2,3-Butanediol Biosynthesis in E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Flavour Management by Citric Acid Negative MLF Starter Cultures Authors: Carsten Heinemeyer, 2B Fermcontrol; Ulrich Hamm DLR-RNH Bad Kreuznach, Germany; Dr
Flavour management by citric acid negative MLF starter cultures Authors: Carsten Heinemeyer, 2B FermControl; Ulrich Hamm DLR-RNH Bad Kreuznach, Germany; Dr. Jürgen Fröhlich, University of Mainz General will be completed in all fermentations. Additionally, acetic acid and a certain residual profile of the metabolic intermediates The malo-lactic fermentation (MLF) is a commonly used remain in the wine. method to convert the aggressive malic acid to lactic acid. This conversion results in a reduction of the titratable acidity, which New MLF Starter Cultures is desired mainly in red wine but also in numerous white wines. The development of a citric acid negative MLF starter culture This process will be done either by the indigenous flora of LAB gave a new opportunity to avoid diacetyl and acetic acid or by selected strains of LAB e.g. Oenococcus oeni. During the production from the citric acid degradation. The fermentation MLF Oenococcus oeni does not convert only malic acid into lactic with a citric acid negative MLF starter culture will best preserve acid, numerous amounts of aroma active by-products will also be the varietal character of the wine. produced. The best known is diacetyl, which gives buttery notes to the wine. Diacetyl will be produced during the MLF by the In a comprehensive four year study at the wine research institute conversion of the natural citric acid in the wine by Oenococcus DLR-RNH Bad Kreuznach in Rhineland-Palatinate-Germany, oeni (Jan Clair Nielsen 1999). Apart from the diacetyl formation different MLF starter strains were tested under practical by Oenococcus oeni, numerous intermediate by-products are winemaking conditions. -

Acetoin Acetate Natural
aurochemicals.com Safety Data Sheet HEALTH 0 FLAMMABILITY 2 REACTIVITY 0 Section 1: PRODUCT AND COMPANY IDENTIFICATION 1.1 Product identifiers Product Name Acetoin Acetate, Natural Product Number 0352600 CAS-No. 4906-24-5 1.2 Product Recommended Use Flavorings 1.3 Preparation Information Company Aurochemicals 7 Nicoll Street Washingtonville, NY 10992- USA Telephone 845-496-6065 Fax 845-496-6248 1.4 Emergency Telephone Number 1-800-535-5053 International - 1-352-323-3500 collect Section 2: HAZARD(s) IDENTIFICATION 2.1 Classification of substance or mixture GHS Classification in accordance with 29 CFR 1910 (OSHA HCS) Combustible Liquid (Category 4) H227 Skin irritation (Category 2) H315 Eye Irritation (Category 2A) H319 Specific Target Organ Toxicity -Single Exposure -Respiratory irritation H335 2.2 GHS Label Elements, Including precautionary statements Pictogram Signal Statement Warning Hazard Statement(s) H227 Combustible liquid H315 Causes skin irritation H319 Causes serious eye irritation H335 May cause respiratory irritation Precautionary Statement(s) PREVENTION P210 Keep away from flames and hot surfaces - No smoking P216 Avoid breathing dust/fumes/gas/mist/vapors/spray P280 Wear protective gloves/protective clothing/face protection 3526 ACETOIN ACETATE Nat sds.doc Printed: April 17, 2019 Page 1 of 7 aurochemicals.com Safety Data Sheet RESPONSE P305+351+338 IF IN EYES: Rinse cautiously with water for several minutes. Remove contact lenses if present and easy to do. Continue rinsing. STORAGE P405 Store locked up DISPOSAL P501 Dispose of contents/container to an approved waste disposal plant 2.3 HNOC (Hazards not otherwise None classified or not covered by GHS Section 3: COMPOSITION / INFORMATION ON INGREDIENTS 3.1 Substances Synonym 2-Acetoxy-3-Butanone Formula C6H10O3 Molecular Weight 130.14 g/mol CAS-No 4906-24-5 EC-No 200-580-7 Hazardous Components Does not contain any hazardous substances Section 4: FIRST AID MEASURES 4.1 Description of first aid measures General Advice Consult a physician. -

