GUCY1A2 Polyclonal Antibody
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Supplementary Materials
Supplementary Materials + - NUMB E2F2 PCBP2 CDKN1B MTOR AKT3 HOXA9 HNRNPA1 HNRNPA2B1 HNRNPA2B1 HNRNPK HNRNPA3 PCBP2 AICDA FLT3 SLAMF1 BIC CD34 TAL1 SPI1 GATA1 CD48 PIK3CG RUNX1 PIK3CD SLAMF1 CDKN2B CDKN2A CD34 RUNX1 E2F3 KMT2A RUNX1 T MIXL1 +++ +++ ++++ ++++ +++ 0 0 0 0 hematopoietic potential H1 H1 PB7 PB6 PB6 PB6.1 PB6.1 PB12.1 PB12.1 Figure S1. Unsupervised hierarchical clustering of hPSC-derived EBs according to the mRNA expression of hematopoietic lineage genes (microarray analysis). Hematopoietic-competent cells (H1, PB6.1, PB7) were separated from hematopoietic-deficient ones (PB6, PB12.1). In this experiment, all hPSCs were tested in duplicate, except PB7. Genes under-expressed or over-expressed in blood-deficient hPSCs are indicated in blue and red respectively (related to Table S1). 1 C) Mesoderm B) Endoderm + - KDR HAND1 GATA6 MEF2C DKK1 MSX1 GATA4 WNT3A GATA4 COL2A1 HNF1B ZFPM2 A) Ectoderm GATA4 GATA4 GSC GATA4 T ISL1 NCAM1 FOXH1 NCAM1 MESP1 CER1 WNT3A MIXL1 GATA4 PAX6 CDX2 T PAX6 SOX17 HBB NES GATA6 WT1 SOX1 FN1 ACTC1 ZIC1 FOXA2 MYF5 ZIC1 CXCR4 TBX5 PAX6 NCAM1 TBX20 PAX6 KRT18 DDX4 TUBB3 EPCAM TBX5 SOX2 KRT18 NKX2-5 NES AFP COL1A1 +++ +++ 0 0 0 0 ++++ +++ ++++ +++ +++ ++++ +++ ++++ 0 0 0 0 +++ +++ ++++ +++ ++++ 0 0 0 0 hematopoietic potential H1 H1 H1 H1 H1 H1 PB6 PB6 PB7 PB7 PB6 PB6 PB7 PB6 PB6 PB6.1 PB6.1 PB6.1 PB6.1 PB6.1 PB6.1 PB12.1 PB12.1 PB12.1 PB12.1 PB12.1 PB12.1 Figure S2. Unsupervised hierarchical clustering of hPSC-derived EBs according to the mRNA expression of germ layer differentiation genes (microarray analysis) Selected ectoderm (A), endoderm (B) and mesoderm (C) related genes differentially expressed between hematopoietic-competent (H1, PB6.1, PB7) and -deficient cells (PB6, PB12.1) are shown (related to Table S1). -
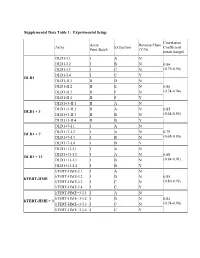
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

