High Throughput Screening and Identification of Coagulopathic Snake Venom Proteins and 2 Peptides Using Nanofractionation and Proteomics Approaches
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

New Records of Snakes (Squamata: Serpentes) from Hoa Binh Province, Northwestern Vietnam
Bonn zoological Bulletin 67 (1): 15–24 May 2018 New records of snakes (Squamata: Serpentes) from Hoa Binh Province, northwestern Vietnam Truong Quang Nguyen1,2,*, Tan Van Nguyen 1,3, Cuong The Pham1,2, An Vinh Ong4 & Thomas Ziegler5 1 Institute of Ecology and Biological Resources, Vietnam Academy of Science and Technology, 18 Hoang Quoc Viet Road, Hanoi, Vietnam 2 Graduate University of Science and Technology, Vietnam Academy of Science and Technology, 18 Hoang Quoc Viet Road, Hanoi, Vietnam 3 Save Vietnam’s Wildlife, Cuc Phuong National Park, Ninh Binh Province, Vietnam 4 Vinh University, 182 Le Duan Road, Vinh City, Nghe An Province, Vietnam 5 AG Zoologischer Garten Köln, Riehler Strasse 173, D-50735 Cologne, Germany * Corresponding author. E-mail: [email protected] Abstract. We report nine new records of snakes from Hoa Binh Province based on a reptile collection from Thuong Tien, Hang Kia-Pa Co, Ngoc Son-Ngo Luong nature reserves, and Tan Lac District, comprising six species of Colubri- dae (Dryocalamus davisonii, Euprepiophis mandarinus, Lycodon futsingensis, L. meridionalis, Sibynophis collaris and Sinonatrix aequifasciata), one species of Pareatidae (Pareas hamptoni) and two species of Viperidae (Protobothrops mu- crosquamatus and Trimeresurus gumprechti). In addition, we provide an updated list of 43 snake species from Hoa Binh Province. The snake fauna of Hoa Binh contains some species of conservation concern with seven species listed in the Governmental Decree No. 32/2006/ND-CP (2006), nine species listed in the Vietnam Red Data Book (2007), and three species listed in the IUCN Red List (2018). Key words. New records, snakes, taxonomy, Hoa Binh Province. -

Medically Important Differences in Snake Venom Composition Are Dictated by Distinct Postgenomic Mechanisms
Medically important differences in snake venom composition are dictated by distinct postgenomic mechanisms Nicholas R. Casewella,b,1, Simon C. Wagstaffc, Wolfgang Wüsterb, Darren A. N. Cooka, Fiona M. S. Boltona, Sarah I. Kinga, Davinia Plad, Libia Sanzd, Juan J. Calveted, and Robert A. Harrisona aAlistair Reid Venom Research Unit and cBioinformatics Unit, Liverpool School of Tropical Medicine, Liverpool L3 5QA, United Kingdom; bMolecular Ecology and Evolution Group, School of Biological Sciences, Bangor University, Bangor LL57 2UW, United Kingdom; and dInstituto de Biomedicina de Valencia, Consejo Superior de Investigaciones Científicas, 11 46010 Valencia, Spain Edited by David B. Wake, University of California, Berkeley, CA, and approved May 14, 2014 (received for review March 27, 2014) Variation in venom composition is a ubiquitous phenomenon in few (approximately 5–10) multilocus gene families, with each snakes and occurs both interspecifically and intraspecifically. family capable of producing related isoforms generated by Venom variation can have severe outcomes for snakebite victims gene duplication events occurring over evolutionary time (1, 14, by rendering the specific antibodies found in antivenoms in- 15). The birth and death model of gene evolution (16) is fre- effective against heterologous toxins found in different venoms. quently invoked as the mechanism giving rise to venom gene The rapid evolutionary expansion of different toxin-encoding paralogs, with evidence that natural selection acting on surface gene families in different snake lineages is widely perceived as the exposed residues of the resulting gene duplicates facilitates main cause of venom variation. However, this view is simplistic subfunctionalization/neofunctionalization of the encoded proteins and disregards the understudied influence that processes acting (15, 17–19). -

WHO Guidance on Management of Snakebites
GUIDELINES FOR THE MANAGEMENT OF SNAKEBITES 2nd Edition GUIDELINES FOR THE MANAGEMENT OF SNAKEBITES 2nd Edition 1. 2. 3. 4. ISBN 978-92-9022- © World Health Organization 2016 2nd Edition All rights reserved. Requests for publications, or for permission to reproduce or translate WHO publications, whether for sale or for noncommercial distribution, can be obtained from Publishing and Sales, World Health Organization, Regional Office for South-East Asia, Indraprastha Estate, Mahatma Gandhi Marg, New Delhi-110 002, India (fax: +91-11-23370197; e-mail: publications@ searo.who.int). The designations employed and the presentation of the material in this publication do not imply the expression of any opinion whatsoever on the part of the World Health Organization concerning the legal status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. Dotted lines on maps represent approximate border lines for which there may not yet be full agreement. The mention of specific companies or of certain manufacturers’ products does not imply that they are endorsed or recommended by the World Health Organization in preference to others of a similar nature that are not mentioned. Errors and omissions excepted, the names of proprietary products are distinguished by initial capital letters. All reasonable precautions have been taken by the World Health Organization to verify the information contained in this publication. However, the published material is being distributed without warranty of any kind, either expressed or implied. The responsibility for the interpretation and use of the material lies with the reader. In no event shall the World Health Organization be liable for damages arising from its use. -

(Apicomplexa: Adeleorina) from the Blood of Echis Pyramidum: Morphology and SSU Rdna Sequence Hepatozoon Pyramidumi Sp
Original Article ISSN 1984-2961 (Electronic) www.cbpv.org.br/rbpv Hepatozoon pyramidumi sp. n. (Apicomplexa: Adeleorina) from the blood of Echis pyramidum: morphology and SSU rDNA sequence Hepatozoon pyramidumi sp. n. (Apicomplexa: Adeleorina) do sangue de Echis pyramidum: morfologia e sequência de SSU rDNA Lamjed Mansour1,2; Heba Mohamed Abdel-Haleem3; Esam Sharf Al-Malki4; Saleh Al-Quraishy1; Abdel-Azeem Shaban Abdel-Baki3* 1 Zoology Department, College of Science, King Saud University, Riyadh, Saudi Arabia 2 Unité de Recherche de Biologie Intégrative et Écologie Évolutive et Fonctionnelle des Milieux Aquatiques, Département de Biologie, Faculté des Sciences de Tunis, Université de Tunis El Manar, Tunisia 3 Zoology Department, Faculty of Science, Beni-Suef University, Beni-Suef, Egypt 4 Department of Biology, College of Sciences, Majmaah University, Majmaah 11952, Riyadh Region, Saudi Arabia How to cite: Mansour L, Abdel-Haleem HM, Al-Malki ES, Al-Quraishy S, Abdel-Baki AZS. Hepatozoon pyramidumi sp. n. (Apicomplexa: Adeleorina) from the blood of Echis pyramidum: morphology and SSU rDNA sequence. Braz J Vet Parasitol 2020; 29(2): e002420. https://doi.org/10.1590/S1984-29612020019 Abstract Hepatozoon pyramidumi sp. n. is described from the blood of the Egyptian saw-scaled viper, Echis pyramidum, captured from Saudi Arabia. Five out of ten viper specimens examined (50%) were found infected with Hepatozoon pyramidumi sp. n. with parasitaemia level ranged from 20-30%. The infection was restricted only to the erythrocytes. Two morphologically different forms of intraerythrocytic stages were observed; small and mature gamonts. The small ganomt with average size of 10.7 × 3.5 μm. Mature gamont was sausage-shaped with recurved poles measuring 16.3 × 4.2 μm in average size. -

Self-Envenomation in an Egyptian Saw-Scaled Viper Using Region of Interest
The biter bit? Investigation of possible in-ovo self- envenomation in an Egyptian saw-scaled viper using region of interest X-ray microtomography John Mulley, Richard E Johnston Proven examples of self-envenomation by venomous snakes, and especially instances of death as a result of these events, are extremely rare, if not non-existent. Here we use Region of Interest X-ray microtomography to investigate a putative case of fatal in-ovo s t self-envenomation in the Egyptian saw-scaled viper, Echis pyramidum. Our analyses have n i provided unprecedented insight into the skeletal anatomy of a late-stage embryonic snake r P and the disposition of the fangs without disrupting or destroying a unique biological e r specimen. P PeerJ PrePrints | http://dx.doi.org/10.7287/peerj.preprints.624v1 | CC-BY 4.0 Open Access | rec: 19 Nov 2014, publ: 19 Nov 2014 1 Title page 2 3 The biter bit? Investigation of possible in-ovo self-envenomation in an Egyptian saw-scaled 4 viper using region of interest X-ray microtomography 5 6 Richard E Johnston1 and John F Mulley2* 7 8 1. College of Engineering, Swansea University, Swansea, SA2 8PP, United Kingdom s 9 2. School of Biological Sciences, Bangor University, Bangor, Gwynedd LL57 2UW, United t n i 10 Kingdom r P 11 e r P 12 *To whom correspondence should be addressed ([email protected]) 13 14 15 16 17 18 19 20 21 22 23 24 25 1 PeerJ PrePrints | http://dx.doi.org/10.7287/peerj.preprints.624v1 | CC-BY 4.0 Open Access | rec: 19 Nov 2014, publ: 19 Nov 2014 26 Abstract 27 Proven examples of self-envenomation by venomous snakes, and especially instances of 28 death as a result of these events, are extremely rare, if not non-existent. -

Biogeography of the Reptiles of the Central African Republic
African Journal of Herpetology, 2006 55(1): 23-59. ©Herpetological Association of Africa Original article Biogeography of the Reptiles of the Central African Republic LAURENT CHIRIO AND IVAN INEICH Muséum National d’Histoire Naturelle Département de Systématique et Evolution (Reptiles) – USM 602, Case Postale 30, 25, rue Cuvier, F-75005 Paris, France This work is dedicated to the memory of our friend and colleague Jens B. Rasmussen, Curator of Reptiles at the Zoological Museum of Copenhagen, Denmark Abstract.—A large number of reptiles from the Central African Republic (CAR) were collected during recent surveys conducted over six years (October 1990 to June 1996) and deposited at the Paris Natural History Museum (MNHN). This large collection of 4873 specimens comprises 86 terrapins and tortois- es, five crocodiles, 1814 lizards, 38 amphisbaenids and 2930 snakes, totalling 183 species from 78 local- ities within the CAR. A total of 62 taxa were recorded for the first time in the CAR, the occurrence of numerous others was confirmed, and the known distribution of several taxa is greatly extended. Based on this material and an additional six species known to occur in, or immediately adjacent to, the coun- try from other sources, we present a biogeographical analysis of the 189 species of reptiles in the CAR. Key words.—Central African Republic, reptile fauna, biogeography, distribution. he majority of African countries have been improved; known distributions of many species Tthe subject of several reptile studies (see are greatly expanded and distributions of some for example LeBreton 1999 for Cameroon). species are questioned in light of our results. -

Testing the Toxicofera: Comparative Reptile Transcriptomics Casts Doubt on the Single, Early Evolution of the Reptile Venom Syst
bioRxiv preprint doi: https://doi.org/10.1101/006031; this version posted June 6, 2014. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Title page 2 3 Title: 4 Testing the Toxicofera: comparative reptile transcriptomics casts doubt on the single, early 5 evolution of the reptile venom system 6 7 Authors: 8 Adam D Hargreaves1, Martin T Swain2, Darren W Logan3 and John F Mulley1* 9 10 Affiliations: 11 1. School of Biological Sciences, Bangor University, Brambell Building, Deiniol Road, 12 Bangor, Gwynedd, LL57 2UW, United Kingdom 13 14 2. Institute of Biological, Environmental & Rural Sciences, Aberystwyth University, 15 Penglais, Aberystwyth, Ceredigion, SY23 3DA, United Kingdom 16 17 3. Wellcome Trust Sanger Institute, Hinxton, Cambridge, CB10 1HH, United Kingdom 18 19 *To whom correspondence should be addressed: [email protected] 20 21 22 Author email addresses: 23 John Mulley - [email protected] 24 Adam Hargreaves - [email protected] 25 Darren Logan - [email protected] 26 Martin Swain - [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/006031; this version posted June 6, 2014. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -

Amphibians and Reptiles of the Mediterranean Basin
Chapter 9 Amphibians and Reptiles of the Mediterranean Basin Kerim Çiçek and Oğzukan Cumhuriyet Kerim Çiçek and Oğzukan Cumhuriyet Additional information is available at the end of the chapter Additional information is available at the end of the chapter http://dx.doi.org/10.5772/intechopen.70357 Abstract The Mediterranean basin is one of the most geologically, biologically, and culturally complex region and the only case of a large sea surrounded by three continents. The chapter is focused on a diversity of Mediterranean amphibians and reptiles, discussing major threats to the species and its conservation status. There are 117 amphibians, of which 80 (68%) are endemic and 398 reptiles, of which 216 (54%) are endemic distributed throughout the Basin. While the species diversity increases in the north and west for amphibians, the reptile diversity increases from north to south and from west to east direction. Amphibians are almost twice as threatened (29%) as reptiles (14%). Habitat loss and degradation, pollution, invasive/alien species, unsustainable use, and persecution are major threats to the species. The important conservation actions should be directed to sustainable management measures and legal protection of endangered species and their habitats, all for the future of Mediterranean biodiversity. Keywords: amphibians, conservation, Mediterranean basin, reptiles, threatened species 1. Introduction The Mediterranean basin is one of the most geologically, biologically, and culturally complex region and the only case of a large sea surrounded by Europe, Asia and Africa. The Basin was shaped by the collision of the northward-moving African-Arabian continental plate with the Eurasian continental plate which occurred on a wide range of scales and time in the course of the past 250 mya [1]. -

Substrate Thermal Properties Influence Ventral Brightness Evolution In
ARTICLE https://doi.org/10.1038/s42003-020-01524-w OPEN Substrate thermal properties influence ventral brightness evolution in ectotherms ✉ Jonathan Goldenberg 1 , Liliana D’Alba 1, Karen Bisschop 2,3, Bram Vanthournout1 & Matthew D. Shawkey 1 1234567890():,; The thermal environment can affect the evolution of morpho-behavioral adaptations of ectotherms. Heat is transferred from substrates to organisms by conduction and reflected radiation. Because brightness influences the degree of heat absorption, substrates could affect the evolution of integumentary optical properties. Here, we show that vipers (Squa- mata:Viperidae) inhabiting hot, highly radiative and superficially conductive substrates have evolved bright ventra for efficient heat transfer. We analyzed the brightness of 4161 publicly available images from 126 species, and we found that substrate type, alongside latitude and body mass, strongly influences ventral brightness. Substrate type also significantly affects dorsal brightness, but this is associated with different selective forces: activity-pattern and altitude. Ancestral estimation analysis suggests that the ancestral ventral condition was likely moderately bright and, following divergence events, some species convergently increased their brightness. Vipers diversified during the Miocene and the enhancement of ventral brightness may have facilitated the exploitation of arid grounds. We provide evidence that integument brightness can impact the behavioral ecology of ectotherms. 1 Evolution and Optics of Nanostructures group, Department -
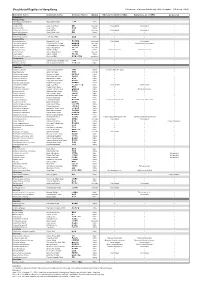
Checklist of Reptiles in Hong Kong © Programme of Ecology & Biodiversity, HKU (Last Update: 10 September 2012)
Checklist of Reptiles in Hong Kong © Programme of Ecology & Biodiversity, HKU (Last Update: 10 September 2012) Scientific name Common name Chinese Name Status Ades & Kendrick (2004) Karsen et al. (1998) Uetz et al. Testudines Platysternidae Platysternon megacephalum Big-headed Terrapin 大頭龜 Native Cheloniidae Caretta caretta Loggerhead Turtle 蠵龜 Uncertain Not included Not included Chelonia mydas Green Turtle 緣海龜 Native Eretmochelys imbricata Hawksbill Turtle 玳瑁 Uncertain Not included Not included Lepidochelys olivacea Pacific Ridley Turtle 麗龜 Native Dermochelyidae Dermochelys coriacea Leatherback Turtle 稜皮龜 Native Geoemydidae Cuora amboinensis Malayan Box Turtle 馬來閉殼龜 Introduced Not included Not included Cuora flavomarginata Yellow-lined Box Terrapin 黃緣閉殼龜 Uncertain Cistoclemmys flavomarginata Cuora trifasciata Three-banded Box Terrapin 三線閉殼龜 Native Mauremys mutica Chinese Pond Turtle 黃喉水龜 Uncertain Mauremys reevesii Reeves' Terrapin 烏龜 Native Chinemys reevesii Chinemys reevesii Ocadia sinensis Chinese Striped Turtle 中華花龜 Uncertain Sacalia bealei Beale's Terrapin 眼斑水龜 Native Trachemys scripta elegans Red-eared Slider 巴西龜 / 紅耳龜 Introduced Trionychidae Palea steindachneri Wattle-necked Soft-shelled Turtle 山瑞鱉 Uncertain Pelodiscus sinensis Chinese Soft-shelled Turtle 中華鱉 / 水魚 Native Squamata - Serpentes Colubridae Achalinus rufescens Rufous Burrowing Snake 棕脊蛇 Native Achalinus refescens (typo) Ahaetulla prasina Jade Vine Snake 綠瘦蛇 Uncertain Amphiesma atemporale Mountain Keelback 無顳鱗游蛇 Native -

