Caveolae, CAVIN1, and Lipid Peroxidation Mediate the Oxidative Stress Response
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mea6 Controls VLDL Transport Through the Coordinated Regulation of COPII Assembly
Cell Research (2016) 26:787-804. npg © 2016 IBCB, SIBS, CAS All rights reserved 1001-0602/16 $ 32.00 ORIGINAL ARTICLE www.nature.com/cr Mea6 controls VLDL transport through the coordinated regulation of COPII assembly Yaqing Wang1, *, Liang Liu1, 2, *, Hongsheng Zhang1, 2, Junwan Fan1, 2, Feng Zhang1, 2, Mei Yu3, Lei Shi1, Lin Yang1, Sin Man Lam1, Huimin Wang4, Xiaowei Chen4, Yingchun Wang1, Fei Gao5, Guanghou Shui1, Zhiheng Xu1, 6 1State Key Laboratory of Molecular Developmental Biology, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China; 2University of Chinese Academy of Sciences, Beijing 100101, China; 3School of Life Science, Shandong University, Jinan 250100, China; 4Institute of Molecular Medicine, Peking University, Beijing 100871, China; 5State Key Laboratory of Reproductive Biology, Institute of Zoology, Chinese Academy of Sciences, Beijing 100101, China; 6Translation- al Medical Center for Stem Cell Therapy, Shanghai East Hospital, Tongji University School of Medicine, Shanghai 200120, China Lipid accumulation, which may be caused by the disturbance in very low density lipoprotein (VLDL) secretion in the liver, can lead to fatty liver disease. VLDL is synthesized in endoplasmic reticulum (ER) and transported to Golgi apparatus for secretion into plasma. However, the underlying molecular mechanism for VLDL transport is still poor- ly understood. Here we show that hepatocyte-specific deletion of meningioma-expressed antigen 6 (Mea6)/cutaneous T cell lymphoma-associated antigen 5C (cTAGE5C) leads to severe fatty liver and hypolipemia in mice. Quantitative lip- idomic and proteomic analyses indicate that Mea6/cTAGE5 deletion impairs the secretion of different types of lipids and proteins, including VLDL, from the liver. -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

RNF11 at the Crossroads of Protein Ubiquitination
biomolecules Review RNF11 at the Crossroads of Protein Ubiquitination Anna Mattioni, Luisa Castagnoli and Elena Santonico * Department of Biology, University of Rome Tor Vergata, Via della ricerca scientifica, 00133 Rome, Italy; [email protected] (A.M.); [email protected] (L.C.) * Correspondence: [email protected] Received: 29 September 2020; Accepted: 8 November 2020; Published: 11 November 2020 Abstract: RNF11 (Ring Finger Protein 11) is a 154 amino-acid long protein that contains a RING-H2 domain, whose sequence has remained substantially unchanged throughout vertebrate evolution. RNF11 has drawn attention as a modulator of protein degradation by HECT E3 ligases. Indeed, the large number of substrates that are regulated by HECT ligases, such as ITCH, SMURF1/2, WWP1/2, and NEDD4, and their role in turning off the signaling by ubiquitin-mediated degradation, candidates RNF11 as the master regulator of a plethora of signaling pathways. Starting from the analysis of the primary sequence motifs and from the list of RNF11 protein partners, we summarize the evidence implicating RNF11 as an important player in modulating ubiquitin-regulated processes that are involved in transforming growth factor beta (TGF-β), nuclear factor-κB (NF-κB), and Epidermal Growth Factor (EGF) signaling pathways. This connection appears to be particularly significant, since RNF11 is overexpressed in several tumors, even though its role as tumor growth inhibitor or promoter is still controversial. The review highlights the different facets and peculiarities of this unconventional small RING-E3 ligase and its implication in tumorigenesis, invasion, neuroinflammation, and cancer metastasis. Keywords: Ring Finger Protein 11; HECT ligases; ubiquitination 1. -