GSE50161, (C) GSE66354, (D) GSE74195 and (E) GSE86574
Figure S1. Boxplots of normalized samples in five datasets. (A) GSE25604, (B) GSE50161, (C) GSE66354, (D) GSE74195 and (E) GSE86574. The x‑axes indicate samples, and the y‑axes represent the expression of genes. Figure S2. Volanco plots of DEGs in five datasets. (A) GSE25604, (B) GSE50161, (C) GSE66354, (D) GSE74195 and (E) GSE86574. Red nodes represent upregulated DEGs and green nodes indicate downregulated DEGs. Cut‑off criteria were P<0.05 and |log2 FC|>1. DEGs, differentially expressed genes; FC, fold change; adj.P.Val, adjusted P‑value. Figure S3. Transcription factor‑gene regulatory network constructed using the Cytoscape iRegulion plug‑in. Table SI. Primer sequences for reverse transcription‑quantitative polymerase chain reaction. Genes Sequences hsa‑miR‑124 F: 5'‑ACACTCCAGCTGGGCAGCAGCAATTCATGTTT‑3' R: 5'‑CTCAACTGGTGTCGTGGA‑3' hsa‑miR‑330‑3p F: 5'‑CATGAATTCACTCTCCCCGTTTCTCCCTCTGC‑3' R: 5'‑CCTGCGGCCGCGAGCCGCCCTGTTTGTCTGAG‑3' hsa‑miR‑34a‑5p F: 5'‑TGGCAGTGTCTTAGCTGGTTGT‑3' R: 5'‑GCGAGCACAGAATTAATACGAC‑3' hsa‑miR‑449a F: 5'‑TGCGGTGGCAGTGTATTGTTAGC‑3' R: 5'‑CCAGTGCAGGGTCCGAGGT‑3' CD44 F: 5'‑CGGACACCATGGACAAGTTT‑3' R: 5'‑TGTCAATCCAGTTTCAGCATCA‑3' PCNA F: 5'‑GAACTGGTTCATTCATCTCTATGG‑3' F: 5'‑TGTCACAGACAAGTAATGTCGATAAA‑3' SYT1 F: 5'‑CAATAGCCATAGTCGCAGTCCT‑3' R: 5'‑TGTCAATCCAGTTTCAGCATCA‑3' U6 F: 5'‑GCTTCGGCAGCACATATACTAAAAT‑3' R: 5'‑CGCTTCACGAATTTGCGTGTCAT‑3' GAPDH F: 5'‑GGAAAGCTGTGGCGTGAT‑3' R: 5'‑AAGGTGGAAGAATGGGAGTT‑3' hsa, homo sapiens; miR, microRNA; CD44, CD44 molecule (Indian blood group); PCNA, proliferating cell nuclear antigen; -

Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’S Disease
SAGE-Hindawi Access to Research International Journal of Alzheimer’s Disease Volume 2010, Article ID 604792, 56 pages doi:10.4061/2010/604792 Research Article Complex and Multidimensional Lipid Raft Alterations in a Murine Model of Alzheimer’s Disease Wayne Chadwick, 1 Randall Brenneman,1, 2 Bronwen Martin,3 and Stuart Maudsley1 1 Receptor Pharmacology Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA 2 Miller School of Medicine, University of Miami, Miami, FL 33124, USA 3 Metabolism Unit, National Institute on Aging, National Institutes of Health, 251 Bayview Boulevard, Suite 100, Baltimore, MD 21224, USA Correspondence should be addressed to Stuart Maudsley, [email protected] Received 17 May 2010; Accepted 27 July 2010 Academic Editor: Gemma Casadesus Copyright © 2010 Wayne Chadwick et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Various animal models of Alzheimer’s disease (AD) have been created to assist our appreciation of AD pathophysiology, as well as aid development of novel therapeutic strategies. Despite the discovery of mutated proteins that predict the development of AD, there are likely to be many other proteins also involved in this disorder. Complex physiological processes are mediated by coherent interactions of clusters of functionally related proteins. Synaptic dysfunction is one of the hallmarks of AD. Synaptic proteins are organized into multiprotein complexes in high-density membrane structures, known as lipid rafts. These microdomains enable coherent clustering of synergistic signaling proteins. -

