RT² Profiler PCR Array (Rotor-Gene® Format) Rat Gap Junctions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Adenylate Cyclase 3: a New Target for Anti‐Obesity Drug Development
obesity reviews doi: 10.1111/obr.12430 Obesity Pharmacotherapy Adenylate cyclase 3: a new target for anti-obesity drug development L. Wu,1 C. Shen,2 M. Seed Ahmed,3 C.-G. Östenson4 and H. F. Gu5,6 1Jiangsu Key Laboratory of Drug Screening, Summary China Pharmaceutical University, Nanjing Obesity has become epidemic worldwide, and abdominal obesity has a negative im- 210009, China, 2Department of Epidemiology, pact on health. Current treatment options on obesity, however, still remain limited. School of Public Health, Nanjing Medical It is then of importance to find a new target for anti-obesity drug development University, Nanjing 211166, China, 3Unit for based upon recent molecular studies in obesity. Adenylate cyclase 3 (ADCY3) is Medical Education, Centre for Learning and the third member of adenylyl cyclase family and catalyses the synthesis of cAMP Knowledge, Department of Learning, from ATP. Genetic studies with candidate gene and genome-wide association study Informatics, Management and Ethics, approaches have demonstrated that ADCY3 genetic polymorphisms are associated Karolinska Institutet, Stockholm 17177, with obesity in European and Chinese populations. Epigenetic studies have indi- Sweden, 4Rolf Luft Center for Diabetes cated that increased DNA methylation levels in the ADCY3 gene are involved in Research and Endocrinology, Department of the pathogenesis of obesity. Furthermore, biological analyses with animal models Molecular Medicine and Surgery, Karolinska have implicated that ADCY3 dysfunction resulted in increased body weight and Institutet, Karolinska University Hospital, Solna, fat mass, while reduction of body weight is partially explained by ADCY3 activa- Stockholm 17176, Sweden, 5Department of tion. In this review, we describe genomic and biological features of ADCY3, sum- Clinical Science, Intervention and marize genetic and epigenetic association studies of the ADCY3 gene with obesity Technologies, Karolinska Institutet, Karolinska and discuss dysfunction and activation of ADCY3. -

Supplemental Table S1
Entrez Gene Symbol Gene Name Affymetrix EST Glomchip SAGE Stanford Literature HPA confirmed Gene ID Profiling profiling Profiling Profiling array profiling confirmed 1 2 A2M alpha-2-macroglobulin 0 0 0 1 0 2 10347 ABCA7 ATP-binding cassette, sub-family A (ABC1), member 7 1 0 0 0 0 3 10350 ABCA9 ATP-binding cassette, sub-family A (ABC1), member 9 1 0 0 0 0 4 10057 ABCC5 ATP-binding cassette, sub-family C (CFTR/MRP), member 5 1 0 0 0 0 5 10060 ABCC9 ATP-binding cassette, sub-family C (CFTR/MRP), member 9 1 0 0 0 0 6 79575 ABHD8 abhydrolase domain containing 8 1 0 0 0 0 7 51225 ABI3 ABI gene family, member 3 1 0 1 0 0 8 29 ABR active BCR-related gene 1 0 0 0 0 9 25841 ABTB2 ankyrin repeat and BTB (POZ) domain containing 2 1 0 1 0 0 10 30 ACAA1 acetyl-Coenzyme A acyltransferase 1 (peroxisomal 3-oxoacyl-Coenzyme A thiol 0 1 0 0 0 11 43 ACHE acetylcholinesterase (Yt blood group) 1 0 0 0 0 12 58 ACTA1 actin, alpha 1, skeletal muscle 0 1 0 0 0 13 60 ACTB actin, beta 01000 1 14 71 ACTG1 actin, gamma 1 0 1 0 0 0 15 81 ACTN4 actinin, alpha 4 0 0 1 1 1 10700177 16 10096 ACTR3 ARP3 actin-related protein 3 homolog (yeast) 0 1 0 0 0 17 94 ACVRL1 activin A receptor type II-like 1 1 0 1 0 0 18 8038 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 1 0 0 0 0 19 8751 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 1 0 0 0 0 20 8728 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 1 0 0 0 0 21 81792 ADAMTS12 ADAM metallopeptidase with thrombospondin type 1 motif, 12 1 0 0 0 0 22 9507 ADAMTS4 ADAM metallopeptidase with thrombospondin type 1 -

Gene and Pathway-Based Second-Wave Analysis of Genome-Wide Association Studies
