Aldehyde Dehydrogenase (Ec 1.2.1.5)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Aldehyde Dehydrogenase ALDH2*2 Allele Exhibits Dominance Over ALDH2*1 in Transduced Hela Cells
The aldehyde dehydrogenase ALDH2*2 allele exhibits dominance over ALDH2*1 in transduced HeLa cells. Q Xiao, … , T Johnston, D W Crabb J Clin Invest. 1995;96(5):2180-2186. https://doi.org/10.1172/JCI118272. Research Article Individuals heterozygous or homozygous for the variant aldehyde dehydrogenase (ALDH2) allele (ALDH2*2), which encodes a protein differing only at residue 487 from the normal protein, have decreased ALDH2 activity in liver extracts and experience cutaneous flushing when they drink alcohol. The mechanisms by which this allele exerts its dominant effect is unknown. To study this effect, the human ALDH2*1 cDNA was cloned and the ALDH2*2 allele was generated by site-directed mutagenesis. These cDNAs were transduced using retroviral vectors into HeLa and CV1 cells, which do not express ALDH2. The normal allele directed synthesis of immunoreactive ALDH2 protein (ALDH2E) with the expected isoelectric point. Extracts of these cells contained increased aldehyde dehydrogenase activity with low Km for the aldehyde substrate. The ALDH2*2 allele directed synthesis of mRNA and immunoreactive protein (ALDH2K), but the protein lacked enzymatic activity. When ALDH2*1-expressing cells were transduced with ALDH2*2 vectors, both mRNAs were expressed and immunoreactive proteins with isoelectric points ranging between those of ALDH2E and ALDH2K were present, indicating that the subunits formed heteromers. ALDH2 activity in these cells was reduced below that of the parental ALDH2*1-expressing cells. Thus, the ALDH2*2 allele is sufficient to cause ALDH2 deficiency in vitro. Find the latest version: https://jci.me/118272/pdf The Aldehyde Dehydrogenase ALDH2*2 Allele Exhibits Dominance over ALDH2*1 in Transduced HeLa Cells Qing Xiao, * Henry Weiner,* Timothy Johnston,* and David W. -

Isocitrate Dehydrogenase Activity Assay Kit (MAK062)
Isocitrate Dehydrogenase Activity Assay Kit Catalog Number MAK062 Storage Temperature –20 C TECHNICAL BULLETIN Product Description Developer 1 vl Isocitrate dehydrogenase (IDH) catalyzes the Catalog Number MAK062E conversion of isocitrate to -ketoglutarate. In eukaryotes, there are three isozymes of IDH, the IDH Positive Control (NADP+) 20 L mitochondrial IDH2 and IDH3, and the cytoplasmic/ Catalog Number MAK062F peroxisomal IDH1. All three IDH family members require the presence of a divalent cation (Mg2+ or Mn2+) NADH Standard, 0.5 mole 1 vl and either the electron-accepting cofactor NADP+ (IDH1 Catalog Number MAK062G and IDH2) or NAD+ (IDH3) for their enzymatic activity. IDH1 and IDH2 mutations resulting in neomorphic Reagents and Equipment Required but Not enzymatic activity are found in certain cancers such as Provided. glioblastoma, acute myeloid leukemia, and colon 96 well flat-bottom plate – It is recommended to use cancer. This neoactivity shows a change in the clear plates for colorimetric assays. substrate specificity resulting in the conversion of Spectrophotometric multiwell plate reader -ketoglutarate to 2-hydroxyglutarate. Mutations in IDH family members are also associated with Ollier disease Precautions and Disclaimer and Maffucci syndrome. This product is for R&D use only, not for drug, household, or other uses. Please consult the Material The Isocitrate Dehydrogenase Activity Assay kit Safety Data Sheet for information regarding hazards provides a simple and direct procedure for measuring and safe handling practices. + + + NADP -dependent, NAD -dependent, or both NADP + and NAD -dependent IDH activity in a variety of Preparation Instructions samples. IDH activity is determined using isocitrate as Briefly centrifuge vials before opening. -

