Doubly Heterozygous LMNA and TTN Mutations Revealed by Exome Sequencing in a Severe Form of Dilated Cardiomyopathy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Lamin-Related Congenital Muscular Dystrophy Alters Mechanical Signaling and Skeletal Muscle Growth
International Journal of Molecular Sciences Article Lamin-Related Congenital Muscular Dystrophy Alters Mechanical Signaling and Skeletal Muscle Growth Daniel J. Owens 1,2 , Julien Messéant 1, Sophie Moog 3, Mark Viggars 2, Arnaud Ferry 1,4, Kamel Mamchaoui 1,5, Emmanuelle Lacène 5 , Norma Roméro 5,6, Astrid Brull 1 , Gisèle Bonne 1 , Gillian Butler-Browne 1 and Catherine Coirault 1,* 1 Center for Research in Myology, Sorbonne Université, INSERM UMRS_974, 75013 Paris, France; [email protected] (D.J.O.); [email protected] (J.M.); [email protected] (A.F.); [email protected] (K.M.); [email protected] (A.B.); [email protected] (G.B.); [email protected] (G.B.-B.) 2 Research Institute for Sport and Exercise Science, Liverpool John Moores University, Liverpool L3 3AF, UK; [email protected] 3 Inovarion, 75005 Paris, France; [email protected] 4 Université de Paris, 75006 Paris, France 5 Neuromuscular Morphology Unit, Institute of Myology, Pitié-Salpêtrière Hospital, 75013 Paris, France; [email protected] (E.L.); [email protected] (N.R.) 6 APHP, Reference Center for Neuromuscular Disorders, Pitié-Salpêtrière Hospital, Institute of Myology, 75013 Paris, France * Correspondence: [email protected]; Tel.: +33-1-1-4216-5708 Abstract: Laminopathies are a clinically heterogeneous group of disorders caused by mutations in the LMNA gene, which encodes the nuclear envelope proteins lamins A and C. The most frequent diseases associated with LMNA mutations are characterized by skeletal and cardiac involvement, and include autosomal dominant Emery–Dreifuss muscular dystrophy (EDMD), limb-girdle muscular Citation: Owens, D.J.; Messéant, J.; dystrophy type 1B, and LMNA-related congenital muscular dystrophy (LMNA-CMD). -

ARVC-Variants.Pdf
Updated gene list responsible for ARVC/D pathology Subtype Gene Location Reference ARVC1 TGFB3 14q24.3 Beffagna G, Occhi G, Nava A, et al. Regulatory mutations in transforming growth factor-beta3 gene cause arrhythmogenic right ventricular cardiomyopathy type 1. Cardiovasc Res. 2005;65:366–73. ARVC2 RYR2 1q43 Tiso N, Stephan DA, Nava A, et al. Identification of mutations in the cardiac ryanodine receptor gene in families affected with arrhythmogenic right ventricular cardiomyopathy type 2 (ARVD2). Hum Mol Genet. 2001;10:189–94. ARVC3 Unknown 14q12-q22 Severini GM, Krajinovic M, Pinamonti B, et al. A new locus for arrhythmogenic right ventricular dysplasia on the long arm of chromosome 14. Genomics. 1996;31:193–200. ARVC4 TTN 2q32.1-q32.3 Taylor M, Graw S, Sinagra G, et al. Genetic variation in titin in arrhythmogenic right ventricular cardiomyopathy-overlap syndromes. Circulation. 2011;124:876–85. ARVC5 TMEM43 3p25.1 Merner ND, Hodgkinson KA, Haywood AF, et al. Arrhythmogenic right ventricular cardiomyopathy type 5 is a fully penetrant, lethal arrhythmic disorder caused by a missense mutation in the TMEM43 gene. Am J Hum Genet. 2008;82:809–21. ARVC6 Unknown 10p14-p12 Li D, Ahmad F, Gardner MJ, et al. The locus of a novel gene responsible for arrhythmogenic right- ventricular dysplasia characterized by early onset and high penetrance maps to chromosome 10p12-p14. Am J Hum Genet. 2000;66:148–56. ARVC7 DES 2q35 Klauke B, Kossmann S, Gaertner A, et al. De novo desmin-mutation N116S is associated with arrhythmogenic right ventricular cardiomyopathy. Hum Mol Genet. 2010;19:4595–607. -

Genetic Mutations and Mechanisms in Dilated Cardiomyopathy
