Package 'Move'
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
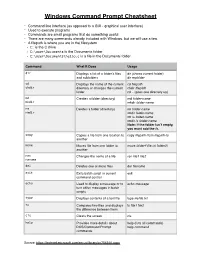
Windows Command Prompt Cheatsheet
Windows Command Prompt Cheatsheet - Command line interface (as opposed to a GUI - graphical user interface) - Used to execute programs - Commands are small programs that do something useful - There are many commands already included with Windows, but we will use a few. - A filepath is where you are in the filesystem • C: is the C drive • C:\user\Documents is the Documents folder • C:\user\Documents\hello.c is a file in the Documents folder Command What it Does Usage dir Displays a list of a folder’s files dir (shows current folder) and subfolders dir myfolder cd Displays the name of the current cd filepath chdir directory or changes the current chdir filepath folder. cd .. (goes one directory up) md Creates a folder (directory) md folder-name mkdir mkdir folder-name rm Deletes a folder (directory) rm folder-name rmdir rmdir folder-name rm /s folder-name rmdir /s folder-name Note: if the folder isn’t empty, you must add the /s. copy Copies a file from one location to copy filepath-from filepath-to another move Moves file from one folder to move folder1\file.txt folder2\ another ren Changes the name of a file ren file1 file2 rename del Deletes one or more files del filename exit Exits batch script or current exit command control echo Used to display a message or to echo message turn off/on messages in batch scripts type Displays contents of a text file type myfile.txt fc Compares two files and displays fc file1 file2 the difference between them cls Clears the screen cls help Provides more details about help (lists all commands) DOS/Command Prompt help command commands Source: https://technet.microsoft.com/en-us/library/cc754340.aspx. -

Mac Keyboard Shortcuts Cut, Copy, Paste, and Other Common Shortcuts
Mac keyboard shortcuts By pressing a combination of keys, you can do things that normally need a mouse, trackpad, or other input device. To use a keyboard shortcut, hold down one or more modifier keys while pressing the last key of the shortcut. For example, to use the shortcut Command-C (copy), hold down Command, press C, then release both keys. Mac menus and keyboards often use symbols for certain keys, including the modifier keys: Command ⌘ Option ⌥ Caps Lock ⇪ Shift ⇧ Control ⌃ Fn If you're using a keyboard made for Windows PCs, use the Alt key instead of Option, and the Windows logo key instead of Command. Some Mac keyboards and shortcuts use special keys in the top row, which include icons for volume, display brightness, and other functions. Press the icon key to perform that function, or combine it with the Fn key to use it as an F1, F2, F3, or other standard function key. To learn more shortcuts, check the menus of the app you're using. Every app can have its own shortcuts, and shortcuts that work in one app may not work in another. Cut, copy, paste, and other common shortcuts Shortcut Description Command-X Cut: Remove the selected item and copy it to the Clipboard. Command-C Copy the selected item to the Clipboard. This also works for files in the Finder. Command-V Paste the contents of the Clipboard into the current document or app. This also works for files in the Finder. Command-Z Undo the previous command. You can then press Command-Shift-Z to Redo, reversing the undo command. -
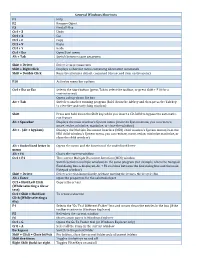
General Windows Shortcuts
General Windows Shortcuts F1 Help F2 Rename Object F3 Find all files Ctrl + Z Undo Ctrl + X Cut Ctrl + C Copy Ctrl + V Paste Ctrl + Y Redo Ctrl + Esc Open Start menu Alt + Tab Switch between open programs Alt + F4 Quit program Shift + Delete Delete item permanently Shift + Right Click Displays a shortcut menu containing alternative commands Shift + Double Click Runs the alternate default command ( the second item on the menu) Alt + Double Click Displays properties F10 Activates menu bar options Shift + F10 Opens a contex t menu ( same as righ t click) Ctrl + Esc or Esc Selects the Start button (press Tab to select the taskbar, or press Shift + F10 for a context menu) Alt + Down Arrow Opens a drop‐down list box Alt + Tab Switch to another running program (hold down the Alt key and then press the Tab key to view the task‐switching window) Alt + Shift + Tab Swit ch b ackward s b etween open appli cati ons Shift Press and hold down the Shift key while you insert a CD‐ROM to bypass the automatic‐ run feature Alt + Spacebar Displays the main window's System menu (from the System menu, you can restore, move, resize, minimize, maximize, or close the window) Alt + (Alt + hyphen) Displays the Multiple Document Interface (MDI) child window's System menu (from the MDI child window's System menu, you can restore, move, resize, minimize maximize, or close the child window) Ctrl + Tab Switch to t h e next child window o f a Multi ple D ocument Interf ace (MDI) pr ogram Alt + Underlined letter in Opens the menu and the function of the underlined letter -

