Anti-CLDN8 (Internal Region) Polyclonal Antibody (CABT-BL4390) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Identification of Genes Concordantly Expressed with Atoh1 During Inner Ear Development
Original Article doi: 10.5115/acb.2011.44.1.69 pISSN 2093-3665 eISSN 2093-3673 Identification of genes concordantly expressed with Atoh1 during inner ear development Heejei Yoon, Dong Jin Lee, Myoung Hee Kim, Jinwoong Bok Department of Anatomy, Brain Korea 21 Project for Medical Science, College of Medicine, Yonsei University, Seoul, Korea Abstract: The inner ear is composed of a cochlear duct and five vestibular organs in which mechanosensory hair cells play critical roles in receiving and relaying sound and balance signals to the brain. To identify novel genes associated with hair cell differentiation or function, we analyzed an archived gene expression dataset from embryonic mouse inner ear tissues. Since atonal homolog 1a (Atoh1) is a well known factor required for hair cell differentiation, we searched for genes expressed in a similar pattern with Atoh1 during inner ear development. The list from our analysis includes many genes previously reported to be involved in hair cell differentiation such as Myo6, Tecta, Myo7a, Cdh23, Atp6v1b1, and Gfi1. In addition, we identified many other genes that have not been associated with hair cell differentiation, including Tekt2, Spag6, Smpx, Lmod1, Myh7b, Kif9, Ttyh1, Scn11a and Cnga2. We examined expression patterns of some of the newly identified genes using real-time polymerase chain reaction and in situ hybridization. For example, Smpx and Tekt2, which are regulators for cytoskeletal dynamics, were shown specifically expressed in the hair cells, suggesting a possible role in hair cell differentiation or function. Here, by re- analyzing archived genetic profiling data, we identified a list of novel genes possibly involved in hair cell differentiation. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -
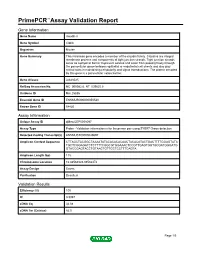
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name claudin 8 Gene Symbol Cldn8 Organism Mouse Gene Summary This intronless gene encodes a member of the claudin family. Claudins are integral membrane proteins and components of tight junction strands. Tight junction strands serve as a physical barrier to prevent solutes and water from passing freely through the paracellular space between epithelial or endothelial cell sheets and also play critical roles in maintaining cell polarity and signal transductions. The protein encoded by this gene is a paracellular cation barrier. Gene Aliases AI648025 RefSeq Accession No. NC_000082.6, NT_039625.8 UniGene ID Mm.25836 Ensembl Gene ID ENSMUSG00000050520 Entrez Gene ID 54420 Assay Information Unique Assay ID qMmuCEP0058097 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000049697 Amplicon Context Sequence CTTAGCTACGGCTAAAATATACACACACAACTACACATACTGACTTTTGGAGTATA TGCTCGGAGATCTCTTTTCGGCGTGGAAACTCCGTTGAGTGGTGCGATGGGATG GTACCGAGTACCTGTAACTGTTGCTCCTTTCAGTA Amplicon Length (bp) 115 Chromosome Location 16:88562328-88562472 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 100 R2 0.9997 cDNA Cq 34.08 cDNA Tm (Celsius) 82.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 23.77 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report Cldn8, Mouse Amplification Plot Amplification of cDNA generated from -

Morphology, Behavior, and the Sonic Hedgehog Pathway in Mouse Models of Down Syndrome
MORPHOLOGY, BEHAVIOR, AND THE SONIC HEDGEHOG PATHWAY IN MOUSE MODELS OF DOWN SYNDROME by Tara Dutka A dissertation submitted to Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland July, 2014 © 2014 Tara Dutka All Rights Reserved Abstract Down Syndrome (DS) is caused by a triplication of human chromosome 21 (Hsa21). Ts65Dn, a mouse model of DS, contains a freely segregating extra chromosome consisting of the distal portion of mouse chromosome 16 (Mmu16), a region orthologous to part of Hsa21, and a non-Hsa21 orthologous region of mouse chromosome 17. All individuals with DS display some level of craniofacial dysmorphology, brain structural and functional changes, and cognitive impairment. Ts65Dn recapitulates these features of DS and aspects of each of these traits have been linked in Ts65Dn to a reduced response to Sonic Hedgehog (SHH) in trisomic cells. Dp(16)1Yey is a new mouse model of DS which has a direct duplication of the entire Hsa21 orthologous region of Mmu16. Dp(16)1Yey’s creators found similar behavioral deficits to those seen in Ts65Dn. We performed a quantitative investigation of the skull and brain of Dp(16)1Yey as compared to Ts65Dn and found that DS-like changes to brain and craniofacial morphology were similar in both models. Our results validate examination of the genetic basis for these phenotypes in Dp(16)1Yey mice and the genetic links for these phenotypes previously found in Ts65Dn , i.e., reduced response to SHH. Further, we hypothesized that if all trisomic cells show a reduced response to SHH, then up-regulation of the SHH pathway might ameliorate multiple phenotypes. -