Retention Indices for Frequently Reported Compounds of Plant Essential Oils
Retention Indices for Frequently Reported Compounds of Plant Essential Oils V. I. Babushok,a) P. J. Linstrom, and I. G. Zenkevichb) National Institute of Standards and Technology, Gaithersburg, Maryland 20899, USA (Received 1 August 2011; accepted 27 September 2011; published online 29 November 2011) Gas chromatographic retention indices were evaluated for 505 frequently reported plant essential oil components using a large retention index database. Retention data are presented for three types of commonly used stationary phases: dimethyl silicone (nonpolar), dimethyl sili- cone with 5% phenyl groups (slightly polar), and polyethylene glycol (polar) stationary phases. The evaluations are based on the treatment of multiple measurements with the number of data records ranging from about 5 to 800 per compound. Data analysis was limited to temperature programmed conditions. The data reported include the average and median values of retention index with standard deviations and confidence intervals. VC 2011 by the U.S. Secretary of Commerce on behalf of the United States. All rights reserved. [doi:10.1063/1.3653552] Key words: essential oils; gas chromatography; Kova´ts indices; linear indices; retention indices; identification; flavor; olfaction. CONTENTS 1. Introduction The practical applications of plant essential oils are very 1. Introduction................................ 1 diverse. They are used for the production of food, drugs, per- fumes, aromatherapy, and many other applications.1–4 The 2. Retention Indices ........................... 2 need for identification of essential oil components ranges 3. Retention Data Presentation and Discussion . 2 from product quality control to basic research. The identifi- 4. Summary.................................. 45 cation of unknown compounds remains a complex problem, in spite of great progress made in analytical techniques over 5. -

Oenological Impact of the Hanseniaspora/Kloeckera Yeast Genus on Wines—A Review
fermentation Review Oenological Impact of the Hanseniaspora/Kloeckera Yeast Genus on Wines—A Review Valentina Martin, Maria Jose Valera , Karina Medina, Eduardo Boido and Francisco Carrau * Enology and Fermentation Biotechnology Area, Food Science and Technology Department, Facultad de Quimica, Universidad de la Republica, Montevideo 11800, Uruguay; [email protected] (V.M.); [email protected] (M.J.V.); [email protected] (K.M.); [email protected] (E.B.) * Correspondence: [email protected]; Tel.: +598-292-481-94 Received: 7 August 2018; Accepted: 5 September 2018; Published: 10 September 2018 Abstract: Apiculate yeasts of the genus Hanseniaspora/Kloeckera are the main species present on mature grapes and play a significant role at the beginning of fermentation, producing enzymes and aroma compounds that expand the diversity of wine color and flavor. Ten species of the genus Hanseniaspora have been recovered from grapes and are associated in two groups: H. valbyensis, H. guilliermondii, H. uvarum, H. opuntiae, H. thailandica, H. meyeri, and H. clermontiae; and H. vineae, H. osmophila, and H. occidentalis. This review focuses on the application of some strains belonging to this genus in co-fermentation with Saccharomyces cerevisiae that demonstrates their positive contribution to winemaking. Some consistent results have shown more intense flavors and complex, full-bodied wines, compared with wines produced by the use of S. cerevisiae alone. Recent genetic and physiologic studies have improved the knowledge of the Hanseniaspora/Kloeckera species. Significant increases in acetyl esters, benzenoids, and sesquiterpene flavor compounds, and relative decreases in alcohols and acids have been reported, due to different fermentation pathways compared to conventional wine yeasts. -

An Original Method for Producing Acetaldehyde and Diacetyl by Yeast
b r a z i l i a n j o u r n a l o f m i c r o b i o l o g y 4 7 (2 0 1 6) 949–954 ht tp://www.bjmicrobiol.com.br/ Industrial Microbiology An original method for producing acetaldehyde and diacetyl by yeast fermentation a a,∗ a b a,c Irina Rosca , Anca Roxana Petrovici , Mihai Brebu , Irina Stoica , Bogdan Minea , a Narcisa Marangoci a “Petru Poni” Institute of Macromolecular Chemistry, Advanced Research Center for Bionanoconjugates and Biopolymers, Aleea GrigoreGhica Voda, Iasi, Romania b SC Zeelandia SRL, R&D Department, Valea Lupului, Iasi, Romania c “Grigore T. Popa” University of Medicine and Pharmacy, Iasi, Romania a r a t i c l e i n f o b s t r a c t Article history: In this study a natural culture medium that mimics the synthetic yeast peptone glucose Received 2 June 2015 medium used for yeast fermentations was designed to screen and select yeasts capable Accepted 29 February 2016 of producing high levels of diacetyl and acetaldehyde. The presence of whey powder and Available online 25 July 2016 sodium citrate in the medium along with manganese and magnesium sulfate enhanced Associate Editor: Jorge Gonzalo both biomass and aroma development. A total of 52 yeasts strains were cultivated in two Farias Avendano different culture media, namely, yeast peptone glucose medium and yeast acetaldehyde- diacetyl medium. The initial screening of the strains was based on the qualitative reaction Keywords: of the acetaldehyde with Schiff’s reagent (violet color) and diacetyl with Brady’s reagent Acetaldehyde (yellow precipitate). -