Genetic mutations and mechanisms in dilated cardiomyopathy Elizabeth M. McNally, … , Jessica R. Golbus, Megan J. Puckelwartz J Clin Invest. 2013;123(1):19-26. https://doi.org/10.1172/JCI62862. Review Series Genetic mutations account for a significant percentage of cardiomyopathies, which are a leading cause of congestive heart failure. In hypertrophic cardiomyopathy (HCM), cardiac output is limited by the thickened myocardium through impaired filling and outflow. Mutations in the genes encoding the thick filament components myosin heavy chain and myosin binding protein C (MYH7 and MYBPC3) together explain 75% of inherited HCMs, leading to the observation that HCM is a disease of the sarcomere. Many mutations are “private” or rare variants, often unique to families. In contrast, dilated cardiomyopathy (DCM) is far more genetically heterogeneous, with mutations in genes encoding cytoskeletal, nucleoskeletal, mitochondrial, and calcium-handling proteins. DCM is characterized by enlarged ventricular dimensions and impaired systolic and diastolic function. Private mutations account for most DCMs, with few hotspots or recurring mutations. More than 50 single genes are linked to inherited DCM, including many genes that also link to HCM. Relatively few clinical clues guide the diagnosis of inherited DCM, but emerging evidence supports the use of genetic testing to identify those patients at risk for faster disease progression, congestive heart failure, and arrhythmia. Find the latest version: https://jci.me/62862/pdf Review series Genetic mutations and mechanisms in dilated cardiomyopathy Elizabeth M. McNally, Jessica R. Golbus, and Megan J. Puckelwartz Department of Human Genetics, University of Chicago, Chicago, Illinois, USA. Genetic mutations account for a significant percentage of cardiomyopathies, which are a leading cause of conges- tive heart failure. -

Nuclear Titin Interacts with A- and B-Type Lamins in Vitro and in Vivo
Research Article 239 Nuclear Titin interacts with A- and B-type lamins in vitro and in vivo Michael S. Zastrow1,*, Denise B. Flaherty2‡, Guy M. Benian2 and Katherine L. Wilson1,§ 1Department of Cell Biology, The Johns Hopkins University School of Medicine, 725 N. Wolfe St, Baltimore, MD 21205, USA 2Department of Pathology, Emory University, Whitehead Biomedical Research Building, Atlanta, GA 30332, USA *Present address: Department of Developmental Biology, Stanford University School of Medicine, Beckman Center, B300, 279 Campus Drive, Stanford, CA 94305, USA ‡Department of Biology, Eckerd College, SHB 105, 4200 54th Ave, St Petersburg, FL 33711, USA §Author for correspondence (e-mail: [email protected]) Accepted 4 October 2005 Journal of Cell Science 119, 239-249 Published by The Company of Biologists 2006 doi:10.1242/jcs.02728 Summary Lamins form structural filaments in the nucleus. Mutations lamin-downregulated [lmn-1(RNAi)] embryos, Ce-titin was in A-type lamins cause muscular dystrophy, undetectable at the nuclear envelope suggesting its cardiomyopathy and other diseases, including progeroid localization or stability requires Ce-lamin. In human cells syndromes. To identify new binding partners for lamin A, (HeLa), antibodies against the titin-specific domain M-is6 we carried out a two-hybrid screen with a human skeletal- gave both diffuse and punctate intranuclear staining by muscle cDNA library, using the Ig-fold domain of lamin A indirect immunofluorescence, and recognized at least three as bait. The C-terminal region of titin was recovered twice. bands larger than 1 MDa in immunoblots of isolated HeLa Previous investigators showed that nuclear isoforms of titin nuclei. -

Lamin A/C Cardiomyopathy: Implications for Treatment
