Rumex, Polygonaceae) Reveals Plasticity of Reproductive
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The 2014 Golden Gate National Parks Bioblitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event
National Park Service U.S. Department of the Interior Natural Resource Stewardship and Science The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 ON THIS PAGE Photograph of BioBlitz participants conducting data entry into iNaturalist. Photograph courtesy of the National Park Service. ON THE COVER Photograph of BioBlitz participants collecting aquatic species data in the Presidio of San Francisco. Photograph courtesy of National Park Service. The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 Elizabeth Edson1, Michelle O’Herron1, Alison Forrestel2, Daniel George3 1Golden Gate Parks Conservancy Building 201 Fort Mason San Francisco, CA 94129 2National Park Service. Golden Gate National Recreation Area Fort Cronkhite, Bldg. 1061 Sausalito, CA 94965 3National Park Service. San Francisco Bay Area Network Inventory & Monitoring Program Manager Fort Cronkhite, Bldg. 1063 Sausalito, CA 94965 March 2016 U.S. Department of the Interior National Park Service Natural Resource Stewardship and Science Fort Collins, Colorado The National Park Service, Natural Resource Stewardship and Science office in Fort Collins, Colorado, publishes a range of reports that address natural resource topics. These reports are of interest and applicability to a broad audience in the National Park Service and others in natural resource management, including scientists, conservation and environmental constituencies, and the public. The Natural Resource Report Series is used to disseminate comprehensive information and analysis about natural resources and related topics concerning lands managed by the National Park Service. -

Atraphaxis Radkanensis (Polygonaceae), a New Species from Iran
Ann. Bot. Fennici 50: 372–374 ISSN 0003-3847 (print) ISSN 1797-2442 (online) Helsinki 10 October 2013 © Finnish Zoological and Botanical Publishing Board 2013 Atraphaxis radkanensis (Polygonaceae), a new species from Iran Solmaz Tavakkoli1, Shahrokh Kazempour Osaloo1,*, Valiollah Mozaffarian2 & Ali Asghar Maassoumi2 1) Department of Plant Biology, Faculty of Biological Sciences, Tarbiat Modares University, Tehran 14115-154, Iran (*corresponding author’ e-mails: [email protected], skosaloo@gmail. com) 2) Department of Botany, Research Institute of Forests and Rangelands, Tehran 13185-116, Iran Received 8 Apr. 2013, final version received 17 June 2013, accepted 20 June 2013 Tavakkoli, S., Kazempour Osaloo, S., Mozaffarian, V. & Maassoumi, A. A. 2013: Atraphaxis rad- kanensis (Polygonaceae), a new species from Iran. — Ann. Bot. Fennici 50: 372–374. Atraphaxis radkanensis Tavakkoli, Kaz. Osaloo & Mozaff. (Polygonaceae) is described and illustrated as a new species from NE Iran. Atraphaxis radkanensis is very similar to A. seravschanica, and both of them are placed in the section Tragopyrum. Atrap- haxis radkanensis is characterized by oblong-ovate leaves, a puberulent indumentum on both surfaces of the leaves and twigs, a mostly terminal inflorescence, as well as by small-sized achenes. Atraphaxis belongs to the tribe Polygoneae of of the floral characters such as the number of Polygonaceae subfam. Polygonoideae (Sanchez perianth segments, stamens and style as well as et al. 2011). The genus comprises 30 species achene shape, Atraphaxis has been divided into distributed in northern Africa and Eurasia, with two subgenera: Atraphaxis (as Euatraphaxis) the greatest diversity in Central Asia (Pavlov and Tragopyrum (Pavlov 1936) or two sec- 1936, Cullen 1967, Rechinger & Schiman- tions: Atraphaxis and Tragopyrum (Rechinger Czeika 1968, Brandbyge 1993, Qaiser 2001, & Schiman-Czeika 1968). -