Anti-COPE Picoband Antibody Catalog # ABO13033
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Anti-COPE Picoband Antibody Catalog # ABO13033 Specification Anti-COPE Picoband Antibody - Product Information Application IHC Primary Accession O14579 Host Rabbit Reactivity Human, Mouse, Rat Clonality Polyclonal Format Lyophilized Description Rabbit IgG polyclonal antibody for Coatomer subunit epsilon(COPE) detection. Tested with WB, IHC-P in Human;Mouse;Rat. Figure 4. IHC analysis of COPE using Reconstitution anti-COPE antibody (ABO13033). Add 0.2ml of distilled water will yield a concentration of 500ug/ml. Anti-COPE Picoband Antibody - Additional Information Gene ID 11316 Other Names Coatomer subunit epsilon, Epsilon-coat protein, Epsilon-COP, COPE Calculated MW 34482 MW KDa Application Details Immunohistochemistry(Paraffin-embedded Section), 0.5-1 µg/ml, Human, Mouse, Rat, By Heat<br> Western blot, 0.1-0.5 µg/ml, Human, Mouse, Rat, <br> <br> Subcellular Localization Cytoplasm . Golgi apparatus membrane ; Peripheral membrane protein ; Cytoplasmic side . Cytoplasmic vesicle, COPI-coated vesicle membrane ; Peripheral membrane protein ; Cytoplasmic side . The coatomer is cytoplasmic or polymerized on the cytoplasmic side of the Golgi, as well as on the vesicles/buds originating from it. Page 1/3 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Contents Each vial contains 5mg BSA, 0.9mg NaCl, 0.2mg Na2HPO4, 0.05mg NaN3. Immunogen E. coli-derived human COPE recombinant protein (Position: E80-A308). Human COPE shares 89.5% amino acid (aa) sequence identity with mouse COPE. Purification Immunogen affinity purified. Cross Reactivity No cross reactivity with other proteins. -

S100A8 Transported by SEC23A Inhibits Metastatic Colonization Via
Sun et al. Cell Death and Disease (2020) 11:650 https://doi.org/10.1038/s41419-020-02835-w Cell Death & Disease ARTICLE Open Access S100A8 transported by SEC23A inhibits metastatic colonization via autocrine activation of autophagy Zhiwei Sun1,2, Bin Zeng1,2,DoudouLiu1,2, Qiting Zhao1,2,JianyuWang1,2 and H. Rosie Xing2,3 Abstract Metastasis is the main cause of failure of cancer treatment. Metastatic colonization is regarded the most rate- limiting step of metastasis and is subjected to regulation by a plethora of biological factors and processes. On one hand, regulation of metastatic colonization by autophagy appears to be stage- and context-dependent, whereas mechanistic characterization remains elusive. On the other hand, interactions between the tumor cells and their microenvironment in metastasis have long been appreciated, whether the secretome of tumor cells can effectively reshape the tumor microenvironment has not been elucidated mechanistically. In the present study, we have identified “SEC23A-S1008-BECLIN1-autophagy axis” in the autophagic regulation of metastatic colonization step, a mechanism that tumor cells can exploit autophagy to exert self-restrain for clonogenic proliferation before the favorable tumor microenvironment is established. Specifically, we employed a paired lung-derived oligometastatic cell line (OL) and the homologous polymetastatic cell line (POL) from human melanoma cell line M14 that differ in colonization efficiency. We show that S100A8 transported by SEC23A inhibits metastatic colonization via autocrine activation of autophagy. Furthermore, we verified the clinical relevance of our experimental findings by bioinformatics analysis of the expression of Sec23a and S100A8 and the clinical- pathological associations. We demonstrate that higher Sec23a and Atg5 expression levels appear to be protective factors and favorable diagnostic (TNM staging) and prognostic (overall survival) markers for skin cutaneous 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; melanoma (SKCM) and colon adenocarcinoma (COAD) patients. -

Site-1 Protease Deficiency Causes Human Skeletal Dysplasia Due to Defective Inter-Organelle Protein Trafficking
Site-1 protease deficiency causes human skeletal dysplasia due to defective inter-organelle protein trafficking Yuji Kondo, … , Patrick M. Gaffney, Lijun Xia JCI Insight. 2018;3(14):e121596. https://doi.org/10.1172/jci.insight.121596. Research Article Cell biology Genetics Graphical abstract Find the latest version: https://jci.me/121596/pdf RESEARCH ARTICLE Site-1 protease deficiency causes human skeletal dysplasia due to defective inter-organelle protein trafficking Yuji Kondo,1 Jianxin Fu,1,2 Hua Wang,3 Christopher Hoover,1,4 J. Michael McDaniel,1 Richard Steet,5 Debabrata Patra,6 Jianhua Song,1 Laura Pollard,7 Sara Cathey,7 Tadayuki Yago,1 Graham Wiley,8 Susan Macwana,8 Joel Guthridge,8 Samuel McGee,1 Shibo Li,3 Courtney Griffin,1 Koichi Furukawa,9 Judith A. James,8 Changgeng Ruan,2 Rodger P. McEver,1,4 Klaas J. Wierenga,3 Patrick M. Gaffney,8 and Lijun Xia1,2,4 1Cardiovascular Biology Research Program, Oklahoma Medical Research Foundation, Oklahoma City, Oklahoma, USA. 2Jiangsu Institute of Hematology, MOH Key Laboratory of Thrombosis and Hemostasis, Collaborative Innovation Center of Hematology, The First Affiliated Hospital of Soochow University, Suzhou, China. 3Department of Pediatrics and 4Department of Biochemistry and Molecular Biology, University of Oklahoma Health Sciences Center, Oklahoma City, Oklahoma, USA. 5Complex Carbohydrate Research Center, University of Georgia, Georgia, Athens, USA. 6Department of Orthopaedic Surgery, Washington University School of Medicine, St. Louis, Missouri, USA. 7Greenwood Genetic Center, Greenwood, South Carolina, USA. 8Division of Genomics and Data Sciences, Arthritis and Clinical Immunology Program, Oklahoma Medical Research Foundation, Oklahoma City, Oklahoma, USA. 9Department of Biochemistry II, Nagoya University Graduate School of Medicine, Nagoya, Japan. -