Isocitrate Dehydrogenase 1 (NADP+) (I5036)
Isocitrate Dehydrogenase 1 (NADP+), human recombinant, expressed in Escherichia coli Catalog Number I5036 Storage Temperature –20 °C CAS RN 9028-48-2 IDH1 and IDH2 have frequent genetic alterations in EC 1.1.1.42 acute myeloid leukemia4 and better understanding of Systematic name: Isocitrate:NADP+ oxidoreductase these mutations may lead to an improvement of (decarboxylating) individual cancer risk assessment.6 In addition other studies have shown loss of IDH1 in bladder cancer Synonyms: IDH1, cytosolic NADP(+)-dependent patients during tumor development suggesting this may isocitrate dehydrogenase, isocitrate:NADP+ be involved in tumor progression and metastasis.7 oxidoreductase (decarboxylating), Isocitric Dehydrogenase, ICD1, PICD, IDPC, ICDC, This product is lyophilized from a solution containing oxalosuccinate decarboxylase Tris-HCl, pH 8.0, with trehalose, ammonium sulfate, and DTT. Product Description Isocitrate dehydrogenase (NADP+) [EC 1.1.1.42] is a Purity: ³90% (SDS-PAGE) Krebs cycle enzyme, which converts isocitrate to a-ketoglutarate. The flow of isocitrate through the Specific activity: ³80 units/mg protein glyoxylate bypass is regulated by phosphorylation of isocitrate dehydrogenase, which competes for a Unit definition: 1 unit corresponds to the amount of 1 common substrate (isocitrate) with isocitrate lyase. enzyme, which converts 1 mmole of DL-isocitrate to The activity of the enzyme is dependent on the a-ketoglutarate per minute at pH 7.4 and 37 °C (NADP formation of a magnesium or manganese-isocitrate as cofactor). The activity is measured by observing the 2 complex. reduction of NADP to NADPH at 340 nm in the 7 presence of 4 mM DL-isocitrate and 2 mM MnSO4. -

(LCHAD) Deficiency / Mitochondrial Trifunctional Protein (MTF) Deficiency
Long chain acyl-CoA dehydrogenase (LCHAD) deficiency / Mitochondrial trifunctional protein (MTF) deficiency Contact details Introduction Regional Genetics Service Long chain acyl-CoA dehydrogenase (LCHAD) deficiency / mitochondrial trifunctional Levels 4-6, Barclay House protein (MTF) deficiency is an autosomal recessive disorder of mitochondrial beta- 37 Queen Square oxidation of fatty acids. The mitochondrial trifunctional protein is composed of 4 alpha London, WC1N 3BH and 4 beta subunits, which are encoded by the HADHA and HADHB genes, respectively. It is characterized by early-onset cardiomyopathy, hypoglycemia, T +44 (0) 20 7762 6888 neuropathy, and pigmentary retinopathy, and sudden death. There is also an infantile F +44 (0) 20 7813 8578 onset form with a hepatic Reye-like syndrome, and a late-adolescent onset form with primarily a skeletal myopathy. Tandem mass spectrometry of organic acids in urine, Samples required and carnitines in blood spots, allows the diagnosis to be unequivocally determined. An 5ml venous blood in plastic EDTA additional clinical complication can occur in the pregnant mothers of affected fetuses; bottles (>1ml from neonates) they may experience maternal acute fatty liver of pregnancy (AFLP) syndrome or Prenatal testing must be arranged hypertension/haemolysis, elevated liver enzymes and low platelets (HELLP) in advance, through a Clinical syndrome. Genetics department if possible. The genes encoding the HADHA and HADHB subunits are located on chromosome Amniotic fluid or CV samples 2p23.3. The pathogenic -

Expression and Characterization of Pantoea CO Dehydrogenase To
www.nature.com/scientificreports OPEN Expression and characterization of Pantoea CO dehydrogenase to utilize CO-containing industrial Received: 31 October 2016 Accepted: 06 February 2017 waste gas for expanding the Published: 14 March 2017 versatility of CO dehydrogenase Eun Sil Choi1,2,*, Kyoungseon Min3,*, Geun-Joong Kim2, Inchan Kwon1 & Yong Hwan Kim3 Although aerobic CO dehydrogenases (CODHs) might be applicable in various fields, their practical applications have been hampered by low activity and no heterologous expression. We, for the first time, could functionally express recombinant PsCODH in E. coli and obtained a highly concentrated recombinant enzyme using an easy and convenient method. Its electron acceptor spectra, optimum −1 conditions (pH 6.5 and 30 °C), and kinetic parameters (kcat of 12.97 s , Km of 0.065 mM, and specific activity of 0.86 Umg−1) were examined. Blast furnace gas (BFG) containing 20% CO, which is a waste gas from the steel-making process, was tested as a substrate for PsCODH. Even with BFG, the recombinant PsCODH retained 88.2% and 108.4% activity compared with those of pure CO and 20% CO, respectively. The results provide not only a promising strategy to utilize CO-containing industrial waste gases as cheap, abundant, and renewable resources but also significant information for further studies about cascade reactions producing value-added chemicals via CO2 as an intermediate produced by a CODH- based CO-utilization system, which would ultimately expand the versatility of CODH. Carbon monoxide (CO), which is a pollutant in the atmosphere, is massively emitted through both natural (e.g., production by plants and volcanic activity) and artificial processes (e.g., incomplete combustion of fuels and industrial processes)1–3. -

The Evolution and Population Genetics of the ALDH2 Locus: Random Genetic Drift, Selection, and Low Levels of Recombination
doi: 10.1046/j.1529-8817.2003.00060.x The evolution and population genetics of the ALDH2 locus: random genetic drift, selection, and low levels of recombination Hiroki Oota1, Andrew J. Pakstis1, Batsheva Bonne-Tamir2, David Goldman3, Elena Grigorenko4, Sylvester L. B. Kajuna5, Nganyirwa J. Karoma5, Selemani Kungulilo6, Ru-Band Lu7, Kunle Odunsi8, Friday Okonofua9, Olga V. Zhukova10, Judith R. Kidd1 and Kenneth K. Kidd1,∗ 1Department of Genetics, Yale University School of Medicine, 333 Cedar Street, P.O. Box 208005, New Haven, CT 06520-8005, USA 2Department of Human Genetics, Sackler School of Medicine, Tel Aviv University, Tel Aviv, Israel 3Laboratory of Neurogenetics, National Institute of Alcohol Abuse and Alcoholism, Rockville, MD 20852, USA 4Department of Psychology, Yale University, New Haven, CT 06520, USA 5The Hubert Kairuki Memorial University, Dar es Salaam, Tanzania 6Muhimbili University College of Health Sciences, Dar es Salaam, Tanzania 7Department of Psychiatry, Tri-Service General hospital, National Defense Medical Center, Taipei, Taiwan, R.O.C. 8Department of Gynecological Oncology, Roswell Park Cancer Institute, Buffalo, NY 14263, USA 9Department of Obstetrics and Gynecology, Faculty of Medicine, University of Benin, Benin City, Nigeria 10N.I. Vavilov Institute of General Genetics RAS, Moscow, Russia Summary The catalytic deficiency of human aldehyde dehydrogenase 2 (ALDH2) is caused by a nucleotide substitution (G1510A; Glu487Lys) in exon 12 of the ALDH2 locus. This SNP,and four non-coding SNPs, including one in the promoter, span 40 kb of ALDH2; these and one downstream STRP have been tested in 37 worldwide populations. Only four major SNP-defined haplotypes account for almost all chromosomes in all populations. -

Is Glyceraldehyde-3-Phosphate Dehydrogenase a Central Redox Mediator?
1 Is glyceraldehyde-3-phosphate dehydrogenase a central redox mediator? 2 Grace Russell, David Veal, John T. Hancock* 3 Department of Applied Sciences, University of the West of England, Bristol, 4 UK. 5 *Correspondence: 6 Prof. John T. Hancock 7 Faculty of Health and Applied Sciences, 8 University of the West of England, Bristol, BS16 1QY, UK. 9 [email protected] 10 11 SHORT TITLE | Redox and GAPDH 12 13 ABSTRACT 14 D-Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is an immensely important 15 enzyme carrying out a vital step in glycolysis and is found in all living organisms. 16 Although there are several isoforms identified in many species, it is now recognized 17 that cytosolic GAPDH has numerous moonlighting roles and is found in a variety of 18 intracellular locations, but also is associated with external membranes and the 19 extracellular environment. The switch of GAPDH function, from what would be 20 considered as its main metabolic role, to its alternate activities, is often under the 21 influence of redox active compounds. Reactive oxygen species (ROS), such as 22 hydrogen peroxide, along with reactive nitrogen species (RNS), such as nitric oxide, 23 are produced by a variety of mechanisms in cells, including from metabolic 24 processes, with their accumulation in cells being dramatically increased under stress 25 conditions. Overall, such reactive compounds contribute to the redox signaling of the 26 cell. Commonly redox signaling leads to post-translational modification of proteins, 27 often on the thiol groups of cysteine residues. In GAPDH the active site cysteine can 28 be modified in a variety of ways, but of pertinence, can be altered by both ROS and 29 RNS, as well as hydrogen sulfide and glutathione. -

How Is Alcohol Metabolized by the Body?
Overview: How Is Alcohol Metabolized by the Body? Samir Zakhari, Ph.D. Alcohol is eliminated from the body by various metabolic mechanisms. The primary enzymes involved are aldehyde dehydrogenase (ALDH), alcohol dehydrogenase (ADH), cytochrome P450 (CYP2E1), and catalase. Variations in the genes for these enzymes have been found to influence alcohol consumption, alcohol-related tissue damage, and alcohol dependence. The consequences of alcohol metabolism include oxygen deficits (i.e., hypoxia) in the liver; interaction between alcohol metabolism byproducts and other cell components, resulting in the formation of harmful compounds (i.e., adducts); formation of highly reactive oxygen-containing molecules (i.e., reactive oxygen species [ROS]) that can damage other cell components; changes in the ratio of NADH to NAD+ (i.e., the cell’s redox state); tissue damage; fetal damage; impairment of other metabolic processes; cancer; and medication interactions. Several issues related to alcohol metabolism require further research. KEY WORDS: Ethanol-to acetaldehyde metabolism; alcohol dehydrogenase (ADH); aldehyde dehydrogenase (ALDH); acetaldehyde; acetate; cytochrome P450 2E1 (CYP2E1); catalase; reactive oxygen species (ROS); blood alcohol concentration (BAC); liver; stomach; brain; fetal alcohol effects; genetics and heredity; ethnic group; hypoxia The alcohol elimination rate varies state of liver cells. Chronic alcohol con- he effects of alcohol (i.e., ethanol) widely (i.e., three-fold) among individ- sumption and alcohol metabolism are on various tissues depend on its uals and is influenced by factors such as strongly linked to several pathological concentration in the blood T chronic alcohol consumption, diet, age, consequences and tissue damage. (blood alcohol concentration [BAC]) smoking, and time of day (Bennion and Understanding the balance of alcohol’s over time. -