Disrupted Nitric Oxide Signaling Due to GUCY1A3 Mutations Increases Risk for Moyamoya Disease, Achalasia and Hypertension
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Clin Genet Manuscript Author . Author manuscript; Manuscript Author available in PMC 2017 October 01. Published in final edited form as: Clin Genet. 2016 October ; 90(4): 351–360. doi:10.1111/cge.12739. Disrupted Nitric Oxide Signaling due to GUCY1A3 Mutations Increases Risk for Moyamoya Disease, Achalasia and Hypertension Stephanie Wallace1, Dong-chuan Guo1, Ellen Regalado1, Lauren Mellor-Crummey1, Michael Banshad2, Deborah A. Nickerson2, Robert Dauser3, Neil Hanchard4, Ronit Marom4, Emil Martin1, Vladimir Berka1, Iraida Sharina1, Vijeya Ganesan5, Dawn Saunders6, Shaine Morris7, and Dianna M. Milewicz, MD1 1Division of Medical Genetics, Cardiology, and Hematology, Department of Internal Medicine, University of Texas Health Science Center, Houston, Texas 2Department of Genome Sciences, University of Washington, Seattle, Washington 3Department of Neurosurgery, Texas Children’s Hospital, Houston, Texas 4Department of Molecular and Human Genetics, Baylor College of Medicine, Houston, Texas 5Neuroscience Unit, University College of London Institute of Child Health, London, UK 6Department of Radiology, Great Ormond Street Hospital, London, UK 7Department of Pediatrics – Cardiology, Texas Children's Hospital and Baylor College of Medicine, Houston, Texas Abstract Moyamoya disease (MMD) is a progressive vasculopathy characterized by occlusion of the terminal portion of the internal carotid arteries and its branches, and the formation of compensatory moyamoya collateral vessels. Homozygous mutations in GUCY1A3 have been reported as a cause of MMD and achalasia. Probands (n = 96) from unrelated families underwent sequencing of GUCY1A3. Functional studies were performed to confirm the pathogenicity of identified GUCY1A3 variants. Two affected individuals from unrelated families were found to have compound heterozygous mutations in GUCY1A3. -

Compromised Global Embryonic Transcriptome Associated with Advanced Maternal Age
Journal of Assisted Reproduction and Genetics (2019) 36:915–924 https://doi.org/10.1007/s10815-019-01438-5 EMBRYO BIOLOGY Compromised global embryonic transcriptome associated with advanced maternal age Blair R. McCallie1,2 & Jason C. Parks1,2 & G. Devon Trahan3 & Kenneth L. Jones3 & Breanne D. Coate4 & Darren K. Griffin2 & William B. Schoolcraft1 & Mandy G. Katz-Jaffe1 Received: 4 February 2019 /Accepted: 12 March 2019 /Published online: 25 April 2019 # The Author(s) 2019 Abstract Purpose To investigate the global transcriptome and associated embryonic molecular networks impacted with advanced maternal age (AMA). Methods Blastocysts derived from donor oocyte IVF cycles with no male factor infertility (< 30 years of age) and AMA blastocysts (≥ 42 years) with no other significant female factor infertility or male factor infertility were collected with informed patient consent. RNA sequencing libraries were prepared using the SMARTer® Ultra® Low Kit (Clontech Laboratories) and sequenced on the Illumina HiSEQ 4000. Bioinformatics included Ingenuity® Pathway Analysis (Qiagen) with ViiA™ 7qPCR utilized for gene expression validation (Applied Biosystems). Results A total of 2688 significant differentially expressed transcripts were identified to distinguish the AMA blastocysts from young, donor controls. 2551 (95%) of these displayed decreased transcription in the blastocysts from older women. Pathway analysis revealed three altered molecular signaling networks known to be critical for embryo and fetal development: CREBBP, ESR1, and SP1. Validation of genes within these networks confirmed the global decreased transcription observed in AMA blastocysts (P <0.05). Conclusions A significant, overall decreased global transcriptome was observed in blastocysts from AMA women. The ESR1/SP1/CREBBP pathway, in particular, was found to be a highly significant upstream regulator impacting biological processes that are vital during embryonic patterning and pre-implantation development. -

A Novel Metabolic Gene Signature-Based Nomogram to Predict Overall Survival in Breast Cancer
367 Original Article Page 1 of 14 A novel metabolic gene signature-based nomogram to predict overall survival in breast cancer Xi Sun1#, Zhi-Rui Zhou2#, Yan Fang1, Shuning Ding1, Shuangshuang Lu1, Zheng Wang1, Hui Wang1, Xiaosong Chen1, Kunwei Shen1 1Department of General Surgery, Comprehensive Breast Health Center, Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China; 2Radiation Oncology Center, Huashan Hospital, Shanghai Medical College, Fudan University, Shanghai, China Contributions: (I) Conception and design: X Sun; (II) Administrative support: X Chen, K Shen; (III) Provision of study materials or patients: S Lu, Z Wang; (IV) Collection and assembly of data: Y Fang, S Ding; (V) Data analysis and interpretation: X Sun, ZR Zhou; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors #These authors contributed equally to this work. Correspondence to: Kunwei Shen; Xiaosong Chen. No. 197, Rui Jin Er Road, Shanghai 200025, China. Email: [email protected]; [email protected]. Background: Breast cancer risk prediction is often based on clinicopathological characteristics despite the high heterogeneity derived from gene expression. Metabolic alteration is a hallmark of cancer, and thus, the integration of a metabolic signature with clinical parameters is necessary to predict disease outcomes in breast cancers. Methods: Metabolic genes were downloaded from the Gene Set Enrichment Analysis (GSEA) dataset. Genes with statistical significance in the univariate analysis were applied in the least absolute shrinkage and selection operator (LASSO) analysis to build a gene signature in the GSE20685 dataset. Clinicopathological characteristics and risk scores with prognostic significance were incorporated into the nomogram to predict the overall survival (OS) of patients. -

