Identifying, Characterizing, and Modulating Regulatory Elements in Their Natural Context
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Diagnosis of Sickle Cell Disease and HBB Haplotyping in the Era of Personalized Medicine: Role of Next Generation Sequencing
Journal of Personalized Medicine Article Diagnosis of Sickle Cell Disease and HBB Haplotyping in the Era of Personalized Medicine: Role of Next Generation Sequencing Adekunle Adekile 1,*, Nagihan Akbulut-Jeradi 2, Rasha Al Khaldi 2, Maria Jinky Fernandez 2 and Jalaja Sukumaran 1 1 Department of Pediatrics, Faculty of Medicine, Kuwait University, P.O. Box 24923, Safat 13110, Kuwait; jalajasukumaran@hotmail 2 Advanced Technology Company, Hawali 32060, Kuwait; [email protected] (N.A.-J.); [email protected] (R.A.); [email protected] (M.J.F.) * Correspondence: [email protected]; Tel.: +965-253-194-86 Abstract: Hemoglobin genotype and HBB haplotype are established genetic factors that modify the clinical phenotype in sickle cell disease (SCD). Current methods of establishing these two factors are cumbersome and/or prone to errors. The throughput capability of next generation sequencing (NGS) makes it ideal for simultaneous interrogation of the many genes of interest in SCD. This study was designed to confirm the diagnosis in patients with HbSS and Sβ-thalassemia, identify any ß-thal mutations and simultaneously determine the ßS HBB haplotype. Illumina Ampliseq custom DNA panel was used to genotype the DNA samples. Haplotyping was based on the alleles on five haplotype-specific SNPs. The patients studied included 159 HbSS patients and 68 Sβ-thal patients, previously diagnosed using high performance liquid chromatography (HPLC). There was Citation: Adekile, A.; considerable discordance between HPLC and NGS results, giving a false +ve rate of 20.5% with a S Akbulut-Jeradi, N.; Al Khaldi, R.; sensitivity of 79% for the identification of Sβthal. -

Adult, Embryonic and Fetal Hemoglobin Are Expressed in Human Glioblastoma Cells
514 INTERNATIONAL JOURNAL OF ONCOLOGY 44: 514-520, 2014 Adult, embryonic and fetal hemoglobin are expressed in human glioblastoma cells MARWAN EMARA1,2, A. ROBERT TURNER1 and JOAN ALLALUNIS-TURNER1 1Department of Oncology, University of Alberta and Alberta Health Services, Cross Cancer Institute, Edmonton, AB T6G 1Z2, Canada; 2Center for Aging and Associated Diseases, Zewail City of Science and Technology, Cairo, Egypt Received September 7, 2013; Accepted October 7, 2013 DOI: 10.3892/ijo.2013.2186 Abstract. Hemoglobin is a hemoprotein, produced mainly in Introduction erythrocytes circulating in the blood. However, non-erythroid hemoglobins have been previously reported in other cell Globins are hemo-containing proteins, have the ability to types including human and rodent neurons of embryonic bind gaseous ligands [oxygen (O2), nitric oxide (NO) and and adult brain, but not astrocytes and oligodendrocytes. carbon monoxide (CO)] reversibly. They have been described Human glioblastoma multiforme (GBM) is the most aggres- in prokaryotes, fungi, plants and animals with an enormous sive tumor among gliomas. However, despite extensive basic diversity of structure and function (1). To date, hemoglobin, and clinical research studies on GBM cells, little is known myoglobin, neuroglobin (Ngb) and cytoglobin (Cygb) repre- about glial defence mechanisms that allow these cells to sent the vertebrate globin family with distinct function and survive and resist various types of treatment. We have tissue distributions (2). During ontogeny, developing erythro- shown previously that the newest members of vertebrate blasts sequentially express embryonic {[Gower 1 (ζ2ε2), globin family, neuroglobin (Ngb) and cytoglobin (Cygb), are Gower 2 (α2ε2), and Portland 1 (ζ2γ2)] to fetal [Hb F(α2γ2)] expressed in human GBM cells. -
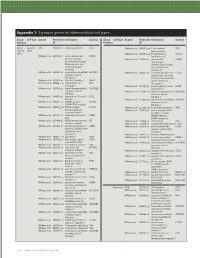
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