Dehydrogenase Classes
Proc. Nati. Acad. Sci. USA Vol. 91, pp. 4980-4984, May 1994 Biochemistry Fundamental molecular differences between alcohol dehydrogenase classes (Drosophila octano dehydrogenase/class m alcohol dehydrogenase/mo ur patterns/zinc enyme famy) OLLE DANIELSSON*, SILVIA ATRIANt, TERESA LUQUEt, LARS HJELMQVIST*, ROSER GONZALEZ-DUARTEt, AND HANS J6RNVALL*f *Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 77 Stockholm, Sweden; tCenter for Biotechnology, Karolinska Institutet, S-141 86 Huddinge, Sweden; and tDepartment of Genetics, University of Barcelona, E-08071 Barcelona, Spain Communicated by Sune Bergstrom, January 18, 1994 ABSTRACT Two types of alcohol dehydrogenase in sepa- ary patterns, with class III being "constant" and class I rate protein families are the "medium-chain" zinc enzymes "variable" (10), result in a consistent picture of the enzyme (including the classical liver and yeast forms) and the "short- system and place the classes of medium-chain alcohol dehy- chain" enzymes (including the insect form). Although the drogenases as separate enzymes in the cellular metabolism. medium-chain family has been characterized in prokaryotes Similarly, another protein family, short-chain dehydroge- and many eukaryotes (fungi, plants, cephalopods, and verte- nases, has also evolved into a family comprising many brates), insects have seemed to possess only the short-chain different enzyme activities, including an alcohol dehydroge- enzyme. We have now also characterized a medium-chain nase (11). This form operates by means of a completely alcohol dehydrogenase in Drosophila. The enzyme is identical different catalytic mechanism and is related to mammalian to insect octanol dehydrogenase. It Is a typical class m alcohol prostaglandin dehydrogenases/carbonyl reductase (12). -

Bioalcohol Production by a New Synthetic Route in a Hyperthermophilic Archaeon
COMMENTARY COMMENTARY Bioalcohol production by a new synthetic route in a hyperthermophilic archaeon Volker Müller1 Department of Molecular Microbiology & Bioenergetics, Institute of Molecular Biosciences, Johann Wolfgang Goethe University Frankfurt am Main, 60438 Frankfurt, Germany The still growing demand for bioethanol is establishing an electrochemical ion gradient currently met by two microbiological pro- across the cytoplasmic membrane that then cesses. The by far most important process is drives ATP synthesis by a membrane-bound the use of first- (sugar) or second- (lignocel- ATP synthase (5). One widespread, ferre- lulosic biomass) generation raw materials and doxin-fueled ion-translocating enyzme is + yeast as catalysts. Yeasts convert the sugars the ferredoxin:NAD oxidoreductase found via glycolysis to pyruvate, which is decar- in many bacteria and few archaea (6). In P. furiosus boxylated, and the resulting acetaldehyde is , the reduced ferredoxin is oxidized Fig. 1. Pathways for ethanol formation from glucose in then reduced to ethanol by a monofunctional by a different enzyme, a membrane-bound, yeasts (A) and bacteria (B). Adh1, alcoholdehydrogenase ethanol dehydrogenase. The second and in- multisubunit hydrogenase (Mbh) with evolu- 1; AdhE, bifunctional CoA-dependent ethanol/aldehyde dustrially less important is the bacterial pro- tionarysimilaritytothecomplexIofbacterial dehydrogenase. cess, in which pyruvate is oxidized to acetyl- and mitochondrial electron transport chains CoA, which is then reduced by a bifunctional (7). The Mbh is a proton-reducing (hydrogen aldehyde dehydrogenase/ethanol dehydroge- evolving) and sodium ion-extruding mem- used to drive acetate reduction and not the nase (AdhE) to ethanol. This pathway is brane protein complex (8, 9). generation of a chemiosmotic ion gradient, found in many fermenting bacteria (Fig. -