Identification of Claudin-8 Interaction Partners During Neural Tube Closure
Identification of Claudin-8 Interaction Partners during Neural Tube Closure Amanda Vaccarella Department of Human Genetics McGill University, Montreal, Canada August 2019 A thesis submitted to McGill University in partial fulfillment of the requirements of the degree of Master of Science © Amanda Vaccarella 2019 1 Abstract Claudins (Cldn) are a family of integral tight junction proteins that play a role in tight junction formation. Some members of the claudin family of integral tight junction proteins play important roles in neural tube closure. Removal of Cldn4 and Cldn8 from the neural ectoderm of chick embryos caused open neural tube defects (NTD) due to a failure of convergent extension and apical constriction. Protein localization at the neural ectoderm apical cell surface was disrupted in Cldn4/Cldn8-depleted embryos. Removal of only Cldn4 had no effect on neural tube closure, suggesting that Cldn8 is required for these events. The claudin cytoplasmic C-terminal tail interacts with signaling and cytoskeletal protein complexes at the tight junction cytoplasmic face. To test the function of the Cldn8 C-terminal domain, three variants at putative phosphorylation sites (S198A, S216A, S216I) and a fourth variant with a deleted PDZ binding domain (Cldn8∆YV) were created in the C-terminal domain of Cldn8. Electroporation of wild type Cldn8, ∆YV, and S216A into chick embryos had no effect on neural tube closure. The S216I variant resulted in open NTDs (p<0.002) and two-thirds of these embryos showed convergent extension defects. S198A caused only NTDs (p<0.002). Based on these data, I hypothesize that protein-protein interactions with the Cldn8 cytoplasmic domain are required for its function during neural tube closure. -

Meta-Analysis of Heterogeneous Down Syndrome Data Reveals
Vilardell et al. BMC Genomics 2011, 12:229 http://www.biomedcentral.com/1471-2164/12/229 RESEARCHARTICLE Open Access Meta-analysis of heterogeneous Down Syndrome data reveals consistent genome-wide dosage effects related to neurological processes Mireia Vilardell1,2*, Axel Rasche1, Anja Thormann1, Elisabeth Maschke-Dutz1, Luis A Pérez-Jurado2,3, Hans Lehrach1 and Ralf Herwig1* Abstract Background: Down syndrome (DS; trisomy 21) is the most common genetic cause of mental retardation in the human population and key molecular networks dysregulated in DS are still unknown. Many different experimental techniques have been applied to analyse the effects of dosage imbalance at the molecular and phenotypical level, however, currently no integrative approach exists that attempts to extract the common information. Results: We have performed a statistical meta-analysis from 45 heterogeneous publicly available DS data sets in order to identify consistent dosage effects from these studies. We identified 324 genes with significant genome- wide dosage effects, including well investigated genes like SOD1, APP, RUNX1 and DYRK1A as well as a large proportion of novel genes (N = 62). Furthermore, we characterized these genes using gene ontology, molecular interactions and promoter sequence analysis. In order to judge relevance of the 324 genes for more general cerebral pathologies we used independent publicly available microarry data from brain studies not related with DS and identified a subset of 79 genes with potential impact for neurocognitive processes. All results have been made available through a web server under http://ds-geneminer.molgen.mpg.de/. Conclusions: Our study represents a comprehensive integrative analysis of heterogeneous data including genome- wide transcript levels in the domain of trisomy 21. -

