Genetic Variants Within the MHC Region Are Associated with Immune Responsiveness to Childhood Vaccinations☆
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Variation Across the Human Olfactory Receptor Repertoire Alters Odor Perception
bioRxiv preprint doi: https://doi.org/10.1101/212431; this version posted November 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Genetic variation across the human olfactory receptor repertoire alters odor perception Casey Trimmer1,*, Andreas Keller2, Nicolle R. Murphy1, Lindsey L. Snyder1, Jason R. Willer3, Maira Nagai4,5, Nicholas Katsanis3, Leslie B. Vosshall2,6,7, Hiroaki Matsunami4,8, and Joel D. Mainland1,9 1Monell Chemical Senses Center, Philadelphia, Pennsylvania, USA 2Laboratory of Neurogenetics and Behavior, The Rockefeller University, New York, New York, USA 3Center for Human Disease Modeling, Duke University Medical Center, Durham, North Carolina, USA 4Department of Molecular Genetics and Microbiology, Duke University Medical Center, Durham, North Carolina, USA 5Department of Biochemistry, University of Sao Paulo, Sao Paulo, Brazil 6Howard Hughes Medical Institute, New York, New York, USA 7Kavli Neural Systems Institute, New York, New York, USA 8Department of Neurobiology and Duke Institute for Brain Sciences, Duke University Medical Center, Durham, North Carolina, USA 9Department of Neuroscience, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA *[email protected] ABSTRACT The human olfactory receptor repertoire is characterized by an abundance of genetic variation that affects receptor response, but the perceptual effects of this variation are unclear. To address this issue, we sequenced the OR repertoire in 332 individuals and examined the relationship between genetic variation and 276 olfactory phenotypes, including the perceived intensity and pleasantness of 68 odorants at two concentrations, detection thresholds of three odorants, and general olfactory acuity. -

OR2W1 (Human) Recombinant Protein
OR2W1 (Human) Recombinant Entrez GeneID: 26692 Protein Gene Symbol: OR2W1 Catalog Number: H00026692-G01 Gene Alias: MGC119162, MGC119163, MGC119165, hs6M1-15 Regulation Status: For research use only (RUO) Gene Summary: Olfactory receptors interact with Product Description: Human OR2W1 full-length ORF odorant molecules in the nose, to initiate a neuronal (NP_112165.1) recombinant protein without tag. response that triggers the perception of a smell. The olfactory receptor proteins are members of a large family Sequence: of G-protein-coupled receptors (GPCR) arising from MDQSNYSSLHGFILLGFSNHPKMEMILSGVVAIFYLITL single coding-exon genes. Olfactory receptors share a VGNTAIILASLLDSQLHTPMYFFLRNLSFLDLCFTTSIIP 7-transmembrane domain structure with many QMLVNLWGPDKTISYVGCIIQLYVYMWLGSVECLLLAV neurotransmitter and hormone receptors and are MSYDRFTAICKPLHYFVVMNPHLCLKMIIMIWSISLANS responsible for the recognition and G protein-mediated VVLCTLTLNLPTCGNNILDHFLCELPALVKIACVDTTTV transduction of odorant signals. The olfactory receptor EMSVFALGIIIVLTPLILILISYGYIAKAVLRTKSKASQRKA gene family is the largest in the genome. The MNTCGSHLTVVSMFYGTIIYMYLQPGNRASKDQGKFL nomenclature assigned to the olfactory receptor genes TLFYTVITPSLNPLIYTLRNKDMKDALKKLMRFHHKSTK and proteins for this organism is independent of other IKRNCKS organisms. [provided by RefSeq] Host: Wheat Germ (in vitro) Theoretical MW (kDa): 36.1 Applications: AP (See our web site product page for detailed applications information) Protocols: See our web site at http://www.abnova.com/support/protocols.asp or product page for detailed protocols Form: Liquid Preparation Method: in vitro wheat germ expression system with proprietary liposome technology Purification: None Recommend Usage: Heating may cause protein aggregation. Please do not heat this product before electrophoresis. Storage Buffer: 25 mM Tris-HCl of pH8.0 containing 2% glycerol. Storage Instruction: Store at -80°C. Aliquot to avoid repeated freezing and thawing. Page 1/1 Powered by TCPDF (www.tcpdf.org). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Extracting Structured Genotype Information from Free-Text HLA