An Unrecognized Hazard in E-Cigarette Vapor: Preliminary Quantification of Methylglyoxal Formation from Propylene Glycol in E-Cigarettes
International Journal of Environmental Research and Public Health Article An Unrecognized Hazard in E-Cigarette Vapor: Preliminary Quantification of Methylglyoxal Formation from Propylene Glycol in E-Cigarettes Parham Azimi 1, Zahra Keshavarz 1, Marianne Lahaie Luna 1,2, Jose Guillermo Cedeno Laurent 1 , Jose Vallarino 1, David C. Christiani 1 and Joseph G. Allen 1,* 1 Department of Environmental Health, Harvard T. H. Chan School of Public Health, Boston, MA 02115, USA; [email protected] (P.A.); [email protected] (Z.K.); [email protected] (M.L.L.); [email protected] (J.G.C.L.); [email protected] (J.V.); [email protected] (D.C.C.) 2 Occupational & Environmental Health Division, Dalla Lana School of Public Health, University of Toronto, Toronto, ON M5T 3M7, Canada * Correspondence: [email protected] Abstract: Up to 95% of the liquid volume in an e-cigarette consists of propylene glycol. Previous research has shown that propylene glycol can generate diacetyl and formaldehyde when heated. New research shows that propylene glycol can also generate methylglyoxal, an alpha di-carbonyl compound recently shown to cause epithelial necrosis at even lower concentrations than diacetyl, the flavoring chemical associated with bronchiolitis obliterans (“Popcorn Lung”). We analyzed chemical emissions from 13 JUUL pod flavors. Diacetyl and methylglyoxal was detected in 100% of samples 3 3 with median concentration (range) of 20 µg/m (less than limit of quantification: 54 µg/m ) and 4219 µg/m3 (677–15,342 µg/m3), respectively. We also detected acetaldehyde (median concentration: Citation: Azimi, P.; Keshavarz, Z.; 341 µg/m3) and propionaldehyde (median concentration: 87 µg/m3) in all samples. -
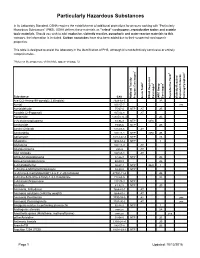
Particularly Hazardous Substances
Particularly Hazardous Substances In its Laboratory Standard, OSHA requires the establishment of additional protections for persons working with "Particularly Hazardous Substances" (PHS). OSHA defines these materials as "select" carcinogens, reproductive toxins and acutely toxic materials. Should you wish to add: explosive, violently reactive, pyrophoric and water-reactve materials to this category, the information is included. Carbon nanotubes have also been added due to their suspected carcinogenic properties. This table is designed to assist the laboratory in the identification of PHS, although it is not definitively conclusive or entirely comprehensive. *Notes on the proper use of this table appear on page 12. 1 6 5 2 3 4 Substance CAS National Toxicity National Program Carcinogen Toxin Acute Regulated OSHA Carcinogen Group IARC Carcinogen Toxin Reproductive Violently Reactive/ Explosive/Peroxide Forming/Pyrophoric A-a-C(2-Amino-9H-pyrido[2,3,b]indole) 2648-68-5 2B Acetal 105-57-7 yes Acetaldehyde 75-07-0 NTP AT 2B Acrolein (2-Propenal) 107-02-8 AT Acetamide 126850-14-4 2B 2-Acetylaminofluorene 53-96-3 NTP ORC Acrylamide 79-06-6 NTP 2B Acrylyl Chloride 814-68-6 AT Acrylonitrile 107-13-1 NTP ORC 2B Adriamycin 23214-92-8 NTP 2A Aflatoxins 1402-68-2 NTP 1 Allylamine 107-11-9 AT Alkylaluminums varies AT Allyl Chloride 107-05-1 AT ortho-Aminoazotoluene 97-56-3 NTP 2B para-aminoazobenzene 60-09-3 2B 4-Aminobiphenyl 92-67-1 NTP ORC 1 1-Amino-2-Methylanthraquinone 82-28-0 NTP (2-Amino-6-methyldipyrido[1,2-a:3’,2’-d]imidazole) 67730-11-4 2B -

