And Regional-Scale Estimations of the Diversity of Stream Fish Using Edna
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Digeneans (Trematoda) Parasitic in Freshwater Fishes (Osteichthyes) of the Lake Biwa Basin in Shiga Prefecture, Central Honshu, Japan
Digeneans (Trematoda) Parasitic in Freshwater Fishes (Osteichthyes) of the Lake Biwa Basin in Shiga Prefecture, Central Honshu, Japan Takeshi Shimazu1, Misako Urabe2 and Mark J. Grygier3 1 Nagano Prefectural College, 8–49–7 Miwa, Nagano City, Nagano 380–8525, Japan and 10486–2 Hotaka-Ariake, Azumino City, Nagano 399–8301, Japan E-mail: [email protected] 2 Department of Ecosystem Studies, School of Environmental Science, The University of Shiga Prefecture, 2500 Hassaka, Hikone City, Shiga 522–8533, Japan 3 Lake Biwa Museum, 1091 Oroshimo, Kusatsu City, Shiga 525–0001, Japan Abstract: The fauna of adult digeneans (Trematoda) parasitic in freshwater fishes (Osteichthyes) from the Lake Biwa basin in Shiga Prefecture, central Honshu, Japan, is studied from the literature and existing specimens. Twenty-four previously known, 2 new, and 4 unidentified species in 17 gen- era and 12 families are recorded. Three dubious literature records are also mentioned. All 30 con- firmed species, except Sanguinicolidae gen. sp. (Aporocotylidae), are described and figured. Life cy- cles are discussed where known. Philopinna kawamutsu sp. nov. (Didymozoidae) was found in the connective tissue between the vertebrae and the air bladder near the esophagus of Nipponocypris tem- minckii (Temminck and Schlegel) (Cyprinidae). Genarchopsis yaritanago sp. nov. (Derogenidae) was found in the intestine of Tanakia lanceolata (Temminck and Schlegel) (Cyprinidae). Asymphylodora innominata (Faust, 1924) comb. nov. is proposed for A. macrostoma Ozaki, 1925 (Lissorchiidae). A key to the families, genera, and species of these digeneans is provided. Host-parasite and parasite- host lists are given. Key words: adult digeneans, Trematoda, parasites, morphology, life cycle, Philopinna kawamutsu sp. -

1 Title 1 Spring-Fed Streams As Flow Refugia: Changes in Fish
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.26.062968; this version posted April 28, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Title 2 Spring-fed streams as flow refugia: Changes in fish assemblages after a rainfall event 3 4 Authors 5 Masaru Sakai1 | Ryoshiro Wakiya2 | Gosuke Hoshi3 6 7 Affiliations 8 1 National Institute for Environmental Studies, Japan (Fukushima Branch), Fukushima, 9 Japan 10 2 Atmosphere and Ocean Research Institute, the University of Tokyo, Chiba, Japan 11 3 Graduate School of Science and Engineering, Chuo University, Tokyo, Japan 12 13 Correspondence 14 Masaru Sakai, National Institute for Environmental Studies, Japan (Fukushima Branch), 15 10-2 Fukasaku, Miharu, Tamura District, Fukushima 963-7700, Japan 16 Email: [email protected] 17 ORCID iD: 0000-0001-5361-0978 18 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.04.26.062968; this version posted April 28, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 19 Abstract 20 Understanding the structure and function of flow refugia in river networks is critical to 21 freshwater conservation in the context of anthropogenic changes in flow regimes. Safety 22 concerns associated with field data collection during floods have largely hindered 23 advances in assessing flow refugia for fishes; however, spring-fed streams can be safely 24 surveyed during rainfall events owing to their stable flow regimes. -

2014 Annual Meeting Program Oral Presentation