"National List of Vascular Plant Species That Occur in Wetlands: 1996 National Summary."
Intro 1996 National List of Vascular Plant Species That Occur in Wetlands The Fish and Wildlife Service has prepared a National List of Vascular Plant Species That Occur in Wetlands: 1996 National Summary (1996 National List). The 1996 National List is a draft revision of the National List of Plant Species That Occur in Wetlands: 1988 National Summary (Reed 1988) (1988 National List). The 1996 National List is provided to encourage additional public review and comments on the draft regional wetland indicator assignments. The 1996 National List reflects a significant amount of new information that has become available since 1988 on the wetland affinity of vascular plants. This new information has resulted from the extensive use of the 1988 National List in the field by individuals involved in wetland and other resource inventories, wetland identification and delineation, and wetland research. Interim Regional Interagency Review Panel (Regional Panel) changes in indicator status as well as additions and deletions to the 1988 National List were documented in Regional supplements. The National List was originally developed as an appendix to the Classification of Wetlands and Deepwater Habitats of the United States (Cowardin et al.1979) to aid in the consistent application of this classification system for wetlands in the field.. The 1996 National List also was developed to aid in determining the presence of hydrophytic vegetation in the Clean Water Act Section 404 wetland regulatory program and in the implementation of the swampbuster provisions of the Food Security Act. While not required by law or regulation, the Fish and Wildlife Service is making the 1996 National List available for review and comment. -

Plants-Derived Biomolecules As Potent Antiviral Phytomedicines: New Insights on Ethnobotanical Evidences Against Coronaviruses
plants Review Plants-Derived Biomolecules as Potent Antiviral Phytomedicines: New Insights on Ethnobotanical Evidences against Coronaviruses Arif Jamal Siddiqui 1,* , Corina Danciu 2,*, Syed Amir Ashraf 3 , Afrasim Moin 4 , Ritu Singh 5 , Mousa Alreshidi 1, Mitesh Patel 6 , Sadaf Jahan 7 , Sanjeev Kumar 8, Mulfi I. M. Alkhinjar 9, Riadh Badraoui 1,10,11 , Mejdi Snoussi 1,12 and Mohd Adnan 1 1 Department of Biology, College of Science, University of Hail, Hail PO Box 2440, Saudi Arabia; [email protected] (M.A.); [email protected] (R.B.); [email protected] (M.S.); [email protected] (M.A.) 2 Department of Pharmacognosy, Faculty of Pharmacy, “Victor Babes” University of Medicine and Pharmacy, 2 Eftimie Murgu Square, 300041 Timisoara, Romania 3 Department of Clinical Nutrition, College of Applied Medical Sciences, University of Hail, Hail PO Box 2440, Saudi Arabia; [email protected] 4 Department of Pharmaceutics, College of Pharmacy, University of Hail, Hail PO Box 2440, Saudi Arabia; [email protected] 5 Department of Environmental Sciences, School of Earth Sciences, Central University of Rajasthan, Ajmer, Rajasthan 305817, India; [email protected] 6 Bapalal Vaidya Botanical Research Centre, Department of Biosciences, Veer Narmad South Gujarat University, Surat, Gujarat 395007, India; [email protected] 7 Department of Medical Laboratory, College of Applied Medical Sciences, Majmaah University, Al Majma’ah 15341, Saudi Arabia; [email protected] 8 Department of Environmental Sciences, Central University of Jharkhand, -