ER-To-Golgi Trafficking and Its Implication in Neurological Diseases
cells Review ER-to-Golgi Trafficking and Its Implication in Neurological Diseases 1,2, 1,2 1,2, Bo Wang y, Katherine R. Stanford and Mondira Kundu * 1 Department of Pathology, St. Jude Children’s Research Hospital, Memphis, TN 38105, USA; [email protected] (B.W.); [email protected] (K.R.S.) 2 Department of Cell and Molecular Biology, St. Jude Children’s Research Hospital, Memphis, TN 38105, USA * Correspondence: [email protected]; Tel.: +1-901-595-6048 Present address: School of Life Sciences, Xiamen University, Xiamen 361102, China. y Received: 21 November 2019; Accepted: 7 February 2020; Published: 11 February 2020 Abstract: Membrane and secretory proteins are essential for almost every aspect of cellular function. These proteins are incorporated into ER-derived carriers and transported to the Golgi before being sorted for delivery to their final destination. Although ER-to-Golgi trafficking is highly conserved among eukaryotes, several layers of complexity have been added to meet the increased demands of complex cell types in metazoans. The specialized morphology of neurons and the necessity for precise spatiotemporal control over membrane and secretory protein localization and function make them particularly vulnerable to defects in trafficking. This review summarizes the general mechanisms involved in ER-to-Golgi trafficking and highlights mutations in genes affecting this process, which are associated with neurological diseases in humans. Keywords: COPII trafficking; endoplasmic reticulum; Golgi apparatus; neurological disease 1. Overview Approximately one-third of all proteins encoded by the mammalian genome are exported from the endoplasmic reticulum (ER) and transported to the Golgi apparatus, where they are sorted for delivery to their final destination in membrane compartments or secretory vesicles [1]. -

Shiga Toxin E. Coli Detection Differentiation Implications for Food
SL440 Shiga Toxin-Producing Escherichia coli: Detection, Differentiation, and Implications for Food Safety1 William J. Zaragoza, Max Teplitski, and Clifton K. Fagerquist2 Introduction for the effective detection of the Shiga-toxin-producing pathogens in a variety of food matrices. Shiga toxin is a protein found within the genome of a type of virus called a bacteriophage. These bacteriophages can There are two types of Shiga toxins—Shiga toxin 1 (Stx1) integrate into the genomes of the bacterium E. coli, giving and Shiga toxin 2 (Stx2). Stx1 is composed of several rise to Shiga toxin-producing E. coli (STEC). Even though subtypes knows as Stx1a, Stx1c, and Stx1d. Stx2 has seven most E. coli are benign or even beneficial (“commensal”) subtypes: a–g. Strains carrying all of the Stx1 subtypes affect members of our gut microbial communities, strains of E. humans, though they are less potent than that of Stx2. Stx2 coli carrying Shiga-toxin encoding genes (as well as other subtypes a,c, and d are frequently associated with human virulence determinants) are highly pathogenic in humans illness, while the other subtypes affect different animals. and other animals. When mammals ingest these bacteria, Subtypes Stx2b and Stx2e affect neonatal piglets, while STECs can undergo phage-driven lysis and deliver these target hosts for Stx2f and Stx2g subtypes are not currently toxins to mammalian guts. The Shiga toxin consists of an known (Fuller et al. 2011). Stx2f was originally isolated A subunit and 5 identical B subunits. The B subunits are from feral pigeons (Schmidt et al. 2000), and Stx2g was involved in binding to gut epithelial cells. -