Mir200-Regulated CXCL12β Promotes Fibroblast Heterogeneity
ARTICLE DOI: 10.1038/s41467-018-03348-z OPEN miR200-regulated CXCL12β promotes fibroblast heterogeneity and immunosuppression in ovarian cancers Anne-Marie Givel1,2, Yann Kieffer1,2, Alix Scholer-Dahirel1,2, Philemon Sirven3, Melissa Cardon1,2, Floriane Pelon1,2, Ilaria Magagna1,2, Géraldine Gentric1,2, Ana Costa1,2, Claire Bonneau1,2, Virginie Mieulet1,2, Anne Vincent-Salomon4 & Fatima Mechta-Grigoriou 1,2 1234567890():,; High-grade serous ovarian cancers (HGSOC) have been subdivided into molecular subtypes. The mesenchymal HGSOC subgroup, defined by stromal-related gene signatures, is invari- ably associated with poor patient survival. We demonstrate that stroma exerts a key function in mesenchymal HGSOC. We highlight stromal heterogeneity in HGSOC by identifying four subsets of carcinoma-associated fibroblasts (CAF-S1-4). Mesenchymal HGSOC show high content in CAF-S1 fibroblasts, which exhibit immunosuppressive functions by increasing attraction, survival, and differentiation of CD25+FOXP3+ T lymphocytes. The beta isoform of the CXCL12 chemokine (CXCL12β) specifically accumulates in the immunosuppressive CAF- S1 subset through a miR-141/200a dependent-mechanism. Moreover, CXCL12β expression in CAF-S1 cells plays a crucial role in CAF-S1 immunosuppressive activity and is a reliable prognosis factor in HGSOC, in contrast to CXCL12α. Thus, our data highlight the differential regulation of the CXCL12α and CXCL12β isoforms in HGSOC, and reveal a CXCL12β- associated stromal heterogeneity and immunosuppressive environment in mesenchymal HGSOC. 1 Institut Curie, Stress and Cancer Laboratory, Equipe labelisée Ligue Nationale Contre le Cancer, PSL Research University, 26, rue d’Ulm, 75005 Paris, France. 2 U830, Inserm, 75005 Paris, France. 3 Institut Curie, Integrative Biology of Human Dendritic Cells and T Cells Laboratory, PSL Research University, U932, Inserm, 26, rue d’Ulm, 75005 Paris, France. -

Supplemental Figures 04 12 2017
Jung et al. 1 SUPPLEMENTAL FIGURES 2 3 Supplemental Figure 1. Clinical relevance of natural product methyltransferases (NPMTs) in brain disorders. (A) 4 Table summarizing characteristics of 11 NPMTs using data derived from the TCGA GBM and Rembrandt datasets for 5 relative expression levels and survival. In addition, published studies of the 11 NPMTs are summarized. (B) The 1 Jung et al. 6 expression levels of 10 NPMTs in glioblastoma versus non‐tumor brain are displayed in a heatmap, ranked by 7 significance and expression levels. *, p<0.05; **, p<0.01; ***, p<0.001. 8 2 Jung et al. 9 10 Supplemental Figure 2. Anatomical distribution of methyltransferase and metabolic signatures within 11 glioblastomas. The Ivy GAP dataset was downloaded and interrogated by histological structure for NNMT, NAMPT, 12 DNMT mRNA expression and selected gene expression signatures. The results are displayed on a heatmap. The 13 sample size of each histological region as indicated on the figure. 14 3 Jung et al. 15 16 Supplemental Figure 3. Altered expression of nicotinamide and nicotinate metabolism‐related enzymes in 17 glioblastoma. (A) Heatmap (fold change of expression) of whole 25 enzymes in the KEGG nicotinate and 18 nicotinamide metabolism gene set were analyzed in indicated glioblastoma expression datasets with Oncomine. 4 Jung et al. 19 Color bar intensity indicates percentile of fold change in glioblastoma relative to normal brain. (B) Nicotinamide and 20 nicotinate and methionine salvage pathways are displayed with the relative expression levels in glioblastoma 21 specimens in the TCGA GBM dataset indicated. 22 5 Jung et al. 23 24 Supplementary Figure 4. -