Chinese Rhubarb)
IJAS_39192 Vol 8, Issue 6, 2020 ISSN- 2321-6832 Review Article GENERAL OVERVIEW OF PHYTOCHEMISTRY AND PHARMACOLOGICAL POTENTIAL OF RHEUM PALMATUM (CHINESE RHUBARB) AAMIR KHAN KHATTAK, SYEDA MONA HASSAN, SHAHZAD SHARIF MUGHAL* Department of Chemistry, Lahore Garrison University, Lahore, Pakistan, Email: [email protected] Received: 21 July 2020 , Revised and Accepted: 11 October 2020 ABSTRACT Recent probe of medicinal plants incorporated in traditional systems for curing infection and sustaining holistic health, has exposed good sum of therapeutic efficiency against deleterious infections and chronic illnesses. Rheum palmatum (Chinese Rhubarb, family Polygonaceae) is a significant medicinal herb, which finds an extensive use in Unani (Traditional) system of medicine. It has been traditionally employed as antiseptic, liver stimulant, diuretic, diabetes, stomachic, purgative/cathartic, anticholesterolemic, antitumor, Alzheimer’s, Parkinson’s, tonic, antidiabetic, and wound healer. The most vital components from Rheum palmatum are the phenolics, flavonoids, terpenoids, saponins, and anthraquinone derivatives such as aloe- emodin, chrysophanol, physcion, rhein, emodin and its glucorhein, and glycoside. Rhubarb also contains tannins which include hydrolysable-tannins, containing glycosidic or ester bonds composed of glucose, gallic acid, and other monosaccharide’s and condensed tannins, resulting principally from the flavone derivatives leukocyanidin and catechin. In recent years, new components such asrevandchinone-1, revandchinone-2, revandchinone-3, revandchinone-4, sulfemodin8-O-b-Dglucoside, and 6-methyl-rhein and aloe-emodin have been reported from the same class. It also encompasses some macro and micro mineral elements such as Ca, K, Mn, Fe, Co, Zn, Na, Cu, and Li. Anthraquinone derivatives demonstrate evidence of anti- microbial, antifungal, anti-proliferative, anti-Parkinson’s, immune enhancing, anticancer, antiulcer, antioxidant, and antiviral activities. -
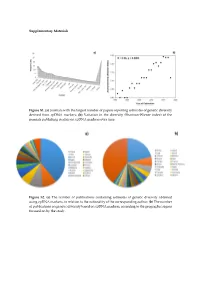
(A) Journals with the Largest Number of Papers Reporting Estimates Of
Supplementary Materials Figure S1. (a) Journals with the largest number of papers reporting estimates of genetic diversity derived from cpDNA markers; (b) Variation in the diversity (Shannon-Wiener index) of the journals publishing studies on cpDNA markers over time. Figure S2. (a) The number of publications containing estimates of genetic diversity obtained using cpDNA markers, in relation to the nationality of the corresponding author; (b) The number of publications on genetic diversity based on cpDNA markers, according to the geographic region focused on by the study. Figure S3. Classification of the angiosperm species investigated in the papers that analyzed genetic diversity using cpDNA markers: (a) Life mode; (b) Habitat specialization; (c) Geographic distribution; (d) Reproductive cycle; (e) Type of flower, and (f) Type of pollinator. Table S1. Plant species identified in the publications containing estimates of genetic diversity obtained from the use of cpDNA sequences as molecular markers. Group Family Species Algae Gigartinaceae Mazzaella laminarioides Angiospermae Typhaceae Typha laxmannii Angiospermae Typhaceae Typha orientalis Angiospermae Typhaceae Typha angustifolia Angiospermae Typhaceae Typha latifolia Angiospermae Araliaceae Eleutherococcus sessiliflowerus Angiospermae Polygonaceae Atraphaxis bracteata Angiospermae Plumbaginaceae Armeria pungens Angiospermae Aristolochiaceae Aristolochia kaempferi Angiospermae Polygonaceae Atraphaxis compacta Angiospermae Apocynaceae Lagochilus macrodontus Angiospermae Polygonaceae Atraphaxis -

Factors Involved in the Maintenance of the Grassy Balds of the Great Smoky Mountains National Park
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Masters Theses Graduate School 3-1968 Factors Involved in the Maintenance of the Grassy Balds of the Great Smoky Mountains National Park Stephen Walker Radford University of Tennessee - Knoxville Follow this and additional works at: https://trace.tennessee.edu/utk_gradthes Part of the Plant Sciences Commons Recommended Citation Radford, Stephen Walker, "Factors Involved in the Maintenance of the Grassy Balds of the Great Smoky Mountains National Park. " Master's Thesis, University of Tennessee, 1968. https://trace.tennessee.edu/utk_gradthes/1446 This Thesis is brought to you for free and open access by the Graduate School at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Masters Theses by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. To the Graduate Council: I am submitting herewith a thesis written by Stephen Walker Radford entitled "Factors Involved in the Maintenance of the Grassy Balds of the Great Smoky Mountains National Park." I have examined the final electronic copy of this thesis for form and content and recommend that it be accepted in partial fulfillment of the equirr ements for the degree of Master of Science, with a major in Botany. Edward E. C. Clebsch, Major Professor We have read this thesis and recommend its acceptance: Ronald H. Peterson, Edward R. Buckner Accepted for the Council: Carolyn R. Hodges Vice Provost and Dean of the Graduate School (Original signatures are on file with official studentecor r ds.) February 28, 1968 To the Graduate Council: I am submitting herewith a thesis written by Stephen Walker Radford entitled "Factors Involved in the Maintenance of the Grassy Balds of the Great Smoky Mountains National Park." I recommend that it be accepted for nine quarter hours of credit in partial fulfillment o�the requirements for the degree of Master of Science, with a major in Botany. -