European Journal of Human Genetics (2010) 18, 111–117 & 2010 Macmillan Publishers Limited All rights reserved 1018-4813/10 $32.00 www.nature.com/ejhg ARTICLE Gene and pathway-based second-wave analysis of genome-wide association studies Gang Peng1, Li Luo2, Hoicheong Siu1, Yun Zhu1, Pengfei Hu1, Shengjun Hong1, Jinying Zhao3, Xiaodong Zhou4, John D Reveille4, Li Jin1, Christopher I Amos5 and Momiao Xiong*,2 Despite the great success of genome-wide association studies (GWAS) in identification of the common genetic variants associated with complex diseases, the current GWAS have focused on single-SNP analysis. However, single-SNP analysis often identifies only a few of the most significant SNPs that account for a small proportion of the genetic variants and offers only a limited understanding of complex diseases. To overcome these limitations, we propose gene and pathway-based association analysis as a new paradigm for GWAS. As a proof of concept, we performed a comprehensive gene and pathway-based association analysis of 13 published GWAS. Our results showed that the proposed new paradigm for GWAS not only identified the genes that include significant SNPs found by single-SNP analysis, but also detected new genes in which each single SNP conferred a small disease risk; however, their joint actions were implicated in the development of diseases. The results also showed that the new paradigm for GWAS was able to identify biologically meaningful pathways associated with the diseases, which were confirmed by a gene-set-rich analysis using gene expression -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Supplementary Materials
Supplementary Materials + - NUMB E2F2 PCBP2 CDKN1B MTOR AKT3 HOXA9 HNRNPA1 HNRNPA2B1 HNRNPA2B1 HNRNPK HNRNPA3 PCBP2 AICDA FLT3 SLAMF1 BIC CD34 TAL1 SPI1 GATA1 CD48 PIK3CG RUNX1 PIK3CD SLAMF1 CDKN2B CDKN2A CD34 RUNX1 E2F3 KMT2A RUNX1 T MIXL1 +++ +++ ++++ ++++ +++ 0 0 0 0 hematopoietic potential H1 H1 PB7 PB6 PB6 PB6.1 PB6.1 PB12.1 PB12.1 Figure S1. Unsupervised hierarchical clustering of hPSC-derived EBs according to the mRNA expression of hematopoietic lineage genes (microarray analysis). Hematopoietic-competent cells (H1, PB6.1, PB7) were separated from hematopoietic-deficient ones (PB6, PB12.1). In this experiment, all hPSCs were tested in duplicate, except PB7. Genes under-expressed or over-expressed in blood-deficient hPSCs are indicated in blue and red respectively (related to Table S1). 1 C) Mesoderm B) Endoderm + - KDR HAND1 GATA6 MEF2C DKK1 MSX1 GATA4 WNT3A GATA4 COL2A1 HNF1B ZFPM2 A) Ectoderm GATA4 GATA4 GSC GATA4 T ISL1 NCAM1 FOXH1 NCAM1 MESP1 CER1 WNT3A MIXL1 GATA4 PAX6 CDX2 T PAX6 SOX17 HBB NES GATA6 WT1 SOX1 FN1 ACTC1 ZIC1 FOXA2 MYF5 ZIC1 CXCR4 TBX5 PAX6 NCAM1 TBX20 PAX6 KRT18 DDX4 TUBB3 EPCAM TBX5 SOX2 KRT18 NKX2-5 NES AFP COL1A1 +++ +++ 0 0 0 0 ++++ +++ ++++ +++ +++ ++++ +++ ++++ 0 0 0 0 +++ +++ ++++ +++ ++++ 0 0 0 0 hematopoietic potential H1 H1 H1 H1 H1 H1 PB6 PB6 PB7 PB7 PB6 PB6 PB7 PB6 PB6 PB6.1 PB6.1 PB6.1 PB6.1 PB6.1 PB6.1 PB12.1 PB12.1 PB12.1 PB12.1 PB12.1 PB12.1 Figure S2. Unsupervised hierarchical clustering of hPSC-derived EBs according to the mRNA expression of germ layer differentiation genes (microarray analysis) Selected ectoderm (A), endoderm (B) and mesoderm (C) related genes differentially expressed between hematopoietic-competent (H1, PB6.1, PB7) and -deficient cells (PB6, PB12.1) are shown (related to Table S1). -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

(Adcy4) in Y1 ADRENOCORTICAL TUMOR CELLS