Recommendation from the Scientific Committee on Occupational Exposure Limits for Diacetyl
Employment, Social Affairs & Inclusion SCOEL Recommendation on Diacetyl Recommendation from the Scientific Committee on Occupational Exposure Limits for Diacetyl SCOEL/SUM/149 June 2014 June 2014 1 Employment, Social Affairs & Inclusion SCOEL Recommendation on Diacetyl Table of Contents 1. Substance identification, physico-chemical properties.............................................. 3 2. Occurrence/use and occupational exposure ........................................................... 3 2.1. Occurrence and use ...................................................................................... 3 2.2. Occupational exposure .................................................................................. 4 2.3. Methods of exposure monitoring and analysis .................................................. 5 3. Health significance ............................................................................................. 6 3.1. Toxicokinetics .............................................................................................. 7 3.1.1. Human data ........................................................................................... 7 3.1.2. Animal data ........................................................................................... 7 3.1.3. Biological monitoring ............................................................................... 8 3.2. Acute toxicity ............................................................................................... 8 3.2.1. Human data .......................................................................................... -

Substituted Deoxyguanosine Nucleosides from 2‐
Synthesis of N2-Substituted Deoxyguanosine UNIT 1.3 Nucleosides from 2-Fluoro-6-O- (Trimethylsilylethyl)-2′-Deoxyinosine This unit describes the synthesis of 2-fluoro-6-O-(trimethylsilylethyl)-2′-deoxyinosine and gives examples of its use for the preparation of N2-substituted deoxyguanosine nucleosides. Such nucleoside derivatives are used for a variety of purposes including chemotherapy, enzyme mechanism studies, nuclear magnetic resonance (NMR) studies (when isotopically labeled), and as synthetic standards for identification of adducts formed by the reaction of DNA with xenobiotics. In addition, the O6-protected 2-fluoro- 2′-deoxyinosine compounds can be converted to phosphoramidites and used in the synthesis of oligonucleotides, thus allowing substitution reactions to be carried out after oligonucleotide assembly. 2-Halopurine derivatives have been used for many years for the preparation of N2-sub- stituted guanosine derivatives, with the 2-fluoro substituent being the most easily dis- placed by nucleophiles (Montgomery and Hewson, 1960; Gerster and Robins, 1965, 1966). The 2-fluoro group is introduced by aqueous diazotization of guanosine in the presence of potassium fluoride or fluoroboric acid. However, these conditions are too harsh for 2′-deoxyguanosine and lead to depurination; hence, different synthetic method- ology is needed for the deoxynucleoside. The fluorine atom can be introduced success- fully by diazotization under anhydrous conditions with t-butyl nitrite as the diazotizing agent and HF in pyridine as the fluoride source (Robins and Uznanski, 1981; Lee et al., 1990; Harris et al., 1991). Success in the fluoridation step requires protection of the C6 oxygen group, which is done by Mitsunobu alkylation (Mitsunobu, 1981) with trimethyl- silylethanol or other alcohols. -