Draft Plant Propagation Protocol
Plant Propagation Protocol for Oxyria digyna ESRM 412 – Native Plant Production Protocol URL: https://courses.washington.edu/esrm412/protocols/OXDI3.pdf Images from google.com TAXONOMY Plant Family Scientific Name Polygonaceae Common Name Buckwheat Species Scientific Name Scientific Name Oxyria digyna Varieties Sub-species Cultivar Common Rheum digynum Synonym(s) Common Name(s) Wood Sorrel, Alpine Sorrel, Alpine Mountainsorrel, Mountain-sorrel, Oxyrie de Montagne Species Code (as OXDI3 per USDA Plants database) GENERAL INFORMATION Geographical range See above map12 Ecological Grows in a wide range of areas, particularly in rocky areas and scree distribution slopes, in alpine and subalpine areas.5,6,10 Does well in disturbed sites11 with soils not too acidic, and moist but well drained. Can grow successfully in open areas with little to no shade, and often grows beneath bird nests due to the high nitrogen content of the soil.4 Climate and elevation range Local habitat and Subalpine forest or wetland riparian species2 abundance Plant strategy type / Produces small, wind dispersed seeds, making it a weed-like pioneer successional stage species, especially in disturbed sites.3 Tolerant of sun stress, but not of interspecies competition, particularly that of Ranunculus glacialis. Is able to germinate after being buried beneath snow.8 Has an intermediate tolerance for shade, a low tolerance for drought and anoxic conditions, and is adapted to coarse to medium soils. Seedlings spread slowly and are moderately vigorous, and the plant does not usually propagate vegetatively.12 Plant characteristics Oxyria digyna is a perennial herb which grows to approx. 2ft tall and 1ft wide.4 It is hairless, having a branching crown and several stems11 with swollen nodes8, and it is reddish tinged11. -

Widespread Paleopolyploidy, Gene Tree Conflict, and Recalcitrant Relationships Among the 3 Carnivorous Caryophyllales1 4 5 Joseph F
bioRxiv preprint doi: https://doi.org/10.1101/115741; this version posted March 10, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 2 Widespread paleopolyploidy, gene tree conflict, and recalcitrant relationships among the 3 carnivorous Caryophyllales1 4 5 Joseph F. Walker*,2, Ya Yang2,5, Michael J. Moore3, Jessica Mikenas3, Alfonso Timoneda4, Samuel F. 6 Brockington4 and Stephen A. Smith*,2 7 8 2Department of Ecology & Evolutionary Biology, University of Michigan, 830 North University Avenue, 9 Ann Arbor, MI 48109-1048, USA 10 3Department of Biology, Oberlin College, Science Center K111, 119 Woodland St., Oberlin, Ohio 44074- 11 1097 USA 12 4Department of Plant Sciences, University of Cambridge, Cambridge CB2 3EA, United Kingdom 13 5 Department of Plant Biology, University of Minnesota-Twin Cities. 1445 Gortner Avenue, St. Paul, MN 14 55108 15 CORRESPONDING AUTHORS: Joseph F. Walker; [email protected] and Stephen A. Smith; 16 [email protected] 17 18 1Manuscript received ____; revision accepted ______. bioRxiv preprint doi: https://doi.org/10.1101/115741; this version posted March 10, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 19 ABSTRACT 20 • The carnivorous members of the large, hyperdiverse Caryophyllales (e.g. -