Powerview Command Reference
PowerView Command Reference TRACE32 Online Help TRACE32 Directory TRACE32 Index TRACE32 Documents ...................................................................................................................... PowerView User Interface ............................................................................................................ PowerView Command Reference .............................................................................................1 History ...................................................................................................................................... 12 ABORT ...................................................................................................................................... 13 ABORT Abort driver program 13 AREA ........................................................................................................................................ 14 AREA Message windows 14 AREA.CLEAR Clear area 15 AREA.CLOSE Close output file 15 AREA.Create Create or modify message area 16 AREA.Delete Delete message area 17 AREA.List Display a detailed list off all message areas 18 AREA.OPEN Open output file 20 AREA.PIPE Redirect area to stdout 21 AREA.RESet Reset areas 21 AREA.SAVE Save AREA window contents to file 21 AREA.Select Select area 22 AREA.STDERR Redirect area to stderr 23 AREA.STDOUT Redirect area to stdout 23 AREA.view Display message area in AREA window 24 AutoSTOre .............................................................................................................................. -

S.Ha.R.K. Installation Howto Tools Knoppix Live CD Linux Fdisk HD
S.Ha.R.K. Installation Tools HowTo • Linux fdisk utility • A copy of Linux installation CD • A copy of Windows® installation CD Tullio Facchinetti University of Pavia - Italy • Some FreeDOS utilities • A copy of S.Ha.R.K. S.Ha.R.K. Workshop S.Ha.R.K. Workshop Knoppix live CD Linux fdisk Command action a toggle a bootable flag Download ISO from b edit bsd disklabel c toggle the dos compatibility flag d delete a partition http://www.knoppix.org l list known partition types m print this menu n add a new partition o create a new empty DOS partition table p print the partition table q quit without saving changes • boot from CD s create a new empty Sun disklabel t change a partition's system id • open a command shell u change display/entry units v verify the partition table • type “su” (become root ), password is empty w write table to disk and exit x extra functionality (experts only) • start fdisk (ex. fdisk /dev/hda ) Command (m for help): S.Ha.R.K. Workshop S.Ha.R.K. Workshop HD partitioning HD partitioning 1st FreeDOS FAT32 FreeDOS must be installed Primary 2nd Windows® FAT32 into the first partition of your HD or it may not boot 3rd Linux / extX Data 1 FAT32 format data partitions as ... Extended FAT32, so that you can share Data n FAT32 your data between Linux, last Linux swap swap Windows® and FreeDOS S.Ha.R.K. Workshop S.Ha.R.K. Workshop 1 HD partitioning Windows ® installation FAT32 Windows® partition type Install Windows®.. -

KEYBOARD SHORTCUTS (Windows)
KEYBOARD SHORTCUTS (Windows) Note: For Mac users, please substitute the Command key for the Ctrl key. This substitution with work for the majority of commands _______________________________________________________________________ General Commands Navigation Windows key + D Desktop to foreground Context menu Right click Alt + underlined letter Menu drop down, Action selection Alt + Tab Toggle between open applications Alt, F + X or Alt + F4 Exit application Alt, Spacebar + X Maximize window Alt, Spacebar + N Minimize window Ctrl + W Closes window F2 Renames a selected file or folder Open Programs To open programs from START menu: Create a program shortcut and drop it into START menu To open programs/files on Desktop: Select first letter, and then press Enter to open Dialog Boxes Enter Selects highlighted button Tab Selects next button Arrow keys Selects next (>) or previous button (<) Shift + Tab Selects previous button _______________________________________________________________________ Microsoft Word Formatting Ctrl + P Print Ctrl + S Save Ctrl + Z Undo Ctrl + Y Redo CTRL+B Make text bold CTRL+I Italicize CTRL+U Underline Ctrl + C Copy Ctrl + V Paste Ctrl + X Copy + delete Shift + F3 Change case of letters Ctrl+Shift+> Increase font size Ctrl+Shift+< Decrease font size Highlight Text Shift + Arrow Keys Selects one letter at a time Shift + Ctrl + Arrow keys Selects one word at a time Shift + End or Home Selects lines of text Change or resize the font CTRL+SHIFT+ > Increase the font size 1 KEYBOARD SHORTCUTS (Windows) CTRL+SHIFT+ < -