J Korean Med Sci. 2020 Mar 30;35(12):e78 https://doi.org/10.3346/jkms.2020.35.e78 eISSN 1598-6357·pISSN 1011-8934 Original Article Extracting Structured Genotype Medical Informatics Information from Free-Text HLA Reports Using a Rule-Based Approach Kye Hwa Lee ,1 Hyo Jung Kim ,2 Yi-Jun Kim ,1 Ju Han Kim ,2 and Eun Young Song 3 1Center for Precision Medicine, Seoul National University Hospital, Seoul, Korea 2Division of Biomedical Informatics, Seoul National University Biomedical Informatics and Systems Biomedical Informatics Research Center, Seoul National University College of Medicine, Seoul, Korea 3Department of Laboratory Medicine, Seoul National University College of Medicine, Seoul, Korea Received: May 16, 2019 ABSTRACT Accepted: Jan 29, 2020 Address for Correspondence: Background: Human leukocyte antigen (HLA) typing is important for transplant patients to Eun Young Song, MD, PhD prevent a severe mismatch reaction, and the result can also support the diagnosis of various Division of Biomedical Informatics, Seoul disease or prediction of drug side effects. However, such secondary applications of HLA National University Biomedical Informatics (SNUBI) and Systems Biomedical Informatics typing results are limited because they are typically provided in free-text format or PDFs on Research Center, Seoul National University electronic medical records. We here propose a method to convert HLA genotype information College of Medicine, 103 Daehak-ro, stored in an unstructured format into a reusable structured format by extracting serotype/ Jongno-gu, Seoul 03080, Korea. allele information. E-mail: [email protected] Methods: We queried HLA typing reports from the clinical data warehouse of Seoul National Kye Hwa Lee, MD, PhD University Hospital (SUPPREME) from 2000 to 2018 as a rule-development data set (64,024 Center for Precision Medicine, Seoul National reports) and from the most recent year (6,181 reports) as a test set. -

Kras, Braf, Erbb2, Tp53
Supplementary Table 1. SBT-EOC cases used for molecular analyses Fresh Frozen Tissue FFPE Tissue Tumour HRM Case ID PCR-Seq HRM component Oncomap (KRAS, BRAF, CNV GE CNV IHC (NRAS) (TP53 Only) ERBB2, TP53) Paired Cases 15043 SBT & INV √ √ √ √ √ √ 65662 SBT & INV √ √ √ √ √ 65661 SBT & INV √ √ √ √ √ √ 9128 SBT & INV √ √ √ √ √ 2044 SBT & INV √ √ √ √ √ 3960 SBT & INV √ √ √ √ √ 5899 SBT & INV √ √ √ √ √ 65663 SBT & INV √ √ √ Inv √ √ √ 65666 SBT √ Inv √ √ √ √ √ 65664 SBT √ √ √ Inv √ √ √ √ √ 15060 SBT √ √ √ Inv √ √ √ √ √ 65665 SBT √ √ √ Inv √ √ √ √ √ 65668 SBT √ √ √ Inv √ √ √ √ 65667 SBT √ √ √ Inv √ √ √ √ 15018 SBT √ Inv √ √ √ √ 15071 SBT √ Inv √ √ √ √ 65670 SBT √ Inv √ √ √ 8982 SBT √ Unpaired Cases 15046 Inv √ √ √ 15014 Inv √ √ √ 65671 Inv √ √ √ 65672 SBT √ √ √ 65673 Inv √ √ √ 65674 Inv √ √ √ 65675 Inv √ √ √ 65676 Inv √ √ √ 65677 Inv √ √ √ 65678 Inv √ √ √ 65679 Inv √ √ √ Fresh Frozen Tissue FFPE Tissue Tumour HRM Case ID PCR-Seq HRM component Oncomap (KRAS, BRAF, CNV GE CNV IHC (NRAS) (TP53 Only) ERBB2, TP53) 65680 Inv √ √ √ 5349 Inv √ √ √ 7957 Inv √ √ √ 8395 Inv √ √ √ 8390 Inv √ √ 2110 Inv √ √ 6328 Inv √ √ 9125 Inv √ √ 9221 Inv √ √ 10701 Inv √ √ 1072 Inv √ √ 11266 Inv √ √ 11368 Inv √ √ √ 12237 Inv √ √ 2064 Inv √ √ 3539 Inv √ √ 2189 Inv √ √ √ 5711 Inv √ √ 6251 Inv √ √ 6582 Inv √ √ 7200 SBT √ √ √ 8633 Inv √ √ √ 9579 Inv √ √ 10740 Inv √ √ 11766 Inv √ √ 3958 Inv √ √ 4723 Inv √ √ 6244 Inv √ √ √ 7716 Inv √ √ SBT-EOC, serous carcinoma with adjacent borderline regions; SBT, serous borderline component of SBT- EOC; Inv, invasive component of SBT-EOC; -