Current Cardiology Reports (2019) 21:160 https://doi.org/10.1007/s11886-019-1224-7 MYOCARDIAL DISEASE (A ABBATE AND G SINAGRA, SECTION EDITORS) Lamin A/C Cardiomyopathy: Implications for Treatment Suet Nee Chen1 & Orfeo Sbaizero1,2 & Matthew R. G. Taylor1 & Luisa Mestroni1 # Springer Science+Business Media, LLC, part of Springer Nature 2019 Abstract Purpose of Review The purpose of this review is to provide an update on lamin A/C (LMNA)-related cardiomyopathy and discuss the current recommendations and progress in the management of this disease. LMNA-related cardiomyopathy, an inherited autosomal dominant disease, is one of the most common causes of dilated cardiomyopathy and is characterized by steady progression toward heart failure and high risks of arrhythmias and sudden cardiac death. Recent Findings We discuss recent advances in the understanding of the molecular mechanisms of the disease including altered cell biomechanics, which may represent novel therapeutic targets to advance the current management approach, which relies on standard heart failure recommendations. Future therapeutic approaches include repurposed molecularly directed drugs, siRNA- based gene silencing, and genome editing. Summary LMNA-related cardiomyopathy is the focus of active in vitro and in vivo research, which is expected to generate novel biomarkers and identify new therapeutic targets. LMNA-related cardiomyopathy trials are currently underway. Keywords Lamin A/C gene . Laminopathy . Heart failure . Arrhythmias . Mechanotransduction . P53 . CRISPR–Cas9 therapy Introduction functions, including maintaining nuclear structural integrity, regulating gene expression, mechanosensing, and Mutations in the lamin A/C gene (LMNA)causelaminopathies, mechanotransduction through the lamina-associated proteins a heterogeneous group of inherited disorders including muscu- [6–11]. -

Hippocampal LMNA Gene Expression Is Increased in Late-Stage Alzheimer’S Disease
International Journal of Molecular Sciences Article Hippocampal LMNA Gene Expression is Increased in Late-Stage Alzheimer’s Disease Iván Méndez-López 1,2,*,†, Idoia Blanco-Luquin 1,†, Javier Sánchez-Ruiz de Gordoa 1,3, Amaya Urdánoz-Casado 1, Miren Roldán 1, Blanca Acha 1, Carmen Echavarri 1,4, Victoria Zelaya 5, Ivonne Jericó 3 and Maite Mendioroz 1,3,* 1 Neuroepigenetics Laboratory-Navarrabiomed, Complejo Hospitalario de Navarra, Universidad Pública de Navarra (UPNA), IdiSNA (Navarra Institute for Health Research), Pamplona, Navarra 31008, Spain; [email protected] (I.B.-L.); [email protected] (J.S.-R.d.G.); [email protected] (A.U.-C.); [email protected] (M.R.); [email protected] (B.A.); [email protected] (C.E.) 2 Department of Internal Medicine, Hospital García Orcoyen, Estella 31200, Spain 3 Department of Neurology, Complejo Hospitalario de Navarra- IdiSNA (Navarra Institute for Health Research), Pamplona, Navarra 31008, Spain; [email protected] 4 Hospital Psicogeriátrico Josefina Arregui, Alsasua, Navarra 31800, Spain 5 Department of Pathology, Complejo Hospitalario de Navarra- IdiSNA (Navarra Institute for Health Research), Pamplona, Navarra 31008, Spain; [email protected] * Correspondence: [email protected] (I.M.-L.); [email protected] (M.M.); Tel.: +34-848-422677 (I.M.-L.) † These authors contributed equally to this work. Received: 30 November 2018; Accepted: 14 February 2019; Published: 18 February 2019 Abstract: Lamins are fibrillary proteins that are crucial in maintaining nuclear shape and function. Recently, B-type lamin dysfunction has been linked to tauopathies. However, the role of A-type lamin in neurodegeneration is still obscure. -

Deficiencies in Lamin B1 and Lamin B2 Cause Neurodevelopmental Defects and Distinct Nuclear Shape Abnormalities in Neurons