Disk Management
IBM i Version 7.2 Systems management Disk management IBM Note Before using this information and the product it supports, read the information in “Notices” on page 141. This edition applies to IBM i 7.2 (product number 5770-SS1) and to all subsequent releases and modifications until otherwise indicated in new editions. This version does not run on all reduced instruction set computer (RISC) models nor does it run on CISC models. This document may contain references to Licensed Internal Code. Licensed Internal Code is Machine Code and is licensed to you under the terms of the IBM License Agreement for Machine Code. © Copyright International Business Machines Corporation 2004, 2013. US Government Users Restricted Rights – Use, duplication or disclosure restricted by GSA ADP Schedule Contract with IBM Corp. Contents Disk management..................................................................................................1 What's new for IBM i 7.2..............................................................................................................................1 PDF file for Disk Management..................................................................................................................... 1 Getting started with disk management.......................................................................................................2 Components of disk storage.................................................................................................................. 2 Finding the logical address for a disk -

Partition.Pdf
Linux Partition HOWTO Anthony Lissot Revision History Revision 3.5 26 Dec 2005 reorganized document page ordering. added page on setting up swap space. added page of partition labels. updated max swap size values in section 4. added instructions on making ext2/3 file systems. broken links identified by Richard Calmbach are fixed. created an XML version. Revision 3.4.4 08 March 2004 synchronized SGML version with HTML version. Updated lilo placement and swap size discussion. Revision 3.3 04 April 2003 synchronized SGML and HTML versions Revision 3.3 10 July 2001 Corrected Section 6, calculation of cylinder numbers Revision 3.2 1 September 2000 Dan Scott provides sgml conversion 2 Oct. 2000. Rewrote Introduction. Rewrote discussion on device names in Logical Devices. Reorganized Partition Types. Edited Partition Requirements. Added Recovering a deleted partition table. Revision 3.1 12 June 2000 Corrected swap size limitation in Partition Requirements, updated various links in Introduction, added submitted example in How to Partition with fdisk, added file system discussion in Partition Requirements. Revision 3.0 1 May 2000 First revision by Anthony Lissot based on Linux Partition HOWTO by Kristian Koehntopp. Revision 2.4 3 November 1997 Last revision by Kristian Koehntopp. This Linux Mini−HOWTO teaches you how to plan and create partitions on IDE and SCSI hard drives. It discusses partitioning terminology and considers size and location issues. Use of the fdisk partitioning utility for creating and recovering of partition tables is covered. The most recent version of this document is here. The Turkish translation is here. Linux Partition HOWTO Table of Contents 1. -

Introduction to Command Line
Introduction To The Command Line Getting started with Terminal The Command Line is a text-based interface that allows you to communicate with your computer to accomplish a wide range of tasks. You will use the skills you develop in this lesson when working with Twarc, a command line tool for collecting and analyzing Twitter data. It is important to have a handle on basic commands so that you can work with the 1 October Twitter Data Collection later on in this tutorial series. Get started by downloading the materials below! Difficulty level: Beginner Optimized for: Mac users. Windows users can view the tutorial here. Prerequisite(s): None Materials: Download ‘walt_whitman.zip’ to your desktop Tutorial Key ● Command Line commands will be displayed in this format ● means you have come to the end of a set of instructions Lesson objectives - Use the command line to navigate your computer - Create and move content around - Make changes to existing files Key Terms ● Terminal - OS X Command Line ○ A text interface for your computer. Terminal receives commands and then passes those commands on to the computer's operating system to run. ● Command ○ A specific order from a user to the computer’s operating system to perform a service ● Graphical-User Interface (GUI) ○ A visual form of user interface that allows users to interact with a computer through icons and other visual indicators ● Filepath ○ A unique address that specifies a location in a file system ● Directory ○ A location for storing files on your computer. A directory is the same thing as a folder; a folder is represented visually in a GUI. -

Introduction: DOS (Disk Operating System) Is an Oldest Type of Operating System
DOS (Disk Operating System) 1 Introduction: DOS (Disk Operating System) is an oldest type of Operating System. Disk Operating System is abbreviated as DOS. DOS is a CUI type of Operating System. In computer science, a generic term describing any operating system is system software which is loaded from disk devices when the system is started or rebooted. DOS is a single-tasking, single-user operating system with a command-line interface. DOS acts on commands. Because DOS is ready to perform when given proper command hence, it is also known as Command Prompt. Commands are certain words of English language or short form of English words. The meaning of these word or short form is already known to DOS. Since, DOS recognized these words and hence acts accordingly. These words and short forms of the English words are better known as commands. Internal Command:-Those commands which are already stored in the “Command.Com” file of DOS are known as internal commands. For example, CLS, VOL, TIME, DATE, COPY etc External Command:-Those commands which are not included in the command.com file of DOS rather included in other files of DOS are known as external commands. It is formatted according to programme. For example, TREE, FORMAT, MODE etc Some Internal Commands:- 1. CLS To clear the screen. \>cls 2. DIR To view the directory and files C:\>Dir 3. DATE To View and change the date C:\>Date Current date is: 01-01-2008 Enter new date (mm/dd/yy):21-03-2009 4. TIME To view and change the time. -