CURLY DOCK SHEEP SORREL Rumex Crisp Us Rumex Acetosella
CURLY DOCK SHEEP SORREL Rumex crisp us Rumex acetosella FAMILY: Polygonaceae GENUS: Rumex SPECIES: crispus acetosella COMMON NAMES: Curly Dock Sheep Sorrel Yellow Dock Sorrell Dock Sour Dock Field Sorrel Narrow-leaved Dock Red Sorrel Wood Sorrel Sour-weed POLLEN GRAINS: Spheroidal, 23 to 27 microns in diameter. A thin exine is finely pitted. Three or four (sometimes six) linear furrows are evenly arranged, each having a small elliptical germ pore. POLLINATING PERIOD: June to August. Earlier in warmer Throughout the summer but areas. heaviest in May and June. DISTRIBUTION: Throughout the United States. ThroughoutmostoftheUnited States. Rarely found in Southern California or desert areas of the Southwest. ALLERGIC IMPORTANCE: Of only moderate Importance In areas of abundance. Curly Dock is a stout perennial 1 ~ to 5 feet Sheep Sorrel is a perennial ~ to 2 feet tall tall. During the first year the plant forms a arising from a slender running rootstock. The dense roseaffe of leaves. Leaves are elon leaves are rather fleshy and vary in size and gated with wavy margins. The upright stems shape from lower to upper. The lower leaves are leafy. The terminal inflorescence is dense are halbard-shaped with two basal lobes. with few or no leaves and is 1 to 2 feet long. In The inflorescence is terminal and mostly leaf fruit in the late summer and fall the tops turn a less. Individual flowers are of separate sex characteristic reddish brown. with the female being red and the male a yellowish green color. The foliage of the docks is acid but especially so in this species. -

Somerset's Ecological Network
Somerset’s Ecological Network Mapping the components of the ecological network in Somerset 2015 Report This report was produced by Michele Bowe, Eleanor Higginson, Jake Chant and Michelle Osbourn of Somerset Wildlife Trust, and Larry Burrows of Somerset County Council, with the support of Dr Kevin Watts of Forest Research. The BEETLE least-cost network model used to produce Somerset’s Ecological Network was developed by Forest Research (Watts et al, 2010). GIS data and mapping was produced with the support of Somerset Environmental Records Centre and First Ecology Somerset Wildlife Trust 34 Wellington Road Taunton TA1 5AW 01823 652 400 Email: [email protected] somersetwildlife.org Front Cover: Broadleaved woodland ecological network in East Mendip Contents 1. Introduction .................................................................................................................... 1 2. Policy and Legislative Background to Ecological Networks ............................................ 3 Introduction ............................................................................................................... 3 Government White Paper on the Natural Environment .............................................. 3 National Planning Policy Framework ......................................................................... 3 The Habitats and Birds Directives ............................................................................. 4 The Conservation of Habitats and Species Regulations 2010 .................................. -

Rhubarb 2018
ONLINE SERIES MONOGRAPHS The Scientific Foundation for Herbal Medicinal Products Rhei radix Rhubarb 2018 www.escop.com The Scientific Foundation for Herbal Medicinal Products RHEI RADIX Rhubarb 2018 ESCOP Monographs were first published in loose-leaf form progressively from 1996 to 1999 as Fascicules 1-6, each of 10 monographs © ESCOP 1996, 1997, 1999 Second Edition, completely revised and expanded © ESCOP 2003 Second Edition, Supplement 2009 © ESCOP 2009 ONLINE SERIES ISBN 978-1-901964-59-2 Rhei radix - Rhubarb © ESCOP 2018 Published by the European Scientific Cooperative on Phytotherapy (ESCOP) Notaries House, Chapel Street, Exeter EX1 1EZ, United Kingdom www.escop.com All rights reserved Except for the purposes of private study, research, criticism or review no part of this text may be reproduced, stored in a retrieval system or transmitted, in any form or by any means, without the written permission of the publisher. Important Note: Medical knowledge is ever-changing. As new research and clinical experience broaden our knowledge, changes in treatment may be required. In their efforts to provide information on the efficacy and safety of herbal drugs and herbal preparations, presented as a substantial overview together with summaries of relevant data, the authors of the material herein have consulted comprehensive sources believed to be reliable. However, in view of the possibility of human error by the authors or publisher of the work herein, or changes in medical knowledge, neither the authors nor the publisher, nor any other party involved in the preparation of this work, warrants that the information contained herein is in every respect accurate or complete, and they are not responsible for any errors or omissions or for results obtained by the use of such information.