The Alexipharmic Mechanisms of Five Licorice Ingredients Involved in CYP450 and Nrf2 Pathways in Paraquat-Induced Mice Acute Lung Injury
Hindawi Oxidative Medicine and Cellular Longevity Volume 2019, Article ID 7283104, 20 pages https://doi.org/10.1155/2019/7283104 Research Article The Alexipharmic Mechanisms of Five Licorice Ingredients Involved in CYP450 and Nrf2 Pathways in Paraquat-Induced Mice Acute Lung Injury Zi-Jing Liu,1,2 Jing Zhong,1 Mei Zhang,1 Ze-Hui Chen,1 Ji-Ye Wang,1 Han-Ying Chen,1,2 Xiao-Qin Wang,1,2 and Bo Zhang 1,2 1Key Laboratory of Xinjiang Endemic Phytomedicine Resources, Ministry of Education, School of Pharmacy, Shihezi University, Shihezi 832002, China 2Pharmacology Department, School of Pharmacy, Shihezi University, Shihezi 832002, China Correspondence should be addressed to Bo Zhang; [email protected] Received 21 May 2018; Revised 30 September 2018; Accepted 3 December 2018; Published 28 April 2019 Academic Editor: Pablo Muriel Copyright © 2019 Zi-Jing Liu et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Oxidative stress is an important mechanism in acute lung injury (ALI) induced by paraquat (PQ), one of the most widely used herbicides in developing countries. In clinical prophylaxis and treatment, licorice is a widely used herbal medicine in China due to its strong alexipharmic characteristics. However, the corresponding biochemical mechanism of antioxidation and detoxification enzymes induced by licorice’s ingredients is still not fully demonstrated. In this study, the detoxification effect of licorice was evaluated in vivo and in vitro. The detoxification and antioxidation effect of its active ingredients involved in the treatment was screened systematically according to Absorption, Distribution, Metabolism, and Excretion (ADME): predictions and evidence-based literature mining methods in silico approach. -

Anti-Hemoglobin Antibody (FITC) Product Number: AC15-0147-12
Anti-Hemoglobin Antibody (FITC) Product Number: AC15-0147-12 Overview Host Species: Goat Clonality: Polyclonal Purity: Hemoglobin Antibody (FITC) is affinity purified. The affinity purified antibody is then conjugated to the fluorescent dye FITC (fluorescein isothiocyanate). Conjugate: FITC Immunogen Type: Anti-hemoglobin antibody (FITC) was rasied against human hemoglobin. Physical Characteristics Amount: 0.05 mg Concentration: 1 mg/ml Volume: 0.05 ml Buffer: 10 mM Sodium Phosphate, 0.15 M NaCl, pH 7.2 with 0.05% (w/v) sodium azide. Storage: FITC conjugated antibody can be stored at 4?C for up to 18 months. For longer storage the conjugate can be stored at -20?C after adding 50% glycerol. Fluorescein conjugated antibodies should always be stored in the dark. Specificity Species Reactivity: Human Specificity: Human hemoglobin Target Information Gene ID Number(s): 3047 (human), 3039 (human), 3040 (human) Alternative Names: 3-prime alpha-globin gene; Alpha globin; alpha one globin; alpha-1 globin; Alpha-globin; Beta globin; CD113t C; CD31antibody (FITC); Erythremia, beta-globin type, included; Gamma 1 globin; Hb FAgamma; HBA 1; HBA 2; HBA; HBA_HUMAN; HBA1antibody (FITC); HBA2; HBB; Hbb-y; HBD; Hbe1; HBG 1antibody (FITC); HBG; HBG1; HBGA; HBGR; HBHantibody (FITC); Hemoglobin alpha 1; hemoglobin alpha 1 globin chain; Hemoglobinalpha chain; Hemoglobin alpha locus; Hemoglobin alpha locus 1antibody (FITC); hemoglobin alpha-1 chain; Hemoglobin beta; Hemoglobin beta chainantibody (FITC); Hemoglobin beta chain complex; Hemoglobin beta locus; Hemoglobingamma -