2014 Annual Meeting Program Papers to be presented at the 47th Annual Meeting of the Ichthyological Society of Japan at Kanagawa Prefectural Museum of Natural History, Novenber 14-17, 2014 Oral Presentation (Each presentation has been allowed 15 min, i.e. 12 min for presentation and 3 min for questions; *speaker) Oral Presentation Room #1 (SEISA Museum Theater, 1st Floor) November 15 (Saturday) Ecology, Conservation 1 09:30- Male and bisexual life history polymorphisms in anadromous fishes-II: Why does Arctic lamprey adopt a bisexual polymorphism? Akira GOTO* and Yuji YAMAZAKI 2 09:45- Habitat use of alien piscivorous chub establishing in small agricultural ditches Yoshihisa KURITA*, Norio ONIKURA and Ryutei INUI 3 10:00- Reproductive characteristics of Pseudorasbora sp. ,"Ushi-motsugo" under captive condition Moe MIYANISHI*, Tetsuro KITAGAWA, Yuka ODA and Kazumi HOSOYA 4 10:15- Spawning habitat selectivity of Honmoroko in inlets to Nishinoko and Ibanaiko Lagoon Takeshi KIKKO*, Daisuke ISHIZAKI, Yoshitaka KATAOKA and Yoshiaki KAI 5 10:30- Ecology of Lefua echigonia in the headwaters of the Tatarazawa, Atugi City-2 Hidetaka SUMIKURA* and Naoyuki SUGURO 6 10:45- Growth and longevity of the Japanese eight-barbel loach in a small marsh in the Kako River system Shigeru AOYAMA*, Tomohiro TABATA, Toshio DOI and Jinsuke AKADA <Break> <General Meeting of Members, 11:15-12:00 (Room #1)> <Lecture of the Awardee, 12:00-12:30 (Room #1)> <Lunch> <Poster Session Core Time on Odd-Numbered Paper, 13:30-14:30> Ecology, Conservation 7 14:30- Life history -

Origin and Phylogenetic Interrelationships of Teleosts Honoring Gloria Arratia
Origin and Phylogenetic Interrelationships of Teleosts Honoring Gloria Arratia Joseph S. Nelson, Hans-Peter Schultze & Mark V. H. Wilson (editors) TELEOSTEOMORPHA TELEOSTEI TELEOCEPHALA s. str. Leptolepis Pholidophorus † Lepisosteus Amia †? †? † †Varasichthyidae †Ichthyodectiformes Elopidae More advanced teleosts crown- group apomorphy-based group stem-based group Verlag Dr. Friedrich Pfeil • München Contents Preface ................................................................................................................................................................ 7 Acknowledgments ........................................................................................................................................... 9 Gloria Arratia’s contribution to our understanding of lower teleostean phylogeny and classifi cation – Joseph S. Nelson ....................................................................................... 11 The case for pycnodont fi shes as the fossil sister-group of teleosts – J. Ralph Nursall ...................... 37 Phylogeny of teleosts based on mitochondrial genome sequences – Richard E. Broughton ............. 61 Occipito-vertebral fusion in actinopterygians: conjecture, myth and reality. Part 1: Non-teleosts – Ralf Britz and G. David Johnson ................................................................................................................... 77 Occipito-vertebral fusion in actinopterygians: conjecture, myth and reality. Part 2: Teleosts – G. David Johnson and Ralf Britz .................................................................................................................. -
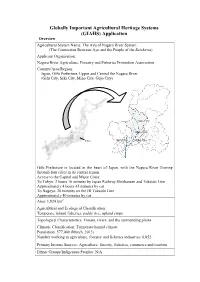
The Ayu of Nagara River System