Microsoft® Official Academic Course: Networking Fundamentals, Exam
Microsoft® Official Academic Course Networking Fundamentals, Exam 98-366 VP & PUBLISHER Barry Pruett SENIOR EXECUTIVE EDITOR Jim Minatel MICROSOFT PRODUCT MANAGER Microsoft Learning SENIOR EDITORIAL ASSISTANT Devon Lewis TECHNICAL EDITOR Ron Handlon CHANNEL MARKETING MANAGER Michele Szczesniak CONTENT MANAGEMENT DIRECTOR Lisa Wojcik CONTENT MANAGER Nichole Urban PRODUCTION COORDINATOR Nicole Repasky PRODUCTION EDITOR Umamaheswari Gnanamani COVER DESIGNER Tom Nery COVER PHOTO: © shutterstock/wavebreakmedia Copyright © 2017 by John Wiley & Sons, Inc. All rights reserved. No part of this publication may be reproduced, stored in a retrieval system or transmitted in any form or by any means, electronic, mechanical, photocopying, recording, scanning or otherwise, except as permitted under Sections 107 or 108 of the 1976 United States Copyright Act, without either the prior written permission of the Publisher, or authorization through payment of the appropriate per-copy fee to the Copyright Clearance Center, Inc. 222 Rosewood Drive, Danvers, MA 01923, (978) 750-8400, fax (978) 646-8600. Requests to the Publisher for permission should be addressed to the Permissions Department, John Wiley & Sons, Inc., 111 River Street, Hoboken, NJ 07030-5774, (201) 748-6011, fax (201) 748-6008. To order books or for customer service, please call 1-800-CALL WILEY (225-5945). Microsoft, Active Directory, AppLocker, Bing, BitLocker, Hyper-V, Internet Explorer, Microsoft Intune, Microsoft Office 365, SQL Server, Visual Studio, Windows Azure, Windows, Windows PowerShell, and Windows Server are either registered trademarks or trademarks of Microsoft Corporation in the United States and/or other countries. Other product and company names mentioned herein may be the trademarks of their respective owners. The example companies, organizations, products, domain names, e-mail addresses, logos, people, places, and events depicted herein are fictitious. -

How to Recover Your VMFS Partition
How to recover your VMFS partition November 8,2007 Signs of a bad day. All or a portion of your VM’s are missing One or more of your VMFS volumes returns: [root@tekitd01 vmhba2]# fdisk -lu /dev/sda Disk /dev/sda: 322.1 GB, 322122547200 bytes 255 heads, 63 sectors/track, 39162 cylinders, total 629145600 sectors Units = sectors of 1 * 512 = 512 bytes Disk /dev/sda doesn't contain a valid partition table . You can list your SAN devices with the following command: [root@tekitd01 vmhba2]# esxcfg-vmhbadevs vmhba0:0:0 /dev/cciss/c0d0 vmhba1:0:1 /dev/sda vmhba1:0:2 /dev/sdb vmhba1:4:2 /dev/sdc . Normal output from fdisk: [root@tekitd01 vmhba2]# fdisk -lu /dev/sdb Disk /dev/sdb: 16.1 GB, 16106127360 bytes 255 heads, 63 sectors/track, 1958 cylinders, total 31457280 sectors Units = sectors of 1 * 512 = 512 bytes Device Boot Start End Blocks Id System /dev/sdb1 128 31455269 15727571 fb Unknown 2 11/08/2007 How to Recover Your VMFS Partition Recovery Steps [root@tekitd01 vmhba2]# fdisk /dev/sda Device contains neither a valid DOS partition table, nor Sun, SGI or OSF disklabel Building a new DOS disklabel. Changes will remain in memory only, until you decide to write them. After that, of course, the previous content won't be recoverable. The number of cylinders for this disk is set to 39162. There is nothing wrong with that, but this is larger than 1024, and could in certain setups cause problems with: 1) software that runs at boot time (e.g., old versions of LILO) 2) booting and partitioning software from other OSs (e.g., DOS FDISK, OS/2 FDISK) Warning: