Outer Membrane Vesicles Mediated Horizontal Transfer of an Aerobic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Aerobic and Anaerobic Bacterial and Fungal Degradation of Pyrene: Mechanism Pathway Including Biochemical Reaction and Catabolic Genes
International Journal of Molecular Sciences Review Aerobic and Anaerobic Bacterial and Fungal Degradation of Pyrene: Mechanism Pathway Including Biochemical Reaction and Catabolic Genes Ali Mohamed Elyamine 1,2 , Jie Kan 1, Shanshan Meng 1, Peng Tao 1, Hui Wang 1 and Zhong Hu 1,* 1 Key Laboratory of Resources and Environmental Microbiology, Department of Biology, Shantou University, Shantou 515063, China; [email protected] (A.M.E.); [email protected] (J.K.); [email protected] (S.M.); [email protected] (P.T.); [email protected] (H.W.) 2 Department of Life Science, Faculty of Science and Technology, University of Comoros, Moroni 269, Comoros * Correspondence: [email protected] Abstract: Microbial biodegradation is one of the acceptable technologies to remediate and control the pollution by polycyclic aromatic hydrocarbon (PAH). Several bacteria, fungi, and cyanobacteria strains have been isolated and used for bioremediation purpose. This review paper is intended to provide key information on the various steps and actors involved in the bacterial and fungal aerobic and anaerobic degradation of pyrene, a high molecular weight PAH, including catabolic genes and enzymes, in order to expand our understanding on pyrene degradation. The aerobic degradation pathway by Mycobacterium vanbaalenii PRY-1 and Mycobactetrium sp. KMS and the anaerobic one, by the facultative bacteria anaerobe Pseudomonas sp. JP1 and Klebsiella sp. LZ6 are reviewed and presented, to describe the complete and integrated degradation mechanism pathway of pyrene. The different microbial strains with the ability to degrade pyrene are listed, and the degradation of Citation: Elyamine, A.M.; Kan, J.; pyrene by consortium is also discussed. -

Pseudomonas Chloritidismutans Sp. Nov., a Non- Denitrifying, Chlorate-Reducing Bacterium
International Journal of Systematic and Evolutionary Microbiology (2002), 52, 2183–2190 DOI: 10.1099/ijs.0.02102-0 Pseudomonas chloritidismutans sp. nov., a non- denitrifying, chlorate-reducing bacterium Laboratory of Microbiology, A. F. W. M. Wolterink, A. B. Jonker, S. W. M. Kengen and A. J. M. Stams Wageningen University, H. van Suchtelenweg 4, 6703 CT Wageningen, Author for correspondence: The Netherlands A. F. W. M. Wolterink. Tel: j31 317 484099. Fax: j31 317 483829. e-mail: Arthur.Wolterink!algemeen.micr.wag-ur.nl A Gram-negative, facultatively anaerobic, rod-shaped, dissimilatory chlorate- reducing bacterium, strain AW-1T, was isolated from biomass of an anaerobic chlorate-reducing bioreactor. Phylogenetic analysis of the 16S rDNA sequence showed 100% sequence similarity to Pseudomonas stutzeri DSM 50227 and 986% sequence similarity to the type strain of P. stutzeri (DSM 5190T). The species P. stutzeri possesses a high degree of genotypic and phenotypic heterogeneity. Therefore, eight genomic groups, termed genomovars, have been proposed based upon ∆Tm values, which were used to evaluate the quality of the pairing within heteroduplexes formed by DNA–DNA hybridization. In this study, DNA–DNA hybridization between strain AW-1T and P. stutzeri strains DSM 50227 and DSM 5190T revealed respectively 805 and 565% similarity. DNA–DNA hybridization between P. stutzeri strains DSM 50227 and DSM 5190T revealed 484% similarity. DNA–DNA hybridization indicated that strain AW-1T is not related at the species level to the type strain of P. stutzeri. However, strain AW-1T and P. stutzeri DSM 50227 are related at the species level. The physiological and biochemical properties of strain AW-1T and the two P. -

Section 4. Guidance Document on Horizontal Gene Transfer Between Bacteria
306 - PART 2. DOCUMENTS ON MICRO-ORGANISMS Section 4. Guidance document on horizontal gene transfer between bacteria 1. Introduction Horizontal gene transfer (HGT) 1 refers to the stable transfer of genetic material from one organism to another without reproduction. The significance of horizontal gene transfer was first recognised when evidence was found for ‘infectious heredity’ of multiple antibiotic resistance to pathogens (Watanabe, 1963). The assumed importance of HGT has changed several times (Doolittle et al., 2003) but there is general agreement now that HGT is a major, if not the dominant, force in bacterial evolution. Massive gene exchanges in completely sequenced genomes were discovered by deviant composition, anomalous phylogenetic distribution, great similarity of genes from distantly related species, and incongruent phylogenetic trees (Ochman et al., 2000; Koonin et al., 2001; Jain et al., 2002; Doolittle et al., 2003; Kurland et al., 2003; Philippe and Douady, 2003). There is also much evidence now for HGT by mobile genetic elements (MGEs) being an ongoing process that plays a primary role in the ecological adaptation of prokaryotes. Well documented is the example of the dissemination of antibiotic resistance genes by HGT that allowed bacterial populations to rapidly adapt to a strong selective pressure by agronomically and medically used antibiotics (Tschäpe, 1994; Witte, 1998; Mazel and Davies, 1999). MGEs shape bacterial genomes, promote intra-species variability and distribute genes between distantly related bacterial genera. Horizontal gene transfer (HGT) between bacteria is driven by three major processes: transformation (the uptake of free DNA), transduction (gene transfer mediated by bacteriophages) and conjugation (gene transfer by means of plasmids or conjugative and integrated elements). -
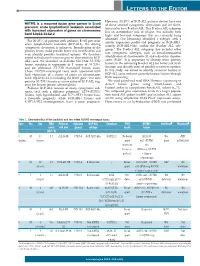
NUTM1 Is a Recurrent Fusion Gene Partner in B-Cell Precursor Acute
LETTERS TO THE EDITOR However, 20-25% of BCP-ALL patients do not have one NUTM1 is a recurrent fusion gene partner in B-cell of these sentinel cytogenetic aberrations and are there- precursor acute lymphoblastic leukemia associated fore said to have B-other ALL. This B-other ALL subgroup with increased expression of genes on chromosome has an intermediate risk of relapse, but includes both band 10p12.31-12.2 high- and low-risk subgroups that are currently being identified. Our laboratory identified a subtype with a For 20-25% of patients with pediatric B-cell precursor similar expression profile and prognosis as BCR-ABL1, acute lymphoblastic leukemia (BCP-ALL), the driving namely BCR-ABL1-like, within the B-other ALL sub- cytogenetic aberration is unknown. Identification of the group.2 The B-other ALL subgroup also includes other primary lesion could provide better risk stratification and rare cytogenetic subtypes, such as intrachromosomal even identify possible treatment options. We therefore amplification of chromosome 21 and a dicentric chromo- aimed to find novel recurrent genetic aberrations in BCP- 1 ALL cases. We identified an in-frame SLC12A6-NUTM1 some (9;20). It is important to identify more primary fusion, resulting in expression of 3’ exons of NUTM1, lesions in the remaining B-other ALL for better risk strat- and six additional NUTM1-rearranged fusion cases. ification and identification of possible treatment options. These NUTM1-rearranged cases were associated with In this study, we aimed to identify recurrent fusions in high expression of a cluster of genes on chromosome BCP-ALL cases without currently known lesions through band 10p12.31-12.2, including the BMI1 gene. -

Effect of Toluene on Pseudomonas Stutzeri ST-9 Morphology: Plasmolysis, Cell Size and 1
Canadian Journal of Microbiology Effect of toluene on Pseudomonas stutzeri ST -9 morphology: Plasmolysis, cell size and formation of outer membrane vesicles Journal: Canadian Journal of Microbiology Manuscript ID cjm-2016-0099.R1 Manuscript Type: Article Date Submitted by the Author: 18-Mar-2016 Complete List of Authors: Michael, Esti; Ariel University, Chemical Engineering Nitzan, YeshayahuDraft ; Bar-Ilan University Langzam, Yakov; Bar-Ilan University, The Mina & Everard Goodman Faculty of Life Sciences Luboshits, Galia ; Ariel University, Cahan, Rivka; Ariel University, Chemical Engineering Keyword: Pseudomonas, plasmolysis, toluene, outer membrane vesicles, morphology https://mc06.manuscriptcentral.com/cjm-pubs Page 1 of 27 Canadian Journal of Microbiology Effect of toluene on Pseudomonas stutzeri ST-9 morphology: Plasmolysis, cell size and 1 formation of outer membrane vesicles 2 3 Esti Michael 1,2 , Yeshayahu Nitzan 2, Yakov Langzam 2, Galia Luboshits 1 and Rivka Cahan 1* 4 1Department of Chemical Engineering, Ariel University, Ariel 40700, Israel 5 2The Mina & Everard Goodman Faculty of Life Sciences, Bar-Ilan University, 6 Ramat-Gan 52900, Israel 7 *Corresponding author 8 9 10 11 Draft 12 13 14 15 16 17 18 19 20 21 22 23 24 25 1 https://mc06.manuscriptcentral.com/cjm-pubs Canadian Journal of Microbiology Page 2 of 27 Abstract 1 Isolated toluene-degrading Pseudomonas stutzeri ST-9 bacteria were grown in a minimal 2 medium containing toluene (100 mg L -1) (MMT) or glucose (MMG) as the sole carbon source, 3 with specific growth rates of 0.019 h -1 and 0.042 h -1, respectively. Scanning (SEM) as well as 4 transmission (TEM) electron microscope analyses showed that the bacterial cells grown to mid 5 log in the presence of toluene possess a plasmolysis space. -

Aerobic and Nitrate Respiration Routes of Carbohydrate Catabolism in Pseudomonas S Tut Zeri
AN ABSTRACT OF THE THESIS OF WILLIAM JAN SPANGLER for the Ph. D. in MICROBIOLOGY (Name) (Degree) (Major) thesis is presented Date 7 / `> /q6! Title AEROBIC AND NITRATE RESPIRATION ROUTES OF CAR- BOHYDRATE CATABOLISM IN PSEUDOMONAS STUTZERI Abstract approved ( Major professor) Pseudomonas stutzeri and other denitrifying bacteria are able to grow under anaerobic conditions, using nitrate -oxygen as the terminal hydrogen acceptor, in a manner analagous to classical aerobic respiration with free -molecular oxygen. This rather unique phenomenon is known as nitrate respiration. Nitrate respiration has been studied with respect to the nitrate reducing enzymes and carrier systems involved in the reduction sequence, but very little emphasis has been placed on the metabolic pathways which are associated with nitrate respiration. This study was carried out in an attempt to establish the metabolic pathways operative, both under aerobic con- ditions and during nitrate respiration, in order to determine whether there was any shift of pathways under conditions of nitrate respira- tion. Primary pathways were determined by the radiorespirometric method using specifically labelled glucose and gluconate. The results, based primarily on the rate of decarboxylation of the C -1 and C -4 positions of glucose, indicated the operation of the Entner- Doudoroff and pentose phosphate pathways under both aerobic condi- tions and conditions of nitrate respiration. Evolution of 14CO2 from the other labels of glucose, as well as incorporation of these labels into the cell, indicated that terminal pathways such as the tricar- boxylic acid cycle or glyoxalate cycle might also be operative under both conditions of oxygen relationship. The secondary pathways were studied using specifically labelled acetate. -

Transposable Elements in Human Cancers by Genome-Wide EST Alignment
Genes Genet. Syst. (2007) 82, p. 145–156 Transposable elements in human cancers by genome-wide EST alignment Dae-Soo Kim1, Jae-Won Huh2 and Heui-Soo Kim1,2* 1PBBRC, Interdisciplinary Research Program of Bioinformatics, Pusan National University, Busan 609-735, Republic of Korea 2Division of Biological Sciences, College of Natural Sciences, Pusan National University, Busan 609-735, Republic of Korea (Received 24 November 2006, accepted 23 January 2007) Transposable elements may affect coding sequences, splicing patterns, and tran- scriptional regulation of human genes. Particles of the transposable elements have been detected in several tissues and tumors. Here, we report genome-wide analysis of gene expression regulated by transposable elements in human cancers. We adopted an analysis pipeline for screening methods to detect cancer- specific expression from expressed human sequences. We developed a database (TECESdb) for understanding the mechanism of cancer development in relation to transposable elements. A total of 999 genes fused with transposable elements were found to be cancer-related in our analysis of the EST database. According to GO (Gene Ontology) analysis, the majority of the 999 cancer-specific genes have functional association with gene receptor, DNA binding, and kinase activity. Our data could contribute greatly to our understanding of human cancers in relation to transposable elements. Key words: Transposable elements, Cancer, Fusion gene, Bioinformatics, EST also appeared in open-reading frames of functional INTRODUCTION human genes (Yulug et al., 1995; Makalowski et al., 1999; The human genome is estimated to be composed of 45% Nekrutenko and Li, 2001; Huh et al., 2006). transposable elements (International Human Genome The L1 5’UTR element is known to have an antisense Sequencing Consortium 2001). -

EWSR1 Gene EWS RNA Binding Protein 1
EWSR1 gene EWS RNA binding protein 1 Normal Function The EWSR1 gene provides instructions for making the EWS protein, whose function is not completely understood. The EWS protein has two regions that contribute to its function. One region, the transcriptional activation domain, allows the EWS protein to turn on (activate) the first step in the production of proteins from genes (transcription). The other region, the RNA-binding domain, allows the EWS protein to attach (bind) to the genetic blueprint for proteins called RNA. The EWS protein may be involved in piecing together this blueprint. Some studies suggest that the RNA-binding domain is able to block (inhibit) the activity of the transcriptional activation domain, and thus regulate the function of the EWS protein. Health Conditions Related to Genetic Changes Ewing sarcoma Mutations involving the EWSR1 gene can cause a type of cancerous tumor known as Ewing sarcoma. These tumors develop in bones or soft tissues, such as nerves and cartilage. There are several types of Ewing sarcoma, including Ewing sarcoma of bone, extraosseous Ewing sarcoma, peripheral primitive neuroectodermal tumor, and Askin tumor. The mutations that cause these tumors are acquired during a person's lifetime and are present only in the tumor cells. This type of genetic change, called a somatic mutation, is not inherited. The most common mutation that causes Ewing sarcoma is a rearrangement (translocation) of genetic material between chromosome 22 and chromosome 11. This translocation, written as t(11;22), fuses part of the EWSR1 gene on chromosome 22 with part of another gene on chromosome 11 called FLI1, creating an EWSR1/FLI1 fusion gene. -

Birth of a Chimeric Primate Gene by Capture of the Transposase Gene
Birth of a chimeric primate gene by capture of the SEE COMMENTARY transposase gene from a mobile element Richard Cordaux*, Swalpa Udit†, Mark A. Batzer*, and Ce´ dric Feschotte†‡ *Department of Biological Sciences, Biological Computation and Visualization Center, Center for BioModular Multi-Scale Systems, Louisiana State University, 202 Life Sciences Building, Baton Rouge, LA 70803; and †Department of Biology, University of Texas, Arlington, TX 76019 Edited by Susan R. Wessler, University of Georgia, Athens, GA, and approved March 27, 2006 (received for review February 10, 2006) The emergence of new genes and functions is of central impor- SETMAR transcript, which consists of these three exons, is tance to the evolution of species. The contribution of various types predicted to encode a protein of 671 amino acids and is of duplications to genetic innovation has been extensively inves- supported by 48 human cDNA clones from 18 different normal tigated. Less understood is the creation of new genes by recycling and͞or cancerous tissues (Table 1, which is published as sup- of coding material from selfish mobile genetic elements. To inves- porting information on the PNAS web site; refs. 14 and 15). tigate this process, we reconstructed the evolutionary history of These data suggest that the SETMAR protein is broadly ex- SETMAR, a new primate chimeric gene resulting from fusion of a pressed and has an important, yet unknown, function in human. SET histone methyltransferase gene to the transposase gene of a Recently, it was shown that the SET domain of the SETMAR mobile element. We show that the transposase gene was recruited protein exhibits histone methyltransferase activity (15), as do all as part of SETMAR 40–58 million years ago, after the insertion of known SET domains (16, 17). -

Isolation and Characterization of a Novel Denitrifying Bacterium with High Nitrate Removal: Pseudomonas Stutzeri
Iran. J. Environ. Health. Sci. Eng., 2010, Vol. 7, No. 4, pp. 313-318 ISOLATION AND CHARACTERIZATION OF A NOVEL DENITRIFYING BACTERIUM WITH HIGH NITRATE REMOVAL: PSEUDOMONAS STUTZERI *1A. Rezaee, 2H. Godini, 1S. Dehestani, 1S. Kaviani 1 Department of Environmental Health, Faculty of Medical Sciences, Tarbiat Modares University, Tehran, Iran 2 Department of Environmental Health, School of Public Health, Lorestan University of Medical Sciences, Khoramabbad, Iran Received 16 August 2009; revised 13 Jully 2010; accepted 20 August 2010 ABSTRACT The aim of this study was to isolate and characterize a high efficiency denitrifier bacterium for reducing nitrate in wastewater. Six denitrifier bacteria with nitrate removal activities were isolated from a petrochemical industry effluent with high salinity and high nitrogen concentrations without treatment. The isolated bacteria were tested for nitrate reomoval activity. One of the bacterium displayed the highest reduction of nitrate. The strain was preliminarily identified using biochemical tests and further identified based on similarity of PCR-16S rRNA using universal primers. Biochemical and molecular experiments showed that the best bacterium with high nitrate removal potential was Pseudomonas stutzeri, a member of the α subclass of the class Proteobacteria. The extent of nitrate removal efficiency was 99% at 200 mg/L NO3 and the nitrite content of the effluent was in the prescribed limit. The experiments showed the ability of Pseudomonas stutzeri to rapidly remove nitrate under anoxic conditions. The strain showed to be potentially good candidate for biodenitrification of high nitrate solutions. Key words: Pseudomonas stutzeri; Denitrification; Polymerase Chain Reaction, Isolation; Characterization INTRODUCTION Biological denitrification is a process carried to respire anaerobically using nitrogen oxides as out by numerous genera of bacteria. -

Selective Induction of Leukemia-Associated Fusion Genes by High-Dose Ionizing Radiation1
[CANCER RESEARCH 58. 421-425. February I. 1<W8| Selective Induction of Leukemia-associated Fusion Genes by High-Dose Ionizing Radiation1 Michael W. N. Deininger, Shikha Bose, Joanna Gora-Tybor, Xiu-Hua Yan, John M. Goldman, and Junia V. Melo2 Leukaemia Research Fumi Centre for Adult Leukaemia. Department of Haemah>li>/;\: Royal Postgraduale Medical School, Ducane Road, London W12 ONN, United Kingdom ABSTRACT event involves the acquisition of the genetic abnormality whose "success" in the production of a leukemic phenotype will depend on There is strong clinical and epidemiológica! evidence that ionizing its capacity to impart to the target cell a proliferative and/or survival radiation can cause leukemia by inducing DNA damage. This crucial advantage over its normal neighbors. In molecular terms, the gener initiation event is believed to be the result of random DNA breakage and misrepair, whereas the subsequent steps, promotion and progression, ation of a potentially successful reciprocal chromosomal translocation must rely on mechanisms of selective pressure to provide the expanding requires that: (a) at least two independent DNA DSBs occur, one in leukemic population with its proliferative/renewal advantage. To investi each chromosome partner; (b) the two breaks occur simultaneously, gate the susceptibility of human cells to external agents at the genetic i.e., within the same cell cycle, so that the two ends of one broken recombination stage of leukemogenesis, we subjected two hematopoietic chromosome are available to interact and be ligated (misrepaired) to cell lines, KG1 and III.6(1, to high doses of y-irradiation. The irradiation the respective complementary broken ends of the other chromosome; induced the formation of fusion genes characteristic of leukemia in both and (c) the recombination observes the polarity of the DNA molecule. -

DNA Transposons and the Evolution of Eukaryotic Genomes
ANRV329-GE41-15 ARI 12 October 2007 11:1 DNA Transposons and the Evolution of Eukaryotic Genomes Cedric´ Feschotte and Ellen J. Pritham Department of Biology, University of Texas, Arlington, Texas 76019; email: [email protected] Annu. Rev. Genet. 2007. 41:331–68 Key Words The Annual Review of Genetics is online at transposable elements, transposase, molecular domestication, http://genet.annualreviews.org chromosomal rearrangements This article’s doi: 10.1146/annurev.genet.40.110405.090448 Abstract Copyright c 2007 by Annual Reviews. Transposable elements are mobile genetic units that exhibit broad All rights reserved by Fordham University on 11/23/12. For personal use only. diversity in their structure and transposition mechanisms. Transpos- 0066-4197/07/1201-0331$20.00 able elements occupy a large fraction of many eukaryotic genomes and their movement and accumulation represent a major force shap- Annu. Rev. Genet. 2007.41:331-68. Downloaded from www.annualreviews.org ing the genes and genomes of almost all organisms. This review fo- cuses on DNA-mediated or class 2 transposons and emphasizes how this class of elements is distinguished from other types of mobile elements in terms of their structure, amplification dynamics, and genomic effect. We provide an up-to-date outlook on the diversity and taxonomic distribution of all major types of DNA transposons in eukaryotes, including Helitrons and Mavericks. We discuss some of the evolutionary forces that influence their maintenance and di- versification in various genomic environments. Finally, we highlight how the distinctive biological features of DNA transposons have contributed to shape genome architecture and led to the emergence of genetic innovations in different eukaryotic lineages.