TRANSCRIPTIONAL REGULATION OF THE MOUSE ADENYLYL CYCLASE TYPE 4 (Adcy4) IN Y1 ADRENOCORTICAL TUMOR CELLS By Xianliang Rui A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Department of Pharmacology and Toxicology University of Toronto © Copyright by Xianliang Rui 2010 Library and Archives Bibliothèque et Canada Archives Canada Published Heritage Direction du Branch Patrimoine de l’édition 395 Wellington Street 395, rue Wellington Ottawa ON K1A 0N4 Ottawa ON K1A 0N4 Canada Canada Your file Votre référence ISBN: 978-0-494-67725-4 Our file Notre référence ISBN: 978-0-494-67725-4 NOTICE: AVIS: The author has granted a non- L’auteur a accordé une licence non exclusive exclusive license allowing Library and permettant à la Bibliothèque et Archives Archives Canada to reproduce, Canada de reproduire, publier, archiver, publish, archive, preserve, conserve, sauvegarder, conserver, transmettre au public communicate to the public by par télécommunication ou par l’Internet, prêter, telecommunication or on the Internet, distribuer et vendre des thèses partout dans le loan, distribute and sell theses monde, à des fins commerciales ou autres, sur worldwide, for commercial or non- support microforme, papier, électronique et/ou commercial purposes, in microform, autres formats. paper, electronic and/or any other formats. The author retains copyright L’auteur conserve la propriété du droit d’auteur ownership and moral rights in this et des droits moraux qui protège cette thèse. Ni thesis. Neither the thesis nor la thèse ni des extraits substantiels de celle-ci substantial extracts from it may be ne doivent être imprimés ou autrement printed or otherwise reproduced reproduits sans son autorisation. -

HHS Public Access Author Manuscript
HHS Public Access Author manuscript Author Manuscript Author ManuscriptBreast Cancer Author Manuscript Res Treat Author Manuscript . Author manuscript; available in PMC 2016 June 01. Published in final edited form as: Breast Cancer Res Treat. 2015 June ; 151(2): 453–463. doi:10.1007/s10549-015-3401-8. Body mass index associated with genome-wide methylation in breast tissue Brionna Y. Hair1, Zongli Xu2, Erin L. Kirk1, Sophia Harlid2, Rupninder Sandhu3, Whitney R. Robinson1,3, Michael C. Wu4, Andrew F. Olshan1, Kathleen Conway1,3, Jack A. Taylor2, and Melissa A. Troester1 1 Department of Epidemiology, University of North Carolina at Chapel Hill, CB #7435, 2101 McGavran-Greenberg Hall, Chapel Hill, NC 27599-7435, USA 2 Epidemiology Branch, and Epigenomics and Stem Cell Biology Laboratory, National Institute of Environmental Health Sciences (NIH), Research Triangle Park, NC, USA 3 Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC, USA 4 Fred Hutchinson Cancer Research Center, Seattle, WA, USA Abstract Gene expression studies indicate that body mass index (BMI) is associated with molecular pathways involved in inflammation, insulin-like growth factor activation, and other carcinogenic processes in breast tissue. The goal of this study was to determine whether BMI is associated with gene methylation in breast tissue and to identify pathways that are commonly methylated in association with high BMI. Epigenome-wide methylation profiles were determined using the Illumina HumanMethylation450 BeadChip array in the non-diseased breast tissue of 81 women undergoing breast surgery between 2009 and 2013 at the University of North Carolina Hospitals. Multivariable, robust linear regression was performed to identify methylation sites associated with BMI at a false discovery rate q value <0.05. -

Increased Plasma Levels of Adenylate Cyclase 8 and Camp Are Associated with Obesity and Type 2 Diabetes: Results from a Cross-Sectional Study
biology Article Increased Plasma Levels of Adenylate Cyclase 8 and cAMP Are Associated with Obesity and Type 2 Diabetes: Results from a Cross-Sectional Study 1, 2, , 2, 2 Samy M. Abdel-Halim y, Ashraf Al Madhoun * y , Rasheeba Nizam y, Motasem Melhem , Preethi Cherian 3, Irina Al-Khairi 3, Dania Haddad 2, Mohamed Abu-Farha 3 , Jehad Abubaker 3, Milad S. Bitar 4 and Fahd Al-Mulla 2,* 1 Department of Oncology, Karolinska University Hospital, 17177 Stockholm, Sweden; [email protected] 2 Department of Genetics and Bioinformatics, Dasman Diabetes Institute, Dasman 15462, Kuwait; [email protected] (R.N.); [email protected] (M.M.); [email protected] (D.H.) 3 Department of Biochemistry and Molecular Biology, Dasman Diabetes Institute, Dasman 15462, Kuwait; [email protected] (P.C.); [email protected] (I.A.