Technologies Available for Licensing
Technologies Available for Licensing Office of Innovation and Industry Alliances www.moffittip.com IMMUNOTHERAPIES CD40: CD40 Agonists for Improved ex vivo TIL Manufacturing 21MA018N T cell: T cells expressing anti-CD3 antibodies autoactivate and decrease expression of T cell receptors to treat GVHD, or make T cells suitable for off-the- 21MA013N shelf treatment of allogeneic subjects ITAM: Human ITAM Mutated Variants For Better Intracellular Signaling Domains 21MA008N For Gamma-Delta CAR T-Cell Activation ESR1: Cancer Vaccine Using Novel ESR1 Derived Peptides For Neoantigen 20MB062N Therapy NOTCH: Single-domain antibodies (nanobodies) targeting the notch ligand DLL4 to 20MB053N Disrupt the interaction of DLL4 and Notch1 for GVHD CD33 CD123 NKG2DL: Bispecific Gamma Delta CAR-T Cells that Recognize 20MB050N CD33, CD123 and NKG2D Ligands for the Treatment of Acute Myeloid Leukemia OR2H1 or OR5V1: Olfactory Receptor Targeting Chimeric Antigen Receptor 20MA027 Expressing T cell (CAR-T) for Solid Tumors HER-2: Combination Therapy of a HER2-DC1 Dendritic Cell Cancer Vaccine and a 20MA016N Probiotic TILs: 12 Chemokine Gene Expression Signature to Increase the Efficacy of 20MA013N Manufacturing Tumor Infiltrating Lymphocytes PERK, IRE1: Method of Enhancing Immunotherapy Using ER Stress Pathway 20MA005 Inhibitors PGC-1α: CAR T Cells Engineered to Express PGC-1 alpha Demonstrate Enhanced 19MB066 Metabolic Fitness PGC-1α: N-Terminal Mutant PGC-1α Overexpression Enhances Metabolic Fitness 19MB066T2 Reducing CAR-T Exhaustion while Maintaining Proliferative Capacity TILs: Fucose increases tumor cell HLA-DRB1 expression increasing CD4+ T-cell 19MB049N activation with synergistic tumor killing activity with anti-PD1 checkpoint inhibitors Antibodies: Fully Human Anti-BDNF Antibodies 19MB048N Antibodies: Fully Human Anti-TSPAN7 Antibodies 19MB047N T-Bet: T-Bet Transcription Factor Armed CAR-T Cells Maintain Memory 19MA035N Phenotypes and Rescue CD4 Cells Leading to Increased Persistence Haskell Adler PhD MBA CLP Charlie Shaw PhD Praba Soundararajan PhD Sr. -

Whole Exome Sequencing in Families at High Risk for Hodgkin Lymphoma: Identification of a Predisposing Mutation in the KDR Gene
Hodgkin Lymphoma SUPPLEMENTARY APPENDIX Whole exome sequencing in families at high risk for Hodgkin lymphoma: identification of a predisposing mutation in the KDR gene Melissa Rotunno, 1 Mary L. McMaster, 1 Joseph Boland, 2 Sara Bass, 2 Xijun Zhang, 2 Laurie Burdett, 2 Belynda Hicks, 2 Sarangan Ravichandran, 3 Brian T. Luke, 3 Meredith Yeager, 2 Laura Fontaine, 4 Paula L. Hyland, 1 Alisa M. Goldstein, 1 NCI DCEG Cancer Sequencing Working Group, NCI DCEG Cancer Genomics Research Laboratory, Stephen J. Chanock, 5 Neil E. Caporaso, 1 Margaret A. Tucker, 6 and Lynn R. Goldin 1 1Genetic Epidemiology Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 2Cancer Genomics Research Laboratory, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 3Ad - vanced Biomedical Computing Center, Leidos Biomedical Research Inc.; Frederick National Laboratory for Cancer Research, Frederick, MD; 4Westat, Inc., Rockville MD; 5Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; and 6Human Genetics Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD, USA ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2015.135475 Received: August 19, 2015. Accepted: January 7, 2016. Pre-published: June 13, 2016. Correspondence: [email protected] Supplemental Author Information: NCI DCEG Cancer Sequencing Working Group: Mark H. Greene, Allan Hildesheim, Nan Hu, Maria Theresa Landi, Jennifer Loud, Phuong Mai, Lisa Mirabello, Lindsay Morton, Dilys Parry, Anand Pathak, Douglas R. Stewart, Philip R. Taylor, Geoffrey S. Tobias, Xiaohong R. Yang, Guoqin Yu NCI DCEG Cancer Genomics Research Laboratory: Salma Chowdhury, Michael Cullen, Casey Dagnall, Herbert Higson, Amy A. -