The Normal Structure and Regulation of Human Globin Gene Clusters
CHAPTER 3 The Normal Structure and Regulation of Human Globin Gene Clusters Bernard G. Forget and Ross C. Hardison The genes encoding the different globin chains of hemoglobin are members of an ancient gene family. In this chapter we will review the structural features of the globin genes, with particular attention to the sequences needed for proper regulation of gene expression. Some of these have been well- conserved during mammalian evolution and therefore are likely to provide a common function in many mammals. Others are only found in higher primates, and may play roles in lineage-specific regulation. We will first describe the structural characteristics of the human globin genes and then provide a comparative analysis of the genomic contexts, regulatory regions and evolutionary conservation of features present in the globin gene clusters. NUMBER AND CHROMOSOMAL LOCALIZATION OF HUMAN GLOBIN GENES Hemoglobin is a heterotetramer that contains two polypeptide subunits related to the α-globin gene subfamily (referred to here as α-like globins) and two polypeptide subunits related to the β-globin gene subfamily (β-like globins). Globin polypeptides bind heme, which in turn allows the hemoglobin in erythrocytes to bind oxygen reversibly and transport it from the lungs to respiring tissues. In humans, as in all vertebrate species studied, different α-like and β-like globin chains are synthesized at Chapter 3 The Normal Structure and Regulation of the Globin Gene Clusters progressive stages of development to produce hemoglobins characteristic of primitive (embryonic) and definitive (fetal and adult) erythroid cells (Figure 3.1). Before precise knowledge of globin gene organization was gained by gene mapping and molecular cloning, a general picture of the number and arrangement of the human globin genes emerged from the genetic analysis of normal and abnormal hemoglobins and their pattern of inheritance. -

Evolution of Hemoglobin and Its Genes
Downloaded from http://perspectivesinmedicine.cshlp.org/ on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Evolution of Hemoglobin and Its Genes Ross C. Hardison Center for Comparative Genomics and Bioinformatics, Huck Institute of Genome Sciences, Department of Biochemistry and Molecular Biology, The Pennsylvania State University, University Park, Pennsylvania 16802 Correspondence: [email protected] Insights into the evolution of hemoglobins and their genes are an abundant source of ideas regarding hemoglobin function and regulation of globin gene expression. This article pre- sents the multiple genes and gene families encoding human globins, summarizes major events in the evolution of the hemoglobin gene clusters, and discusses how these studies provide insights into regulation of globin genes. Although the genes in and around the a-like globin gene complex are relatively stable, the b-like globin gene clusters are more dynamic, showing evidence of transposition to a new locus and frequent lineage-specific expansions and deletions. The cis-regulatory modules controlling levels and timing of gene expression are a mix of conserved and lineage-specific DNA, perhaps reflecting evolutionary constraint on core regulatory functions shared broadly in mammals and adaptive fine-tuning in different orders of mammals. wide range of animals, vertebrate and inver- globin genes, and attempts to modulate globin Atebrate, use hemoglobins to transport oxy- gene expression are a fundamental approach to gen, carrying it from lungs, gills, or -

Gene Expression Profiling of Peripheral Blood in Patients With
FLORE Repository istituzionale dell'Università degli Studi di Firenze Gene expression profiling of peripheral blood in patients with abdominal aortic aneurysm Questa è la Versione finale referata (Post print/Accepted manuscript) della seguente pubblicazione: Original Citation: Gene expression profiling of peripheral blood in patients with abdominal aortic aneurysm / Giusti B; Rossi L; Lapini I; Magi A; Pratesi G; Lavitrano M; Blasi GM; Pulli R; Pratesi C; Abbate R. - In: EUROPEAN JOURNAL OF VASCULAR AND ENDOVASCULAR SURGERY. - ISSN 1078-5884. - STAMPA. - 38(2009), pp. 104-112. [10.1016/j.ejvs.2009.01.020] Availability: This version is available at: 2158/369023 since: 2018-03-01T22:38:47Z Published version: DOI: 10.1016/j.ejvs.2009.01.020 Terms of use: Open Access La pubblicazione è resa disponibile sotto le norme e i termini della licenza di deposito, secondo quanto stabilito dalla Policy per l'accesso aperto dell'Università degli Studi di Firenze (https://www.sba.unifi.it/upload/policy-oa-2016-1.pdf) Publisher copyright claim: (Article begins on next page) 24 September 2021 Eur J Vasc Endovasc Surg (2009) 38, 104e112 Gene Expression Profiling of Peripheral Blood in Patients with Abdominal Aortic Aneurysm B. Giusti a,*, L. Rossi a, I. Lapini a, A. Magi a, G. Pratesi b, M. Lavitrano c, G.M. Biasi c, R. Pulli d, C. Pratesi d, R. Abbate a a Department of Medical and Surgical Critical Care and DENOTHE Center, University of Florence, Viale Morgagni 85, 50134 Florence, Italy b Vascular Surgery Unit, Department of Surgery, University of Rome ‘‘Tor Vergata’’, Rome, Italy c Department of Surgical Sciences, University of Milano-Bicocca, Italy d Department of Vascular Surgery, University of Florence, Italy Submitted 17 September 2008; accepted 15 January 2009 Available online 23 February 2009 KEYWORDS Abstract Object: Abdominal aortic aneurysm (AAA) pathogenesis remains poorly understood. -