In Vivo Mapping of a GPCR Interactome Using Knockin Mice
In vivo mapping of a GPCR interactome using knockin mice Jade Degrandmaisona,b,c,d,e,1, Khaled Abdallahb,c,d,1, Véronique Blaisb,c,d, Samuel Géniera,c,d, Marie-Pier Lalumièrea,c,d, Francis Bergeronb,c,d,e, Catherine M. Cahillf,g,h, Jim Boulterf,g,h, Christine L. Lavoieb,c,d,i, Jean-Luc Parenta,c,d,i,2, and Louis Gendronb,c,d,i,j,k,2 aDépartement de Médecine, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; bDépartement de Pharmacologie–Physiologie, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; cFaculté de Médecine et des Sciences de la Santé, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; dCentre de Recherche du Centre Hospitalier Universitaire de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; eQuebec Network of Junior Pain Investigators, Sherbrooke, QC J1H 5N4, Canada; fDepartment of Psychiatry and Biobehavioral Sciences, University of California, Los Angeles, CA 90095; gSemel Institute for Neuroscience and Human Behavior, University of California, Los Angeles, CA 90095; hShirley and Stefan Hatos Center for Neuropharmacology, University of California, Los Angeles, CA 90095; iInstitut de Pharmacologie de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; jDépartement d’Anesthésiologie, Université de Sherbrooke, Sherbrooke, QC J1H 5N4, Canada; and kQuebec Pain Research Network, Sherbrooke, QC J1H 5N4, Canada Edited by Brian K. Kobilka, Stanford University School of Medicine, Stanford, CA, and approved April 9, 2020 (received for review October 16, 2019) With over 30% of current medications targeting this family of attenuates pain hypersensitivities in several chronic pain models proteins, G-protein–coupled receptors (GPCRs) remain invaluable including neuropathic, inflammatory, diabetic, and cancer pain therapeutic targets. -

1 the TRAPP Complex Mediates Secretion Arrest Induced by Stress Granule Assembly Francesca Zappa1, Cathal Wilson1, Giusepp
bioRxiv preprint doi: https://doi.org/10.1101/528380; this version posted February 5, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. The TRAPP complex mediates secretion arrest induced by stress granule assembly Francesca Zappa1, Cathal Wilson1, Giuseppe Di Tullio1, Michele Santoro1, Piero Pucci2, Maria Monti2, Davide D’Amico1, Sandra Pisonero Vaquero1, Rossella De Cegli1, Alessia Romano1, Moin A. Saleem3, Elena Polishchuk1, Mario Failli1, Laura Giaquinto1, Maria Antonietta De Matteis1, 2 1 Telethon Institute of Genetics and Medicine, Pozzuoli (Naples), Italy 2 Federico II University, Naples, Italy 3 Bristol Renal, Bristol Medical School, University of Bristol, UK Correspondence to: [email protected], [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/528380; this version posted February 5, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. The TRAnsport-Protein-Particle (TRAPP) complex controls multiple membrane trafficking steps and is thus strategically positioned to mediate cell adaptation to diverse environmental conditions, including acute stress. We have identified TRAPP as a key component of a branch of the integrated stress response that impinges on the early secretory pathway. TRAPP associates with and drives the recruitment of the COPII coat to stress granules (SGs) leading to vesiculation of the Golgi complex and an arrest of ER export. -

The Popeye Domain Containing Genes and Their Function in Striated Muscle
Journal of Cardiovascular Development and Disease Review The Popeye Domain Containing Genes and Their Function in Striated Muscle Roland F. R. Schindler 1, Chiara Scotton 2, Vanessa French 1, Alessandra Ferlini 2 and Thomas Brand 1,* 1 Developmental Dynamics, Harefield Heart Science Centre, National Heart and Lung Institute, Imperial College London, Hill End Road, Harefield UB9 6JH, UK; [email protected] (R.F.R.S.); [email protected] (V.F.) 2 Department of Medical Sciences, Medical Genetics Unit, University of Ferrara, Ferrara 44121, Italy; [email protected] (C.S.); fl[email protected] (A.F.) * Correspondence: [email protected]; Tel.: +44-189-582-8900 Academic Editors: Robert E. Poelmann and Monique R.M. Jongbloed Received: 26 April 2016; Accepted: 13 June 2016; Published: 15 June 2016 Abstract: The Popeye domain containing (POPDC) genes encode a novel class of cAMP effector proteins, which are abundantly expressed in heart and skeletal muscle. Here, we will review their role in striated muscle as deduced from work in cell and animal models and the recent analysis of patients carrying a missense mutation in POPDC1. Evidence suggests that POPDC proteins control membrane trafficking of interacting proteins. Furthermore, we will discuss the current catalogue of established protein-protein interactions. In recent years, the number of POPDC-interacting proteins has been rising and currently includes ion channels (TREK-1), sarcolemma-associated proteins serving functions in mechanical stability (dystrophin), compartmentalization (caveolin 3), scaffolding (ZO-1), trafficking (NDRG4, VAMP2/3) and repair (dysferlin) or acting as a guanine nucleotide exchange factor for Rho-family GTPases (GEFT). -
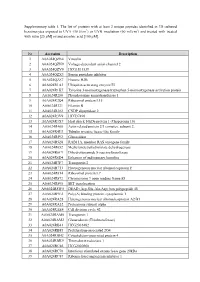
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31