Moldx : BCKDHB Gene Test
Local Coverage Article: Billing and Coding: MolDX: BCKDHB Gene Test (A55099) Links in PDF documents are not guaranteed to work. To follow a web link, please use the MCD Website. Contractor Information CONTRACTOR NAME CONTRACT TYPE CONTRACT JURISDICTION STATE(S) NUMBER Noridian Healthcare Solutions, A and B MAC 01111 - MAC A J - E California - Entire State LLC Noridian Healthcare Solutions, A and B MAC 01112 - MAC B J - E California - Northern LLC Noridian Healthcare Solutions, A and B MAC 01182 - MAC B J - E California - Southern LLC Noridian Healthcare Solutions, A and B MAC 01211 - MAC A J - E American Samoa LLC Guam Hawaii Northern Mariana Islands Noridian Healthcare Solutions, A and B MAC 01212 - MAC B J - E American Samoa LLC Guam Hawaii Northern Mariana Islands Noridian Healthcare Solutions, A and B MAC 01311 - MAC A J - E Nevada LLC Noridian Healthcare Solutions, A and B MAC 01312 - MAC B J - E Nevada LLC Noridian Healthcare Solutions, A and B MAC 01911 - MAC A J - E American Samoa LLC California - Entire State Guam Hawaii Nevada Northern Mariana Islands Article Information General Information Article ID Original Effective Date Created on 12/19/2019. Page 1 of 6 A55099 10/17/2016 Article Title Revision Effective Date Billing and Coding: MolDX: BCKDHB Gene Test 12/01/2019 Article Type Revision Ending Date Billing and Coding N/A AMA CPT / ADA CDT / AHA NUBC Copyright Retirement Date Statement N/A CPT codes, descriptions and other data only are copyright 2018 American Medical Association. All Rights Reserved. Applicable FARS/HHSARS apply. Current Dental Terminology © 2018 American Dental Association. -
Supplementary Information Effect of Oncogene Activating Mutations And
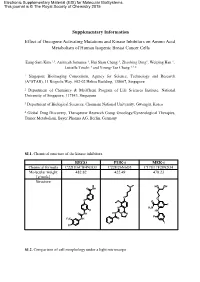
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is © The Royal Society of Chemistry 2015 Supplementary Information Effect of Oncogene Activating Mutations and Kinase Inhibitors on Amino Acid Metabolism of Human Isogenic Breast Cancer Cells Eung-Sam Kim 1,3, Animesh Samanta 1, Hui Shan Cheng 1, Zhaobing Ding1, Weiping Han 1, Luisella Toschi 4 and Young-Tae Chang 1,2,* 1 Singapore Bioimaging Consortium, Agency for Science, Technology and Research (A*STAR), 11 Biopolis Way, #02-02 Helios Building, 138667, Singapore 2 Department of Chemistry & MedChem Program of Life Sciences Institute, National University of Singapore, 117543, Singapore. 3 Department of Biological Sciences, Chonnam National University, Gwangju, Korea 4 Global Drug Discovery, Therapeutic Research Group Oncology/Gynecological Therapies, Tumor Metabolism, Bayer Pharma AG, Berlin, Germany SI.1. Chemical structure of the kinase inhibitors. REGO PI3K-i MEK-i Chemical formula C22H16ClF4N3O3 C22H26N6O3 C17H17F2IN2O4 Molecular weight 482.82 422.49 478.23 [g/mole] Structure N HO OH O NH O O O O O F N H2N F HN O NH HN N N F F3C NH O N I Cl SI.2. Comparison of cell morphology under a light microscope Scale bar: 100 µm 10x objective lens at Ti microscope (Nikon) SI.3. Scheme of the experimental method for metabolite extraction SI.4. Gradient profile of HPLC run. A : Buffer solution B: Mobile phase H O Time 2 (40 mM NaHPO4, (MeOH/ACN/H2O, Flow rate pH7.8 at RT) 45:45:10, v/v/v) (ml/min) (min) (%) (%) (%) 0 100 0 0 1.0 3 90 10 0 1.0 15 87 13 0 1.0 25 70 30 0 1.0 37 65 35 0 1.0 43 58 42 0 1.0 52 43 57 0 1.0 55 0 100 0 1.0 60 0 0 100 1.0 65 to 80 100 0 0 1.0 SI.5.