1 Evolution of Hemoglobin Loci and Their Regulatory Elements Sjaak
Evolution of hemoglobin loci and their regulatory elements Sjaak Philipsen1 and Ross C. Hardison2 1 Department of Cell Biology Ee1071b, Erasmus MC, P.O. Box 2040, 3000 CA Rotterdam, The Netherlands. Email [email protected] Tel. +31-10-7044282 2 Department of Biochemistry and Molecular Biology, Huck Institute for Comparative Genomics and Bioinformatics, The Pennsylvania State University, University Park, PA 16802, USA. Email [email protected] Tel. +1-814-8630113 Key words: transcriptional regulation; globin genes; locus control regions; super-enhancers; evolution Word count Abstract: 178 Review: 5793 This article was accepted to Blood Cells, Molecules and Diseases in August 2017. The DOI is doi.org/10.1016/j.bcmd.2017.08.001 The link to the article at BCMD is https://doi.org/10.1016/j.bcmd.2017.08.001 1 Abstract Across the expanse of vertebrate evolution, each species produces multiple forms of hemoglobin in erythroid cells at appropriate times and in the proper amounts. The multiple hemoglobins are encoded in two globin gene clusters in almost all species. One globin gene cluster, linked to the gene NPRL3, is preserved in all vertebrates, including a gene cluster encoding the highly divergent globins from jawless vertebrates. This preservation of synteny may reflect the presence of a powerful enhancer of globin gene expression in the NPRL3 gene. Despite substantial divergence in noncoding DNA sequences among mammals, several epigenetic features of the globin gene regulatory regions are preserved across vertebrates. The preserved features include multiple DNase hypersensitive sites, at least one of which is an enhancer, and binding by key lineage-restricted transcription factors such as GATA1 and TAL1, which in turn recruit coactivators such as P300 that catalyze acetylation of histones. -

A Benchmark of Hemoglobin Blocking During Library Preparation for Mrna
www.nature.com/scientificreports OPEN A benchmark of hemoglobin blocking during library preparation for mRNA-Sequencing of human blood samples Florian Uellendahl-Werth1, Markus Wolfen2, Andre Franke1, Olaf Wolkenhauer2,3 & David Ellinghaus1* RNA-Sequencing (RNA-Seq) of peripheral blood can be a valuable source of information for investigating the status and mechanism of diseases. However, blood contains 50–80% unwanted hemoglobin (Hb) transcripts. Lexogen’s QuantSeq mRNA-Seq-Kit for Illumina RNA-Seq features a ‘Globin Block’ (GB) module that depletes Hb cDNAs during library preparation. Here, we aimed to assess GB’s efectiveness and checked for technical biases attributable to GB. Using whole blood total RNA samples of 91 healthy individuals, we sequenced 91 pairs of GB and non-blocked samples (noGB) on Illumina HiSeq2500 and 8 pairs of GB/noGB technical replicates on HiSeq4000. GB reduced the fraction of Hb transcripts from 43% (s.d. 14%) to 8.0% (s.d. 4.3%). From GB samples we detected 1,397 more expressed genes at approximately 11 million reads per RNA-isolate. Enrichment and diferential expression analyses did not reveal signifcant diferences for GB and noGB samples with respect to molecular function. In contrast to results from studies that have examined the performance of GB during RNA isolation, we were able to assign GB to corresponding noGB samples (from multiple sequencing runs on HiSeq2500) with at least 89.8% accuracy from the complete correlation matrix of all GB/GB, noGB/noGB and GB/noGB pairs. However, the use of diferent sequencers (HiSeq2500 vs HiSeq4000) impaired assignment of technical replicates, whereas assignment of GB to corresponding noGB samples worked perfectly when sequencing on one lane on HiSeq4000. -