Herbicide Assays for Predicting Or Determining Plant Responses in Aquatic Systems
J. Aquat. Plant Manage. 56s: 67–73 Herbicide assays for predicting or determining plant responses in aquatic systems GREG MACDONALD AND MICHAEL NETHERLAND* INTRODUCTION GROWTH RESPONSE STUDIES Herbicides are an important component of aquatic plant Growth studies can provide useful information, particu- management, but represent a relatively small sector of larly as it relates to activity on a whole plant basis. In herbicide use compared to agriculture and other commer- addition to visual assessment, differential response to a cial production systems. There are over 150 active herbicide herbicide can be measured through stem length, leaf ingredients registered for use in cropping systems, but only number, plant biomass, growth rates, reproductive devel- 13 are registered in aquatic systems in the United States. opment (propagule number and viability), etc. Regression There is a considerable amount of information regarding analysis to model the response can be used to generate the activity and responses of terrestrial plant species (both inhibition (I50,I90 values) to a particular parameter as a weeds and crops) to herbicides, but comparatively little for function of herbicide rate. These studies are useful to aquatic plants and aquatic herbicides. With this in mind, the evaluate the response of different populations of the same purpose of this article is to provide a framework for species to assess changes or differences in herbicide researchers and practitioners who seek to better understand response. such as the development of resistance (Puri et and evaluate the response of both invasive and native al. 2007). Growth experiments are also useful in determining aquatic plants to herbicides. the level, as function of herbicide rate, of off-target damage We will focus on herbicides registered for use in aquatic to desirable species. -

Diacetyl Take a Single Short Sniff
beer flavor standard 2,3-butanedione OFF-FLAVOR KIT available from www.cicerone.org ASSESSMENT CONFUSIONS Without covering the glass, swirl • Butyric acid the beer to release the aroma. • Vanillin Diacetyl Take a single short sniff. Repeat • Isobutyraldehyde “like butter, or as necessary. IMPORTANCE butter popcorn” THRESHOLD stouts 10 – 40 µg/l and lagers, eg in other lager beers. Considerable ORIGINS efforts are made by breweries to Produced in beer from a precursor formed by yeast during fermentation. Can also be formed REMARKS by contaminant lactic acid bacteria. 2,3-Butanedione is one of two vicinal diketones found in beer. O The ratio of diacetyl to CH3 pentanedione concentrations can H3 C be used as an indicator of bacterial contamination in beer. TION O A www.aroxa.com CAS NUMBER [email protected] 431-03-8 ©2013 CARA TECHNOLOGY LTD % POPUL RANDALLS ROAD LEATHERHEAD, SURREY FLAVOR THRESHOLD KT22 7RY, UK beer flavor standard dimethyl sulfide OFF-FLAVOR KIT available from www.cicerone.org ASSESSMENT CONFUSIONS Without covering the glass, swirl • Methyl thioacetate the beer to release the aroma. • Ethanethiol DMS Take a single short sniff. Repeat • Dimethyl trisulphide “like sweetcorn or as necessary. IMPORTANCE tomato sauce” THRESHOLD 30 - 50 µg/l beers. Excessive levels are indicative ORIGINS of growth of contaminant bacteria Formed from malt-derived during fermentation. precursors, primarily during wort production and – to a lesser extent – REMARKS during fermentation. The perception of dimethyl sulfide H3 C CH3 aromatic higher -

Thermophilic Fermentation of Acetoin and 2, 3-Butanediol by a Novel
Xiao et al. Biotechnology for Biofuels 2012, 5:88 http://www.biotechnologyforbiofuels.com/content/5/1/88 RESEARCH Open Access Thermophilic fermentation of acetoin and 2, 3-butanediol by a novel Geobacillus strain Zijun Xiao1*, Xiangming Wang1, Yunling Huang1, Fangfang Huo1, Xiankun Zhu1, Lijun Xi1 and Jian R Lu2* Abstract Background: Acetoin and 2,3-butanediol are two important biorefinery platform chemicals. They are currently fermented below 40°C using mesophilic strains, but the processes often suffer from bacterial contamination. Results: This work reports the isolation and identification of a novel aerobic Geobacillus strain XT15 capable of producing both of these chemicals under elevated temperatures, thus reducing the risk of bacterial contamination. The optimum growth temperature was found to be between 45 and 55°C and the medium initial pH to be 8.0. In addition to glucose, galactose, mannitol, arabionose, and xylose were all acceptable substrates, enabling the potential use of cellulosic biomass as the feedstock. XT15 preferred organic nitrogen sources including corn steep liquor powder, a cheap by-product from corn wet-milling. At 55°C, 7.7 g/L of acetoin and 14.5 g/L of 2,3-butanediol could be obtained using corn steep liquor powder as a nitrogen source. Thirteen volatile products from the cultivation broth of XT15 were identified by gas chromatography–mass spectrometry. Acetoin, 2,3-butanediol, and their derivatives including a novel metabolite 2,3-dihydroxy-3-methylheptan-4-one, accounted for a total of about 96% of all the volatile products. In contrast, organic acids and other products were minor by-products.