Globally Important Agricultural Heritage Systems (GIAHS) Application Overview Agricultural System Name: The Ayu of Nagara River System (The Connection Between Ayu and the People of the Satokawa) Applicant Organisation: Nagara River Agriculture, Forestry and Fisheries Promotion Association Country/Area/Region: Japan, Gifu Prefecture, Upper and Central the Nagara River (Gifu City, Seki City, Mino City, Gujo City) Gujo City Mino City Tokyo Gifu City Seki City Gifu Prefecture is located in the heart of Japan, with the Nagara River flowing through four cities in its central region. Access to the Capital and Major Cities: To Tokyo: 2 hours 10 minutes by Japan Railway Shinkansen and Tokaido Line Approximately 4 hours 45 minutes by car To Nagoya: 20 minutes on the JR Tokaido Line Approximately 50 minutes by car Area: 1,824 km2 Agricultural and Ecological Classification: Temperate, inland fisheries, paddy rice, upland crops Topological Characteristics: Forests, rivers, and the surrounding plains Climatic Classification: Temperate humid climate Population: 577,000 (March, 2013) Number working in agriculture, forestry and fisheries industries: 6,052 Primary Income Sources: Agriculture, forestry, fisheries, commerce and tourism Ethnic Groups/Indigenous Peoples: N/A Explanation of Agricultural Heritage System On the upper and middle courses of the Nagara River located in Gifu Prefecture exist thriving inland fisheries which revolve around a species of Japanese sweetfish called “ayu” (Plecoglossus altivelis altivelis). Despite flowing through urban and residential areas, the pristine Nagara River that runs through the site’s centre boasts an abundance of clear, high quality water, and is also considered one of Japan’s three clearest rivers. The people of the region receive the river’s bounty and in turn strive to conserve it for future generations. -

Bayesian Node Dating Based on Probabilities of Fossil Sampling Supports Trans-Atlantic Dispersal of Cichlid Fishes
Supporting Information Bayesian Node Dating based on Probabilities of Fossil Sampling Supports Trans-Atlantic Dispersal of Cichlid Fishes Michael Matschiner,1,2y Zuzana Musilov´a,2,3 Julia M. I. Barth,1 Zuzana Starostov´a,3 Walter Salzburger,1,2 Mike Steel,4 and Remco Bouckaert5,6y Addresses: 1Centre for Ecological and Evolutionary Synthesis (CEES), Department of Biosciences, University of Oslo, Oslo, Norway 2Zoological Institute, University of Basel, Basel, Switzerland 3Department of Zoology, Faculty of Science, Charles University in Prague, Prague, Czech Republic 4Department of Mathematics and Statistics, University of Canterbury, Christchurch, New Zealand 5Department of Computer Science, University of Auckland, Auckland, New Zealand 6Computational Evolution Group, University of Auckland, Auckland, New Zealand yCorresponding author: E-mail: [email protected], [email protected] 1 Supplementary Text 1 1 Supplementary Text Supplementary Text S1: Sequencing protocols. Mitochondrial genomes of 26 cichlid species were amplified by long-range PCR followed by the 454 pyrosequencing on a GS Roche Junior platform. The primers for long-range PCR were designed specifically in the mitogenomic regions with low interspecific variability. The whole mitogenome of most species was amplified as three fragments using the following primer sets: for the region between position 2 500 bp and 7 300 bp (of mitogenome starting with tRNA-Phe), we used forward primers ZM2500F (5'-ACG ACC TCG ATG TTG GAT CAG GAC ATC C-3'), L2508KAW (Kawaguchi et al. 2001) or S-LA-16SF (Miya & Nishida 2000) and reverse primer ZM7350R (5'-TTA AGG CGT GGT CGT GGA AGT GAA GAA G-3'). The region between 7 300 bp and 12 300 bp was amplified using primers ZM7300F (5'-GCA CAT CCC TCC CAA CTA GGW TTT CAA GAT GC-3') and ZM12300R (5'-TTG CAC CAA GAG TTT TTG GTT CCT AAG ACC-3'). -
Genetic Catalogue, Biological Reference Collections and Online Database of European Marine Fishes