M BoC | ARTICLE Deficiencies in lamin B1 and lamin B2 cause neurodevelopmental defects and distinct nuclear shape abnormalities in neurons Catherine Coffiniera, Hea-Jin Jungb, Chika Nobumoria, Sandy Changa, Yiping Tua, Richard H. Barnes IIa, Yuko Yoshinagac, Pieter J. de Jongc, Laurent Vergnesd, Karen Reued, Loren G. Fonga, and Stephen G. Younga,d aDepartment of Medicine and bMolecular Biology Institute, University of California, Los Angeles, Los Angeles, CA 90095; cChildren’s Hospital Oakland Research Institute, Oakland, CA 94609; dDepartment of Human Genetics, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, CA 90095 ABSTRACT Neuronal migration is essential for the development of the mammalian brain. Monitoring Editor Here, we document severe defects in neuronal migration and reduced numbers of neurons in Thomas Michael Magin lamin B1–deficient mice. Lamin B1 deficiency resulted in striking abnormalities in the nuclear University of Leipzig shape of cortical neurons; many neurons contained a solitary nuclear bleb and exhibited an Received: Jun 13, 2011 asymmetric distribution of lamin B2. In contrast, lamin B2 deficiency led to increased numbers Revised: Sep 9, 2011 of neurons with elongated nuclei. We used conditional alleles for Lmnb1 and Lmnb2 to create Accepted: Sep 23, 2011 forebrain-specific knockout mice. The forebrain-specificLmnb1- and Lmnb2-knockout models had a small forebrain with disorganized layering of neurons and nuclear shape abnormalities, similar to abnormalities identified in the conventional knockout mice. A more severe pheno- type, complete atrophy of the cortex, was observed in forebrain-specific Lmnb1/Lmnb2 double-knockout mice. This study demonstrates that both lamin B1 and lamin B2 are essential for brain development, with lamin B1 being required for the integrity of the nuclear lamina, and lamin B2 being important for resistance to nuclear elongation in neurons. -
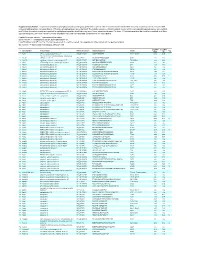
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Mechanosensing Defects and YAP-Signaling in LMNA-Related Congenital Muscular Dystrophy
Mechanosensing defects and YAP-signaling in LMNA-related congenital muscular dystrophy Dissertation in fulfillment of the requirements for the Joint Degree (Cotutelle) “Doctor rerum naturalium (Dr. rer. nat.)” integrated in the International Graduate School for Myology “MyoGrad” in the Department of Biology, Chemistry, Pharmacy at Freie Universität Berlin and in Cotutelle Agreement with the Ecole Doctorale 515 “Complexité du Vivant” at the Université Pierre et Marie Curie Paris 6 Submitted by Martina Fischer Supervised by Catherine Coirault and Prof. Dr. Petra Knaus Thesis defense: June 28th, 2017 Thesis jury: Thesis supervisor, UPMC: Catherine Coirault, PhD Thesis supervisor, first reviewer of Freie Universität Berlin: Prof. Dr. Petra Knaus Reviewer UPMC: Jean-Thomas Vilquin, PhD Second reviewer, Freie Universität Berlin: Prof. Dr. Sigmar Stricker External Expert: Prof. Dr. Henning Wackerhage External Expert: Prof. Dr. Michael Gotthardt External Expert: Prof. Dr. Markus Schülke-Gerstenfeld Postdoctoral Fellow, Freie Universität Berlin: Dr. Maria Reichenbach ACKNOWLEDGEMENTS Firstly, I would like to express my sincere gratitude to my advisors Catherine Coirault and Prof. Dr. Petra Knaus for the continuous support of my PhD study, for their example, effort and knowledge. Their guidance was essential in all the time of my research and writing. Besides my advisors, I would like to thank the members of my former thesis committee and todays defense committee: Prof. Dr. Sigmar Stricker, Prof. Dr. Michael Gotthardt, Prof. Dr. Henning Wackerhage, Jean-Thomas Vilquin, Dr. Maria Reichenbach, Danièle Lacasa, Athanassia Sotiropoulos, Sigolène Meilhac and Dr. Christian Hiepen for their insightful comments and support. I also own thanks to the founders and coordinators of MyoGrad, especially Gisèle Bonne and Simone Spuler for creating this fruitful environment. -