Checklist of Amphibians and Reptiles of Morocco: a Taxonomic Update and Standard Arabic Names
Herpetology Notes, volume 14: 1-14 (2021) (published online on 08 January 2021) Checklist of amphibians and reptiles of Morocco: A taxonomic update and standard Arabic names Abdellah Bouazza1,*, El Hassan El Mouden2, and Abdeslam Rihane3,4 Abstract. Morocco has one of the highest levels of biodiversity and endemism in the Western Palaearctic, which is mainly attributable to the country’s complex topographic and climatic patterns that favoured allopatric speciation. Taxonomic studies of Moroccan amphibians and reptiles have increased noticeably during the last few decades, including the recognition of new species and the revision of other taxa. In this study, we provide a taxonomically updated checklist and notes on nomenclatural changes based on studies published before April 2020. The updated checklist includes 130 extant species (i.e., 14 amphibians and 116 reptiles, including six sea turtles), increasing considerably the number of species compared to previous recent assessments. Arabic names of the species are also provided as a response to the demands of many Moroccan naturalists. Keywords. North Africa, Morocco, Herpetofauna, Species list, Nomenclature Introduction mya) led to a major faunal exchange (e.g., Blain et al., 2013; Mendes et al., 2017) and the climatic events that Morocco has one of the most varied herpetofauna occurred since Miocene and during Plio-Pleistocene in the Western Palearctic and the highest diversities (i.e., shift from tropical to arid environments) promoted of endemism and European relict species among allopatric speciation (e.g., Escoriza et al., 2006; Salvi North African reptiles (Bons and Geniez, 1996; et al., 2018). Pleguezuelos et al., 2010; del Mármol et al., 2019). -

List of Reptile Species in Hong Kong
List of Reptile Species in Hong Kong Family No. of Species Common Name Scientific Name Order TESTUDOFORMES Cheloniidae 4 Loggerhead Turtle Caretta caretta Green Turtle Chelonia mydas Hawksbill Turtle Eretmochelys imbricata Olive Ridley Turtle Lepidochelys olivacea Dermochelyidae 1 Leatherback Turtle Dermochelys coriacea Emydidae 1 Red-eared Slider * Trachemys scripta elegans Geoemydidae 3 Three-banded Box Turtle Cuora trifasciata Reeves' Turtle Mauremys reevesii Beale's Turtle Sacalia bealei Platysternidae 1 Big-headed Turtle Platysternon megacephalum Trionychidae 1 Chinese Soft-shelled Turtle Pelodiscus sinensis Order SQUAMATA Suborder LACERTILIA Agamidae 1 Changeable Lizard Calotes versicolor Lacertidae 1 Grass Lizard Takydromus sexlineatus ocellatus Scincidae 11 Chinese Forest Skink Ateuchosaurus chinensis Long-tailed Skink Eutropis longicaudata Chinese Skink Plestiodon chinensis chinensis Five-striped Blue-tailed Skink Plestiodon elegans Blue-tailed Skink Plestiodon quadrilineatus Vietnamese Five-lined Skink Plestiodon tamdaoensis Slender Forest Skink Scincella modesta Reeve's Smooth Skink Scincella reevesii Brown Forest Skink Sphenomorphus incognitus Indian Forest Skink Sphenomorphus indicus Chinese Waterside Skink Tropidophorus sinicus Varanidae 1 Common Water Monitor Varanus salvator Dibamidae 1 Bogadek's Burrowing Lizard Dibamus bogadeki Gekkonidae 8 Four-clawed Gecko Gehyra mutilata Chinese Gecko Gekko chinensis Tokay Gecko Gekko gecko Bowring's Gecko Hemidactylus bowringii Brook's Gecko* Hemidactylus brookii House Gecko* Hemidactylus