QUALITY OF LIFE AND MANAGEMENT OF LIVING RESOURCES PROGRAMME (1998-2002) Genetic Catalogue, Biological Reference Collections and Online Database of European Marine Fishes FINAL REPORT ANNEXES Contract number: QLRI-CT-2002-02755 Project acronym: FishTrace QoL action line: Area 14.1, Infrastructures Reporting period: 01/01/03 - 30/06/06 QLRI-CT-2002-02755 FishTrace INDEX Annexes Annex I: 1st FishTrace Plenary Meeting. Madrid, 12-14th February, 2003. Annex II: FishTrace Database Structuration Meeting. Paris, 13-14th May, 2003. Annex III: FishTrace Standardization Meeting. Stockholm, 4-7th June, 2003. Annex IV: 2nd FishTrace Plenary Meeting. Las Palmas de Gran Canaria (Spain), 19-21st November, 2003. Annex V: FishTrace Meeting on Data Validation. Ispra, Italy. 15-16th April, 2004. Annex VI: 3rd FishTrace Plenary Meeting. Paris, 25-27th November, 2004. Annex VII: FishTrace Database & Web Interface Meeting. Madrid, 11th March, 2005. Annex VIII: 4th FishTrace Plenary Meeting. Funchal (Portugal), 24-26th October, 2005. Annex IX: FishTrace Concluding Meeting. Kavala (Greece), May 10-12th, 2006. Annex X: Sampling and Taxonomy (WP2) Protocol. Annex XI: Results from Molecular Genetic Procedures Standardisation. Annex XII: Molecular identification and DNA barcoding (WP3). Protocol and PCR conditions. Annex XIII: Phylogenetic Validation of Sequences. Guidelines. Annex XIV: Preparing sequence files for the Sequin tool. Annex XV: Reference Collections (WP5) Protocol, Loan Request Form and Invoice of Specimens. Annex XVI: Guidelines for Validation purposes (WP7) including a protocol defining format for the Validation tasks. Annex XVII: Concise Protocol for the online validation process. Annex XVIII: The Bibliography Module in the FishTrace Database. Annex XIX: Results from the genetic population structure analysis. -

Of Mitochondrial Genomes for Protonibea Diacanthus
Table S1. Codon Number, Frequency and RSCU (Relative Synonymous Codon Usage) of mitochondrial genomes for Protonibea diacanthus. Amino acid Codon Number Frequency(%) RSCU Ala GCC 170 4.4702 1.9048 Ala GCA 115 3.0239 1.2885 Ala GCT 58 1.5251 0.6499 Ala GCG 14 0.3681 0.1569 Arg CGA 46 1.2096 2.3000 Arg CGC 22 0.5785 1.1000 Arg CGG 6 0.1578 0.3000 Arg CGT 6 0.1578 0.3000 Asn AAC 88 2.3140 1.4915 Asn AAT 30 0.7889 0.5085 Asp GAC 62 1.6303 1.6104 Asp GAT 15 0.3944 0.3896 Cys TGC 20 0.5259 1.5385 Cys TGT 6 0.1578 0.4615 Gln CAA 79 2.0773 1.7363 Gln CAG 12 0.3155 0.2637 Glu GAA 73 1.9195 1.4314 Glu GAG 29 0.7626 0.5686 Gly GGC 106 2.7873 1.7890 Gly GGA 62 1.6303 1.0464 Gly GGG 37 0.9729 0.6245 Gly GGT 32 0.8414 0.5401 His CAC 77 2.0247 1.4528 His CAT 29 0.7626 0.5472 Ile ATC 147 3.8654 1.0352 Ile ATT 137 3.6024 0.9648 Leu CTC 215 5.6534 1.9225 Leu CTA 180 4.7331 1.6095 Leu CTT 134 3.5235 1.1982 Leu TTA 74 1.9458 0.6617 Leu CTG 48 1.2622 0.4292 Leu TTG 20 0.5259 0.1788 Lys AAA 69 1.8144 1.7692 Lys AAG 9 0.2367 0.2308 Met ATA 88 2.3140 1.2754 Met ATG 50 1.3148 0.7246 Phe TTC 151 3.9705 1.2227 Phe TTT 96 2.5243 0.7773 Pro CCC 123 3.2343 2.1674 Pro CCA 51 1.3410 0.8987 Pro CCT 41 1.0781 0.7225 Pro CCG 12 0.3155 0.2115 Ser TCC 79 2.0773 2.1161 Ser TCA 61 1.6040 1.6339 Ser AGC 34 0.8940 0.9107 Ser TCT 28 0.7363 0.7500 Ser AGT 14 0.3681 0.3750 Ser TCG 8 0.2104 0.2143 Stp TAA 4 0.1052 2.0000 Stp AGA 2 0.0526 1.0000 Stp TAG 2 0.0526 1.0000 Stp AGG 0 0.0000 0.0000 Thr ACC 136 3.5761 1.8133 Thr ACA 103 2.7084 1.3733 Thr ACT 51 1.3410 0.6800 Thr ACG 10 0.2630 0.1333 Trp TGA 93 2.4454 1.5630 Trp TGG 26 0.6837 0.4370 Tyr TAC 81 2.1299 1.3729 Tyr TAT 37 0.9729 0.6271 Val GTA 61 1.6040 1.2513 Val GTC 60 1.5777 1.2308 Val GTT 50 1.3148 1.0256 Val GTG 24 0.6311 0.4923 Tab. -