Kif1b Rab7a Lmna
Title: Charcot-Marie-Tooth Neuropathy Type 2 GeneReview Molecular Genetics: Less Commonly Involved Genes Author: Bird TD Updated: March 2016 KIF1B Gene structure. KIF1B comprises 47 exons and 167.13 kb of DNA. Pathogenic allelic variants. See Table A, Locus Specific and HGMD Normal gene product. Kinesin-like protein KIF1B is involved in axonal transport of synaptic vesicle precursors [Zhao et al 2001]. The kinesin superfamily of proteins is essential for intracellular transport along microtubules. Abnormal gene product. There may be a defect in the transport of synaptic vesicles. RAB7A Gene structure. RAB7A has six exons and 87.9 kb of DNA. Pathogenic allelic variants. See Table A. Normal gene product. Ras-related protein Rab-7a belongs to the RAB family of Ras- related GTPases essential for the regulation of intracellular membrane trafficking. Rab- 7a is involved in transport between late endosomes and lysosomes. RAB-interacting lysosomal protein (RILP) induces the recruitment of dynein-dynactin motors and regulates transport toward the minus-end of microtubules [Verhoeven et al 2003]. Abnormal gene product. Abnormal Rab-7a may cause malfunction of lysosomes and inhibit neurite outgrowth [Spinosa et al 2008, Bucci & Deluca 2012]. LMNA Gene structure. LMNA has 12 exons spread over 24 kb of genomic DNA. Pathogenic allelic variants. The most common pathogenic variant found in individuals with CMT2B1 is p.Arg298Cys, a founder mutation in North Africa [Bouhouche et al 2007, De Sandre-Giovannoli et al 2002]. See also Table A. Table 5. Selected LMNA Variants DNA Nucleotide Protein Amino Acid Class of Variant Allele Reference Sequences Change Change Benign c.1908C>T p.= 1 c.398G>T p.Arg133Leu NM_170707.2 c.892C>T p.Arg298Cys Pathogenic NP_733821.1 c.1411C>T p.Arg471Cys c.1579C>T p.Arg527Cys Note on variant classification: Variants listed in the table have been provided by the author. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

LMNA-Related Disorders
LMNA-Related Disorders Indications for Ordering Genetics To confirm a clinical diagnosis of a LMNA-related disorder, Genes: LMNA such as: • Hutchinson-Gilford progeria syndrome (HGPS) Inheritance: See table • Emery-Dreifuss muscular dystrophy type 2 (EDMD2) Penetrance: Varies by syndrome • Limb-Girdle muscular dystrophy 1B (LGMD1B) • Charcot-Marie-Tooth 2B1 (CMT2B1) Structure/Function • Familial partial lipodystrophy, Dunnigan type (FPLD) • Composed of 12 exons • LMNA-related dilated cardiomyopathy (DCM) • LMNA encodes isoforms A and C of the lamin protein • Mandibulo-acral dysplasia (MAD) o Structural component of the nuclear membrane • Atypical Werner syndrome (WS) o Anchors heterochromatin to the inner nuclear • Restrictive dermopathy (RD) membrane • Other, intermediate phenotypes Variants Test Description • Alternative splicing of the LMNA gene produces two proteins (lamin A and C) Polymerase chain reaction (PCR) followed by bidirectional • Variants occur throughout the gene sequencing of all coding regions and intron/exon o Predominantly missense boundaries of the LMNA gene o p.G608G variant in exon 11 . Present in all individuals with HGPS Tests to Consider Test Interpretation Primary Tests LMNA-Related Disorders (LMNA) Sequencing 2004543 Sensitivity/Specificity • Confirm suspected laminopathy caused by LMNA variants, • Clinical sensitivity: dependent on the specific LMNA- including HGPS, EDMD2, LGMD1B, CMT2B1, FPLD , DCM, related disorder MAD, WS, or RD • Analytic sensitivity/specificity: 99% Related Test Results Familial Mutation,