BET Inhibitors Enhance Embryonic and Fetal Globin Expression in Erythroleukemia Cell Lines by John Z
BET inhibitors enhance embryonic and fetal globin expression in erythroleukemia cell lines by John Z. Cao, Kristina Bigelow, Amittha Wickrema, and Lucy A. Godley Received: March 16, 2021. Accepted: August 20, 2021. Citation: John Z. Cao, Kristina Bigelow, Amittha Wickrema, and Lucy A. Godley. BET inhibitors enhance embryonic and fetal globin expression in erythroleukemia cell lines. Haematologica. 2021 Aug 26. doi: 10.3324/haematol.2021.278791. [Epub ahead of print] Publisher's Disclaimer. E-publishing ahead of print is increasingly important for the rapid dissemination of science. Haematologica is, therefore, E-publishing PDF files of an early version of manuscripts that have completed a regular peer review and have been accepted for publication. E-publishing of this PDF file has been approved by the authors. After having E-published Ahead of Print, manuscripts will then undergo technical and English editing, typesetting, proof correction and be presented for the authors' final approval; the final version of the manuscript will then appear in a regular issue of the journal. All legal disclaimers that apply to the journal also pertain to this production process. BET inhibitors enhance embryonic and fetal globin expression in erythroleukemia cell lines John Z. Cao1, Kristina Bigelow2, Amittha Wickrema1,2, and Lucy A. Godley1,2* 1 Committee on Cancer Biology, Biological Sciences Division, The University of Chicago, Chicago IL 2 Section of Hematology/Oncology, Department of Medicine, The University of Chicago, Chicago IL *Corresponding author Contact information: 5841 S. Maryland Ave. MC 2115 Chicago, IL 60637 Phone: 773-702-4140 FAX: 773-702-9268 Email: [email protected] Author Contribution J.Z.C. -

Elabscience.Com ® E-Mail:[email protected] Elabscience Elabscience Biotechnology Inc
Tel:240-252-7368(USA) Fax:240-252-7376(USA) www.elabscience.com ® E-mail:[email protected] Elabscience Elabscience Biotechnology Inc. HBE1 Polyclonal Antibody Catalog No. E-AB-18898 Reactivity H Storage Store at -20℃. Avoid freeze / thaw cycles. Host Rabbit Applications WB,IHC,ELISA Isotype IgG Note: Centrifuge before opening to ensure complete recovery of vial contents. Images Immunogen Information Immunogen Fusion protein of human HBE1 Gene Accession BC015537 Swissprot P02100 Synonyms Epsilon-globin,HBE,HBE1,Hemoglobin epsilon chain,Hemoglobin subunit epsilon Western blot analysis of Human Product Information placenta tissue lysate using HBE1 Calculated MW 16 kDa Polyclonal Antibody at dilution of Observed MW Refer to figures 1:300 Buffer PBS with 0.05% NaN3 and 40% Glycerol,pH7.4 Purify Antigen affinity purification Dilution WB 1:500-1:2000, IHC 1:30-1:150, ELISA 1:5000-1:10000 Background The hemoglobin molecule is a tetramer consisting of two α-globin-like polypeptide chains and two β-globin-like chains. The human hemoglobin Immunohistochemistry of paraffin- genes are expressed in a tightly developmentally controlled fashion.ε- embedded Human tonsil tissue using globin (HBE1) is the predominantly expressed gene during the embryonic HBE1 Polyclonal Antibody at dilution stage. The epsilon hemoglobin chain seems to be the best marker for fetal of 1:30(×200) nucleated red blood cells (NRBCs). Anti-HBE1 may be used to label and isolate fetal cells from maternal blood and can be useful in prenatal diagnosis. This antibody specifically recognizes the HBE1 and doesn't cross-react with other globin chains. For Research Use Only Focus on your research Thank you for your recent purchase.