Horizontal and Vertical Distribution of Cottus Reinii (Pisces: Cottidae) in Lake Biwa, Central Japan
Biogeography 17. 13–20. Sep. 20, 2015 Horizontal and vertical distribution of Cottus reinii (Pisces: Cottidae) in Lake Biwa, central Japan Yasuhiro Fujioka1,2*, Akihisa Sakai1 and Atsuhiko Ide1 1 Shiga Prefecture Fisheries Experiment Station, Hassaka 2138-3, Hikone, Shiga, Japan 2 Lake Biwa Museum, Oroshimo 1091, Kusatsu, Shiga, 525-0001 Japan Abstract. The distribution of Utsusemi-kazika Cottus reinii in Lake Biwa was investigated by means of small trawl-net, commercial trawl-nets, and commercial set-nets. C. reinii were abundant in the coastal bot- tom waters deeper than 20 m between June and August and deeper than 30 m in October. These were mostly small fish of less than 4 cm SL, showing a mode in the 1-1.99 cm class in June. Fish of more than 4 cm SL also were caught in coastal waters (10-20 m in depth) and offshore deep waters (deeper than 40 m) throughout the year. These results suggest that this species leaves the plankton to begin its benthic life at a size of about 1 cm SL at depths of 20-30 m. After growing to 3-4 cm SL by October at depths of 30-40 m, they expand their range to bottom waters deeper than 40 m. The bottom sediment of the lake habitat is mainly fine mud, very different from the reported of gravel-bottom habitat of this fish in rivers. Key words: Cottus reinii, Utsusemi-kazika, Distribution, Lake Biwa Introduction is chiefly distributed in shallow coastal waters with stony bottoms and occasionally in muddy deep wa- The genus Cottus has a circumpolar distribution ters, and it spawns in rivers (Nakamura, 1975; Goto, in the northern hemisphere and includes more than 1995). -

Complete Mitochondrial Genome of the Amur Sculpin Cottus Szanaga (Cottoidei: Cottidae)
Mitochondrial DNA Part B Resources ISSN: (Print) 2380-2359 (Online) Journal homepage: http://www.tandfonline.com/loi/tmdn20 Complete mitochondrial genome of the Amur sculpin Cottus szanaga (Cottoidei: Cottidae) Evgeniy S. Balakirev, Pavel A. Saveliev & Francisco J. Ayala To cite this article: Evgeniy S. Balakirev, Pavel A. Saveliev & Francisco J. Ayala (2016) Complete mitochondrial genome of the Amur sculpin Cottus szanaga (Cottoidei: Cottidae), Mitochondrial DNA Part B, 1:1, 737-738, DOI: 10.1080/23802359.2016.1233470 To link to this article: http://dx.doi.org/10.1080/23802359.2016.1233470 © 2016 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group. Published online: 03 Oct 2016. Submit your article to this journal View related articles View Crossmark data Full Terms & Conditions of access and use can be found at http://www.tandfonline.com/action/journalInformation?journalCode=tmdn20 Download by: [The UC Irvine Libraries] Date: 03 October 2016, At: 11:39 MITOCHONDRIAL DNA PART B: RESOURCES, 2016 VOL. 1, NO. 1, 737–738 http://dx.doi.org/10.1080/23802359.2016.1233470 MITOGENOME ANNOUNCEMENT Complete mitochondrial genome of the Amur sculpin Cottus szanaga (Cottoidei: Cottidae)* Evgeniy S. Balakireva,b,c, Pavel A. Savelievb and Francisco J. Ayalaa aDepartment of Ecology and Evolutionary Biology, University of California, Irvine, CA, USA; bA. V. Zhirmunsky Institute of Marine Biology, Far Eastern Branch, Russian Academy of Science, Vladivostok, Russia; cFar Eastern Federal University, School of Natural Sciences, Vladivostok, Russia ABSTRACT ARTICLE HISTORY The complete mitochondrial genome was sequenced in two individuals of the Amur sculpin Cottus sza- Received 2 September 2016 naga. -

Scorpaeniformes: Platycephalidae) and the Phylogenetic Relationships Within Scorpaeniformes Based on Whole Mitogenomes
