radial Æ Bilateral Æ Embryo development provides information about how animal groups are related Blastula: hallow with a single layer of cells Gastrula: results in two layers of cells and cavity (gut) with one opening (blastopore) Cavity reaches the other side and the gut is like a tube Some cells from a third layer of cells A second cavityyg forms between the gut and the outside of the animal 2 4/14/2014 Animals have different number of true tissue layers and different type of gut No true tissuesÆ Two tissue layers Æ Three tissue layersÆ No gutÆ Sac like gutÆ Tube like gutÆ Phylum Porifera: Simplest of Animals Sponges: No tissues, no symmetry Intracellular digestion, no digestive system or cavity Collar cells or choanocytes Support by spicules or spongin fibers 3 4/14/2014 Procedure 1 • Grantia sponge Locate osculum • Sponge spicules Bell Labs Research on Deep-Sea Sponge Yields Substantial Mechanical Engineering Insights 4 4/14/2014 Medications from Sponges Thirty percent of all potential new natural medicine has been isolated in sponges. About 75% of the recently registered and patented material to fight cancer comes from sponges. Furthermore, it appears that medicine from sponges helps, for example, asthma and psoriasis; therefore it offers enormous possibilities for research. Eribulin, a novel chemotherapy drug derived from a sea sponge, improves survival in heavily-pretreated metastatic breast cancer. Phylum Cnidaria Coral Sea Anemone Man-of-war Hydra Jellyfish 5 4/14/2014 Phylum Cnidaria Tissues: Endoderm Ectoderm Type of gut: Symmetry: Radial Cnidocytes or Stinging cells Polyp or Medusa form Importance Some jellyfish are considered a delicacy Corals: Medicines cabinets for the 21st century cancer cell inhibitor Sunscreen 6 4/14/2014 Procedure 2 2. -

Animal Evolution: Looking for the First Nervous System
Dispatch R655 cellular or symbiotic functions, cannot genome reduction in the symbiont, 10. Nakabachi, A., Ishida, K., Hongoh, Y., Ohkuma, M., and Miyagishima, S. (2014). be lost without harmful or fatal results. HGT from various sources to the host Aphid gene of bacterial origin encodes a But these genes can be replaced, genome to maintain symbiont function, protein transported to an obligate and perhaps ever more easily as and now the targeting of protein endosymbiont. Curr. Biol. 24, R640–R641. 11. Moran, N.A., and Jarvik, T. (2010). Lateral the interaction networks of the products from host to symbiont has transfer of genes from fungi underlies endosymbiont reduce in complexity, even been found [4,10]. These make carotenoid production in aphids. Science 328, 624–627. thus reducing pressures on proteins clean separation of endosymbiont from 12. Jorgenson, M.A., Chen, Y., Yahashiri, A., to co-evolve. In a sense, when genes organelle more difficult to see, Popham, D.L., and Weiss, D.S. (2014). The are transferred from symbiont or prompting us not to look for the point bacterial septal ring protein RlpA is a lytic transglycosylase that contributes to rod organelle to the host nuclear genome when a symbiont ‘becomes’ an shape and daughter cell separation in and their proteins targeted back, this is organelle, but rather to ask, ‘Is there Pseudomonas aeruginosa. Mol. Microbiol. 93, 113–128. a compensatory change [4,9,14,17]. really anything so special about 13. Acuna, R., Padilla, B.E., Florez-Ramos, C.P., But other compensatory transfers that organelles?’ Rubio, J.D., Herrera, J.C., Benavides, P., affect a symbiont or organelle can also Lee, S.J., Yeats, T.H., Egan, A.N., Doyle, J.J., et al. -