Cluster-Specific Gene Markers Enhance Shigella and Enteroinvasive Escherichia Coli in Silico Serotyping Xiaomei Zhang1, Michael
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.30.428723; this version posted February 1, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Cluster-specific gene markers enhance Shigella and Enteroinvasive Escherichia coli in 2 silico serotyping 3 4 Xiaomei Zhang1, Michael Payne1, Thanh Nguyen1, Sandeep Kaur1, Ruiting Lan1* 5 6 1School of Biotechnology and Biomolecular Sciences, University of New South Wales, Sydney, 7 New South Wales, Australia 8 9 10 11 *Corresponding Author 12 Email: [email protected] 13 Phone: 61-2-9385 2095 14 Fax: 61-2-9385 1483 15 16 Keywords: Phylogenetic clusters, cluster-specific gene markers, Shigella/EIEC serotyping 17 Running title: Shigella EIEC Cluster Enhanced Serotyping 18 Repositories: Raw sequence data are available from NCBI under the BioProject number 19 PRJNA692536. 20 21 22 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.01.30.428723; this version posted February 1, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 23 Abstract 24 Shigella and enteroinvasive Escherichia coli (EIEC) cause human bacillary dysentery with 25 similar invasion mechanisms and share similar physiological, biochemical and genetic 26 characteristics. The ability to differentiate Shigella and EIEC from each other is important for 27 clinical diagnostic and epidemiologic investigations. The existing genetic signatures may not 28 discriminate between Shigella and EIEC. However, phylogenetically, Shigella and EIEC 29 strains are composed of multiple clusters and are different forms of E. -
European Patent Office of Opposition to That Patent, in Accordance with the Implementing Regulations
(19) TZZ Z_T (11) EP 2 884 280 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: G01N 33/566 (2006.01) 09.05.2018 Bulletin 2018/19 (21) Application number: 13197310.9 (22) Date of filing: 15.12.2013 (54) Method for evaluating the scent performance of perfumes and perfume mixtures Verfahren zur Bewertung des Duftverhaltens von Duftstoffen und Duftstoffmischungen Procédé d’evaluation de senteur performance du parfums et mixtures de parfums (84) Designated Contracting States: (56) References cited: AL AT BE BG CH CY CZ DE DK EE ES FI FR GB WO-A2-03/091388 GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR • BAGHAEI KAVEH A: "Deorphanization of human olfactory receptors by luciferase and Ca-imaging (43) Date of publication of application: methods.",METHODS IN MOLECULAR BIOLOGY 17.06.2015 Bulletin 2015/25 (CLIFTON, N.J.) 2013, vol. 1003, 19 June 2013 (2013-06-19), pages229-238, XP008168583, ISSN: (73) Proprietor: Symrise AG 1940-6029 37603 Holzminden (DE) • KAVEH BAGHAEI ET AL: "Olfactory receptors coded by segregating pseudo genes and (72) Inventors: odorants with known specific anosmia.", 33RD • Hatt, Hanns ANNUAL MEETING OF THE ASSOCIATION FOR 44789 Bochum (DE) CHEMORECEPTION, 1 April 2011 (2011-04-01), • Gisselmann, Günter XP055111507, 58456 Witten (DE) • TOUHARA ET AL: "Deorphanizing vertebrate • Ashtibaghaei, Kaveh olfactory receptors: Recent advances in 44801 Bochum (DE) odorant-response assays", NEUROCHEMISTRY • Panten, Johannes INTERNATIONAL, PERGAMON PRESS, 37671 Höxter (DE) OXFORD, GB, vol. -