-K.); [email protected] (M.A.-F.); [email protected] (J.A.) 4 Department of Pharmacology and Toxicology, Faculty of Medicine, Kuwait University, Jabriya 046302, Kuwait; [email protected] * Correspondence: [email protected] (A.A.M.); [email protected] (F.A.-M.); Tel.: +965-2224-2999 (ext. 2805) (A.A.M.); +965-6777-1040 (F.A.-M.) These authors equally contributed to this work. y Received: 5 July 2020; Accepted: 20 August 2020; Published: 24 August 2020 Abstract: Adenylate cyclases (ADCYs) catalyze the conversion of ATP to cAMP,an important co-factor in energy homeostasis. Giving ADCYs role in obesity, diabetes and inflammation, we questioned whether calcium-stimulated ADCY isoforms may be variably detectable in human plasma. -
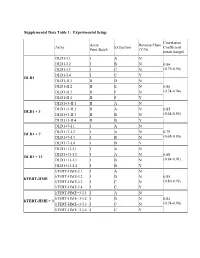
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Loss-Of-Function Variants in ADCY3 Increase Risk of Obesity and Type 2 Diabetes
BRIEF COMMUNICATION https://doi.org/10.1038/s41588-017-0022-7 Loss-of-function variants in ADCY3 increase risk of obesity and type 2 diabetes Niels Grarup 1, Ida Moltke 2, Mette K. Andersen 1, Maria Dalby2, Kristoffer Vitting-Seerup2,3, Timo Kern1, Yuvaraj Mahendran1, Emil Jørsboe 2, Christina V. L. Larsen4,5, Inger K. Dahl-Petersen4, Arthur Gilly6, Daniel Suveges6, George Dedoussis7, Eleftheria Zeggini 6, Oluf Pedersen1, Robin Andersson 2, Peter Bjerregaard4,5, Marit E. Jørgensen4,5,8*, Anders Albrechtsen 2* and Torben Hansen 1,9* We have identified a variant in ADCY3 (encoding adenylate carriers had type 2 diabetes (P = 7.8 × 10−5; Table 1), while one had cyclase 3) associated with markedly increased risk of obesity impaired fasting glucose and one had impaired glucose tolerance. and type 2 diabetes in the Greenlandic population. The variant Notably, the association with type 2 diabetes remained significant disrupts a splice acceptor site, and carriers have decreased after adjustment for BMI (P = 6.5 × 10−4), suggesting that it is not ADCY3 RNA expression. Additionally, we observe an enrich- simply mediated by increased BMI. The effects on BMI and type 2 ment of rare ADCY3 loss-of-function variants among indi- diabetes were also observed, although with smaller sizes, when data viduals with type 2 diabetes in trans-ancestry cohorts. These were analyzed according to an additive genetic model (Table 1). findings provide new information on disease etiology relevant However, when we compared the recessive and additive models for future treatment strategies. with the full genotype model, we rejected the additive model (BMI, Identification of homozygous loss-of-function mutations in P = 0.002; type 2 diabetes, P = 0.004) but not the recessive model humans may readily provide information about the biological (BMI, P = 0.17; type 2 diabetes, P = 0.095). -

Epigenetic Modifications to Cytosine and Alzheimer's Disease
University of Kentucky UKnowledge Theses and Dissertations--Chemistry Chemistry 2017 EPIGENETIC MODIFICATIONS TO CYTOSINE AND ALZHEIMER’S DISEASE: A QUANTITATIVE ANALYSIS OF POST-MORTEM TISSUE Elizabeth M. Ellison University of Kentucky, [email protected] Digital Object Identifier: https://doi.org/10.13023/ETD.2017.398 Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Ellison, Elizabeth M., "EPIGENETIC MODIFICATIONS TO CYTOSINE AND ALZHEIMER’S DISEASE: A QUANTITATIVE ANALYSIS OF POST-MORTEM TISSUE" (2017). Theses and Dissertations--Chemistry. 86. https://uknowledge.uky.edu/chemistry_etds/86 This Doctoral Dissertation is brought to you for free and open access by the Chemistry at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Chemistry by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained needed written permission statement(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine) which will be submitted to UKnowledge as Additional File. I hereby grant to The University of Kentucky and its agents the irrevocable, non-exclusive, and royalty-free license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known.