On Some Hydroids (Cnidaria) from the Coast of Pakistan
Pakistan J. Zool., vol. 38(3), pp. 225-232, 2006. On Some Hydroids (Cnidaria) from the Coast of Pakistan NASEEM MOAZZAM AND MOHAMMAD MOAZZAM Institute of Marine Sciences, University of Karachi, Karachi 75270, Pakistan (NM) and Marine Fisheries Department, Government of Pakistan, Fish Harbour, West Wharf, Karachi 74900, Pakistan (MM) Abstract .- The paper deals with the occurrence of eleven species of the hydroids from the coast of Pakistan. All the species are reported for the first time from Pakistan. These species are Hydractinia epidocleensis, Pennaria disticha, Eudendrium capillare, Orthopyxis cf. crenata, Clytia noliformis, C. hummelincki, Dynamena crisioides, D. quadridentata, Sertularia distans, Pycnotheca mirabilis and Macrorhynchia philippina. Key words: Hydroids, Coelenterata, Pakistan, Hydractinia, Pennaria, Eudendrium, Orthopyxis, Clytia, Dynamena, Sertularia, Pycnotheca, Macrorhynchia. INTRODUCTION used in the paper are derived from Millard (1975), Gibbons and Ryland (1989), Ryland and Gibbons (1991). In comparison to other invertebrates, TAXONOMIC ENUMERATION hydroids are one of the least known groups of marine animals from the coast of Pakistan Haque Family BOUGAINVILLIIDAE (1977) reported a few Cnidaria from the Pakistani Genus HYDRACTINIA Van Beneden, 1841 coast including two hydroids i.e. Plumularia flabellum Allman, 1883 (= P. insignis Allman, 1. Hydractinia epidocleensis Leloup, 1931 1883) and Campanularia juncea Allman, 1874 (= (Fig. 1) Thyroscyphus junceus (Allman, 1876) from Keamari and Bhit Island, Karachi, respectively. Ahmed and Hameed (1999), Ahmed et al. (1978) and Haq et al. (1978) have mentioned the presence of hydroids in various habitats along the coast of Pakistan. Javed and Mustaquim (1995) reported Sertularia turbinata (Lamouroux, 1816) from Manora Channel, Karachi. The present paper describes eleven species of Cnidaria collected from the Pakistani coast all of which are new records for Pakistan. -

Cnidarian Immunity and the Repertoire of Defense Mechanisms in Anthozoans
biology Review Cnidarian Immunity and the Repertoire of Defense Mechanisms in Anthozoans Maria Giovanna Parisi 1,* , Daniela Parrinello 1, Loredana Stabili 2 and Matteo Cammarata 1,* 1 Department of Earth and Marine Sciences, University of Palermo, 90128 Palermo, Italy; [email protected] 2 Department of Biological and Environmental Sciences and Technologies, University of Salento, 73100 Lecce, Italy; [email protected] * Correspondence: [email protected] (M.G.P.); [email protected] (M.C.) Received: 10 August 2020; Accepted: 4 September 2020; Published: 11 September 2020 Abstract: Anthozoa is the most specious class of the phylum Cnidaria that is phylogenetically basal within the Metazoa. It is an interesting group for studying the evolution of mutualisms and immunity, for despite their morphological simplicity, Anthozoans are unexpectedly immunologically complex, with large genomes and gene families similar to those of the Bilateria. Evidence indicates that the Anthozoan innate immune system is not only involved in the disruption of harmful microorganisms, but is also crucial in structuring tissue-associated microbial communities that are essential components of the cnidarian holobiont and useful to the animal’s health for several functions including metabolism, immune defense, development, and behavior. Here, we report on the current state of the art of Anthozoan immunity. Like other invertebrates, Anthozoans possess immune mechanisms based on self/non-self-recognition. Although lacking adaptive immunity, they use a diverse repertoire of immune receptor signaling pathways (PRRs) to recognize a broad array of conserved microorganism-associated molecular patterns (MAMP). The intracellular signaling cascades lead to gene transcription up to endpoints of release of molecules that kill the pathogens, defend the self by maintaining homeostasis, and modulate the wound repair process. -