Genetics and Extracellular Vesicles of Pediatrics Sleep Disordered Breathing and Epilepsy
International Journal of Molecular Sciences Review Genetics and Extracellular Vesicles of Pediatrics Sleep Disordered Breathing and Epilepsy Abdelnaby Khalyfa 1,2,* and David Sanz-Rubio 2 1 Department of Pediatrics, Section of Sleep Medicine, The University of Chicago, Chicago, IL 60637, USA 2 Department of Child Health and the Child Health Research Institute, University of Missouri School of Medicine, Columbia, MO 65201, USA; [email protected] * Correspondence: [email protected] Received: 20 August 2019; Accepted: 28 October 2019; Published: 4 November 2019 Abstract: Sleep remains one of the least understood phenomena in biology, and sleep disturbances are one of the most common behavioral problems in childhood. The etiology of sleep disorders is complex and involves both genetic and environmental factors. Epilepsy is the most popular childhood neurological condition and is characterized by an enduring predisposition to generate epileptic seizures, and the neurobiological, cognitive, psychological, and social consequences of this condition. Sleep and epilepsy are interrelated, and the importance of sleep in epilepsy is less known. The state of sleep also influences whether a seizure will occur at a given time, and this differs considerably for various epilepsy syndromes. The development of epilepsy has been associated with single or multiple gene variants. The genetics of epilepsy is complex and disorders exhibit significant genetic heterogeneity and variability in the expressivity of seizures. Phenobarbital (PhB) is the most widely used antiepileptic drug. With its principal mechanism of action to prolong the opening time of the γ-aminobutyric acid (GABA)-A receptor-associated chloride channel, it enhances chloride anion influx into neurons, with subsequent hyperpolarization, thereby reducing excitability. -
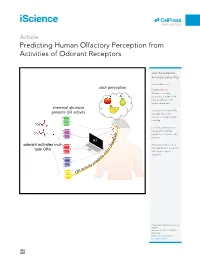
Predicting Human Olfactory Perception from Activities of Odorant Receptors
iScience ll OPEN ACCESS Article Predicting Human Olfactory Perception from Activities of Odorant Receptors Joel Kowalewski, Anandasankar Ray [email protected] odor perception HIGHLIGHTS Machine learning predicted activity of 34 human ORs for ~0.5 million chemicals chemical structure Activities of human ORs predicts OR activity could predict odor character using machine learning Few OR activities were needed to optimize r predictions of each odor e t c percept a AI r a odorant activates mul- h Behavior predictions in c Drosophila also need few r tiple ORs o olfactory receptor d o activities ts ic ed pr ity tiv ac OR Kowalewski & Ray, iScience 23, 101361 August 21, 2020 ª 2020 The Author(s). https://doi.org/10.1016/ j.isci.2020.101361 iScience ll OPEN ACCESS Article Predicting Human Olfactory Perception from Activities of Odorant Receptors Joel Kowalewski1 and Anandasankar Ray1,2,3,* SUMMARY Odor perception in humans is initiated by activation of odorant receptors (ORs) in the nose. However, the ORs linked to specific olfactory percepts are unknown, unlike in vision or taste where receptors are linked to perception of different colors and tastes. The large family of ORs (~400) and multiple receptors activated by an odorant present serious challenges. Here, we first use machine learning to screen ~0.5 million compounds for new ligands and identify enriched structural motifs for ligands of 34 human ORs. We next demonstrate that the activity of ORs successfully predicts many of the 146 different perceptual qualities of chem- icals. Although chemical features have been used to model odor percepts, we show that biologically relevant OR activity is often superior.