Associations Between Human Disease Genes and Overlapping Gene Groups and Multiple Amino Acid Runs
Associations between human disease genes and overlapping gene groups and multiple amino acid runs Samuel Karlin*†, Chingfer Chen*, Andrew J. Gentles*, and Michael Cleary‡ Departments of *Mathematics and ‡Pathology, Stanford University, Stanford, CA 94305 Contributed by Samuel Karlin, October 30, 2002 Overlapping gene groups (OGGs) arise when exons of one gene are Chr 22. Such rearrangements may be mediated by recombination contained within the introns of another. Typically, the two over- events based on region-specific low copy repeats. The DGS lapping genes are encoded on opposite DNA strands. OGGs are region of 22q11.2 is particularly rich with segmental duplications, often associated with specific disease phenotypes. In this report, which can induce deletions, translocations, and genomic insta- we identify genes with OGG architecture and genes encoding bility (4). There are several anomalous sequence features asso- multiple long amino acid runs and examine their relations to ciated with OGGs, including Alu sequences intersecting exons, diseases. OGGs appear to be susceptible to genomic rearrange- pseudogenes occupying introns, and single-exon (intronless) ments as happens commonly with the loci of the DiGeorge syn- genes that often result from a processed multiexon gene. drome on human chromosome 22. We also examine the degree of At least 28 genes in Chr 21 are related to diseases, as conservation of OGGs between human and mouse. Our analyses characterized in the GeneCards database (5), as are 64 genes in suggest that (i) a high proportion of genes in OGG regions are Chr 22. Specific disorders that have been mapped to genes on disease-associated, (ii) genomic rearrangements are likely to occur Chr 21 and that involve OGG structures include: amyotrophic within OGGs, possibly as a consequence of anomalous sequence lateral sclerosis (ALS, Lou Gehrig’s disease), linked to the features prevalent in these regions, and (iii) multiple amino acid GRIK1 ionotrophic kainate 1 glutamate receptor gene at 21q22 runs are also frequently associated with pathologies. -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

Investigating Contributions of Trisomy 21 in Down Syndrome to Alzheimer Disease Phenotypes in a Novel Mouse Cross
Investigating contributions of trisomy 21 in Down syndrome to Alzheimer disease phenotypes in a novel mouse cross A thesis presented in partial fulfilment of the requirements for the degree of Doctor of Philosophy from University College London Xun Yu Choong Department of Neurodegenerative Disease Institute of Neurology University College London 2016 1 Declaration I, Xun Yu Choong, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. 2 Acknowledgements The work done in this project was only possible with the support of numerous friends and colleagues, with whom I have grown an incredible amount. Foremost thanks go to Prof. Elizabeth Fisher and Dr. Frances Wiseman, who have been inexhaustibly dedicated supervisors and inspirational figures. Special mention also goes to two unofficial mentors who have looked out for me throughout the PhD, Dr. Karen Cleverley and Dr. Sarah Mizielinska. The Fisher and Isaacs groups have been a joy to work with and have always unhesitatingly offered time and help. Thank you to the Down syndrome group: Olivia Sheppard, Dr. Toby Collins, Dr. Suzanna Noy, Amy Nick, Laura Pulford, Justin Tosh, Matthew Rickman; other members of Lizzy’s group: Dr. Anny Devoy, Dr. Rosie Bunton-Stasyshyn, Dr. Rachele Saccon, Julian Pietrzyk, Dr. Beverley Burke, Heesoon Park, Julian Jaeger; the Isaacs group: Dr. Adrian Isaacs, Dr. Emma Clayton, Dr. Roberto Simone, Charlotte Ridler, Frances Norona. The PhD office has also been a second home in more ways than one, thanks to: Angelos Armen, Dr. -

Human Chromosome 21 Gene Expression Atlas in the Mouse
letters to nature Microarray data analysis .............................................................. We identified all of the positive oligonucleotides with the threshold values R ¼ 13 and D ¼ 12Q (ref. 21). R and D are threshold values for the ratio and the difference between Human chromosome 21 gene perfect match intensity and mismatch intensity, respectively. Thus varying these values gives different measures of sensitivity and specificity. We then used BLAST to identify expression atlas in the mouse conserved blocks that corresponded to at least two positive oligonucleotides so as to reduce the number of false positives. Alexandre Reymond*†, Valeria Marigo†‡, Murat B. Yaylaoglu†§, Received 16 September; accepted 30 October 2002; doi:10.1038/nature01251. Antonio Leoni‡, Catherine Ucla*, Nathalie Scamuffa*, 1. Hardison, R. C., Oeltjen, J. & Miller, W. Long human–mouse sequence alignments reveal novel Cristina Caccioppoli‡, Emmanouil T. Dermitzakis*, Robert Lyle*, regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997). Sandro Banfi , Gregor Eichele , Stylianos E. Antonarakis 2. O’Brien, S. J. et al. The promise of comparative genomics in mammals. Science 286, 458–462 (1999) ‡ § * 479–481. & Andrea Ballabio‡k 3. Shabalina, S. A., Ogurtsov, A. Y., Kondrashov, V. A. & Kondrashov, A. S. Selective constraint in intergenic regions of human and mouse genomes. Trends Genet. 17, 373–376 (2001). * Division of Medical Genetics, University of Geneva Medical School and 4. Hardison, R. C. Conserved noncoding sequences are reliable guides to regulatory elements. Trends University Hospital of Geneva, CMU, 1, rue Michel Servet, 1211 Geneva, Genet. 16, 369–372 (2000). Switzerland 5. Dermitzakis, E. T. & Clark, A. G. Evolution of transcription factor binding sites in mammalian gene regulatory regions: conservation and turnover. -