Carbonic Anhydrases: an Ancient Tool in Calcareous Sponge Biomineralization
fgene-12-624533 March 30, 2021 Time: 13:30 # 1 BRIEF RESEARCH REPORT published: 07 April 2021 doi: 10.3389/fgene.2021.624533 Carbonic Anhydrases: An Ancient Tool in Calcareous Sponge Biomineralization Oliver Voigt1*, Benedetta Fradusco1, Carolin Gut1, Charalampos Kevrekidis1, Sergio Vargas1 and Gert Wörheide1,2,3 1 Department of Earth and Environmental Sciences, Palaeontology and Geobiology, Ludwig-Maximilians-Universität München, Munich, Germany, 2 GeoBio-Center, Ludwig-Maximilians-Universität München, Munich, Germany, 3 SNSB-Bayerische Staatssammlung für Paläontologie und Geologie, Munich, Germany Enzymes of the a-carbonic anhydrase gene family (CAs) are essential for the deposition of calcium carbonate biominerals. In calcareous sponges (phylum Porifera, class Calcarea), specific CAs are involved in the formation of calcite spicules, a unique trait and synapomorphy of this class. However, detailed studies on the CA repertoire of calcareous sponges exist for only two species of one of the two Calcarea subclasses, the Calcaronea. The CA repertoire of the second subclass, the Calcinea, has not been investigated so far, leaving a considerable gap in our knowledge about this Edited by: Melanie Debiais-Thibaud, gene family in Calcarea. Here, using transcriptomic analysis, phylogenetics, and in situ Université de Montpellier, France hybridization, we study the CA repertoire of four additional species of calcareous Reviewed by: sponges, including three from the previously unsampled subclass Calcinea. Our data Helena Cetkovi´ c,´ Rudjer Boskovic Institute, Croatia indicate that the last common ancestor of Calcarea had four ancestral CAs with defined Ana Riesgo, subcellular localizations and functions (mitochondrial/cytosolic, membrane-bound, and Natural History Museum, secreted non-catalytic). The evolution of membrane-bound and secreted CAs involved United Kingdom gene duplications and losses, whereas mitochondrial/cytosolic and non-catalytic CAs *Correspondence: Oliver Voigt are evidently orthologous genes. -
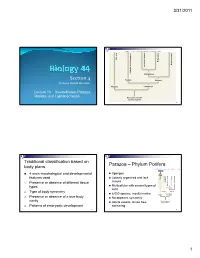
S I Section 4
3/31/2011 Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Porifera Ecdysozoa Deuterostomia Lophotrochozoa Cnidaria and Ctenophora Cnidaria and Protostomia SSiection 4 Radiata Bilateria Professor Donald McFarlane Parazoa Eumetazoa Lecture 13 Invertebrates: Parazoa, Radiata, and Lophotrochozoa Ancestral colonial choanoflagellate 2 Traditional classification based on Parazoa – Phylum Porifera body plans Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Parazoa 4 main morphological and developmental Sponges features used Loosely organized and lack Porifera Ecdysozoa Cnidaria and Ctenophora tissues euterostomia photrochozoa D 1. o Presence or absence of different tissue L Multicellular with several types of types Protostomia cells 2. Type of body symmetry Radiata Bilateria 8,000 species, mostly marine Parazoa Eumetazoa 3. Presence or absence of a true body No apparent symmetry cavity Ancestral colonial Adults sessile, larvae free- choanoflagellate 4. Patterns of embryonic development swimming 3 4 1 3/31/2011 Water drawn through pores (ostia) into spongocoel Flows out through osculum Reproduce Choanocytes line spongocoel Sexually Most hermapppgggphrodites producing eggs and sperm Trap and eat small particles and plankton Gametes are derived from amoebocytes or Mesohyl between choanocytes and choanocytes epithelial cells Asexually Amoebocytes absorb food from choanocytes, Small fragment or bud may detach and form a new digest it, and carry -

Basal Metazoans - Dirk Erpenbeck, Simion Paul, Michael Manuel, Paulyn Cartwright, Oliver Voigt and Gert Worheide
EVOLUTION OF PHYLOGENETIC TREE OF LIFE - Basal Metazoans - Dirk Erpenbeck, Simion Paul, Michael Manuel, Paulyn Cartwright, Oliver Voigt and Gert Worheide BASAL METAZOANS Dirk Erpenbeck Ludwig-Maximilians Universität München, Germany Simion Paul and Michaël Manuel Université Pierre et Marie Curie in Paris, France. Paulyn Cartwright University of Kansas USA. Oliver Voigt and Gert Wörheide Ludwig-Maximilians Universität München, Germany Keywords: Metazoa, Porifera, sponges, Placozoa, Cnidaria, anthozoans, jellyfishes, Ctenophora, comb jellies Contents 1. Introduction on ―Basal Metazoans‖ 2. Phylogenetic relationships among non-bilaterian Metazoa 3. Porifera (Sponges) 4. Placozoa 5. Ctenophora (Comb-jellies) 6. Cnidaria 7. Cultural impact and relevance to human welfare Glossary Bibliography Biographical Sketch Summary Basal metazoans comprise the four non-bilaterian animal phyla Porifera (sponges), Cnidaria (anthozoans and jellyfishes), Placozoa (Trichoplax) and Ctenophora (comb jellies). The phylogenetic position of these taxa in the animal tree is pivotal for our understanding of the last common metazoan ancestor and the character evolution all Metazoa,UNESCO-EOLSS but is much debated. Morphological, evolutionary, internal and external phylogenetic aspects of the four phyla are highlighted and discussed. SAMPLE CHAPTERS 1. Introduction on “Basal Metazoans” In many textbooks the term ―lower metazoans‖ still refers to an undefined assemblage of invertebrate phyla, whose phylogenetic relationships were rather undefined. This assemblage may contain both bilaterian and non-bilaterian taxa. Currently, ―Basal Metazoa‖ refers to non-bilaterian animals only, four phyla that lack obvious bilateral symmetry, Porifera, Placozoa, Cnidaria and Ctenophora. ©Encyclopedia of Life Support Systems (EOLSS) EVOLUTION OF PHYLOGENETIC TREE OF LIFE - Basal Metazoans - Dirk Erpenbeck, Simion Paul, Michael Manuel, Paulyn Cartwright, Oliver Voigt and Gert Worheide These four phyla have classically been known as ―diploblastic‖ Metazoa.