Detection of H3k4me3 Identifies Neurohiv Signatures, Genomic
viruses Article Detection of H3K4me3 Identifies NeuroHIV Signatures, Genomic Effects of Methamphetamine and Addiction Pathways in Postmortem HIV+ Brain Specimens that Are Not Amenable to Transcriptome Analysis Liana Basova 1, Alexander Lindsey 1, Anne Marie McGovern 1, Ronald J. Ellis 2 and Maria Cecilia Garibaldi Marcondes 1,* 1 San Diego Biomedical Research Institute, San Diego, CA 92121, USA; [email protected] (L.B.); [email protected] (A.L.); [email protected] (A.M.M.) 2 Departments of Neurosciences and Psychiatry, University of California San Diego, San Diego, CA 92103, USA; [email protected] * Correspondence: [email protected] Abstract: Human postmortem specimens are extremely valuable resources for investigating trans- lational hypotheses. Tissue repositories collect clinically assessed specimens from people with and without HIV, including age, viral load, treatments, substance use patterns and cognitive functions. One challenge is the limited number of specimens suitable for transcriptional studies, mainly due to poor RNA quality resulting from long postmortem intervals. We hypothesized that epigenomic Citation: Basova, L.; Lindsey, A.; signatures would be more stable than RNA for assessing global changes associated with outcomes McGovern, A.M.; Ellis, R.J.; of interest. We found that H3K27Ac or RNA Polymerase (Pol) were not consistently detected by Marcondes, M.C.G. Detection of H3K4me3 Identifies NeuroHIV Chromatin Immunoprecipitation (ChIP), while the enhancer H3K4me3 histone modification was Signatures, Genomic Effects of abundant and stable up to the 72 h postmortem. We tested our ability to use H3K4me3 in human Methamphetamine and Addiction prefrontal cortex from HIV+ individuals meeting criteria for methamphetamine use disorder or not Pathways in Postmortem HIV+ Brain (Meth +/−) which exhibited poor RNA quality and were not suitable for transcriptional profiling. -

Wo2018/232195
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2018/232195 Al 20 December 2018 (20.12.2018) W !P O PCT (51) International Patent Classification: MANIAN, Ayshwarya; 77 Massachusetts Ave., Cam A61K 31/00 (2006.01) C07K 16/28 (2006.01) bridge, MA 02139 (US). C07K 14/705 (2006.01) C12Q 1/68 (2018.01) (72) Inventor: ROZENBLATT-ROSEN, Orit; 415 Main C07K 16/24 (2006.01) G06F 19/00 (2018.01) Street, Cambridge, MA 02142 (US). (21) International Application Number: (74) Agent: NIX, F., Brent et al.; Johnson, Marcou & Isaacs, PCT/US20 18/037662 LLC, P.O. Box 691, Hoschton, GA 30548 (US). (22) International Filing Date: (81) Designated States (unless otherwise indicated, for every 14 June 2018 (14.06.2018) kind of national protection available): AE, AG, AL, AM, (25) Filing Language: English AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, (26) Publication Langi English DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, (30) Priority Data: HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, 62/5 19,788 14 June 2017 (14.06.2017) US KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, [US/US]; (71) Applicants: THE BROAD INSTITUTE, INC. OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, 415 Main Street, Cambridge, MA 02142 (US).