G C A T T A C G G C A T genes Article The First Complete Mitochondrial Genome of the Flathead Cociella crocodilus (Scorpaeniformes: Platycephalidae) and the Phylogenetic Relationships within Scorpaeniformes Based on Whole Mitogenomes 1, 1, 1 2 1, 1, Lei Cui y, Rongbo Cao y, Yuelei Dong , Xingchen Gao , Jingyi Cen * and Songhui Lu * 1 Key Laboratory of Eutrophication and Red Tide Prevention, Research Center for Harmful Algae and Marine Biology, Jinan University, Guangzhou 510632, China 2 Chinese Sturgeon Research Institute, Three Gorges Corporation, Yichang 443100, China * Correspondence: [email protected] (J.C.); [email protected] (S.L.); Tel.: +86-20-8522-5913 (J.C.); +86-20-8522-2720 (S.L.) These authors contributed equally to this work. y Received: 21 May 2019; Accepted: 12 July 2019; Published: 15 July 2019 Abstract: Complete mitochondrial genomes (mitogenomes) are important molecular markers for understanding the phylogenetics of various species. Although recent studies on the mitogenomes of the Scorpaeniformes species have been greatly advanced, information regarding molecular studies and the taxonomic localization of Platycephalidae is still sparse. To further analyze the phylogeny of Platycephalidae, we sequenced the complete mitogenome of Cociella crocodilus of the Platycephalidae family within Scorpaeniformes for the first time. The mitogenome was 17,314 bp in length, contained two ribosomal RNA genes (rRNAs), 22 transfer RNA genes (tRNAs), 13 protein-coding genes (PCGs), and two typical noncoding control regions (the control region (CR) and origin of the light strand (OL)). All PCGs used standard initiation codons ATG, apart from cox1. The majority of the tRNA genes could be folded into cloverleaf secondary structures, whereas the secondary structure of tRNASer (AGN) lacked a dihydrouridine (DHU) stem. -

Species Divers. 20(1): 73-81
Species Diversity 20: 73–81 25 May 2015 DOI: 10.12782/sd.20.1.073 Rediscovery of a Fish Acanthocephalan, Acanthocephalus minor (Echinorhynchida: Echinorhynchidae), in the Lake Biwa Basin, Central Japan, with a Review of the Fish Acanthocephalan Fauna of the Basin Kazuya Nagasawa1,2 and Masato Nitta1 1 Graduate School of Biosphere Science, Hiroshima University, 1-4-4 Kagamiyama, Higashi-Hiroshima, Hiroshima 739-8528, Japan E-mail: [email protected] (KN); [email protected] (MN) 2 Corresponding author (Received 28 January 2015; Accepted 30 April 2015) Specimens of the echinorhynchid acanthocephalan Acanthocephalus minor Yamaguti, 1935 were collected from the rectum of a dark sleeper, Odontobutis obscura (Temminck and Schlegel, 1845), in an irrigation canal near Lake Biwa, Shiga Prefecture, west-central Japan. This represents a rediscovery of A. minor in the Lake Biwa basin after a gap of nearly 80 years. It appears not to be distributed in Lake Biwa proper, but to occur rarely in rivers and irrigation canals of a limited coastal area around the lake. To date, 10 nominal species of acanthocephalan in four families and three orders have been re- ported from fish in the Lake Biwa basin. Among these, striking morphological similarities between Acanthocephalus aculea- tus Van Cleave, 1931 and its congener A. opsariichthydis Yamaguti, 1935 are noted. It is furthermore suggested that A. gotoi Van Cleave, 1925 could not maintain its population after the basin’s wild population of its major host, Anguilla japonica Temminck and Schlegel, 1846, disappeared in the mid-1960s. Key Words: Acanthocephalus minor, Acanthocephala, Odontobutis obscura, fish parasite, faunistic review, Lake Biwa basin, Japan.