Emerging Roles of Claudins in Human Cancer
Int. J. Mol. Sci. 2013, 14, 18148-18180; doi:10.3390/ijms140918148 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Review Emerging Roles of Claudins in Human Cancer Mi Jeong Kwon 1,2 1 College of Pharmacy, Kyungpook National University, 80 Daehak-ro, Buk-gu, Daegu 702-701, Korea; E-Mail: [email protected]; Tel.: +82-53-950-8581; Fax: +82-53-950-8557 2 Research Institute of Pharmaceutical Sciences, College of Pharmacy, Kyungpook National University, 80 Daehak-ro, Buk-gu, Daegu 702-701, Korea Received: 12 August 2013; in revised form: 23 August 2013 / Accepted: 27 August 2013 / Published: 4 September 2013 Abstract: Claudins are major integral membrane proteins of tight junctions. Altered expression of several claudin proteins, in particular claudin-1, -3, -4 and -7, has been linked to the development of various cancers. Although their dysregulation in cancer suggests that claudins play a role in tumorigenesis, the exact underlying mechanism remains unclear. The involvement of claudins in tumor progression was suggested by their important role in the migration, invasion and metastasis of cancer cells in a tissue-dependent manner. Recent studies have shown that they play a role in epithelial to mesenchymal transition (EMT), the formation of cancer stem cells or tumor-initiating cells (CSCs/TICs), and chemoresistance, suggesting that claudins are promising targets for the treatment of chemoresistant and recurrent tumors. A recently identified claudin-low breast cancer subtype that is characterized by the enrichment of EMT and stem cell-like features is significantly associated with disease recurrence, underscoring the importance of claudins as predictors of tumor recurrence. -

Gene Expression Profiling Separates Chromophobe Renal Cell Carcinoma from Oncocytoma and Identifies Vesicular Transport and Cell
Human Cancer Biology Gene Expression Profiling Separates Chromophobe Renal Cell Carcinoma from Oncocytoma and Identifies Vesicular Transport andCellJunctionProteinsasDifferentiallyExpressedGenes Stephen Rohan,1Jiangling J. Tu,1Jean Kao,1Piali Mukherjee,2 Fabien Campagne,2 Xi K. Zhou,3 Elizabeth Hyjek,1Miguel A. Alonso,4 and Yao-Tseng Chen1 Abstract Purpose:To compare gene expression profiles of chromophobe renal cell carcinoma (RCC) and benign oncocytoma, aiming at identifying differentially expressed genes. Experimental Design: Nine cases each of chromophobe RCC and oncocytoma were analyzed by oligonucleotide microarray. Candidate genes that showed consistent differential expression were validated by reverse transcription-PCR using 25 fresh-frozen and 15 formalin-fixed, paraffin-embedded tumor samples. Immunohistochemical analysis was also done for two selected gene products, claudin 8 and MAL2. Results: Unsupervised hierarchical clustering separated the chromophobe RCC and oncocy- toma into two distinct groups. By a combination of data analysis approaches, we identified 11 candidate genes showing consistent differential expression between chromophobe RCC and oncocytoma. Five of these genes, AP1M2, MAL2, PROM2, PRSS8,andFLJ20171,wereshown to effectively separate these two tumor groups by quantitative reverse transcription-PCR using fresh tissue samples, with similar trends seen on formalin-fixed tissues. Immunohistochemical analysis revealed selective expression of MAL2 and claudin 8 in distal renal tubules, with MAL2 antibody showing differential expression between chromophobe RCC and oncocytoma. Functional analyses suggest that genes encoding tight junction proteins and vesicular membrane trafficking proteins, normally expressed in distal nephrons, are retained in chromophobe RCC and lost or consistently down-regulated in oncocytoma, indicating that these two tumor types, believed to be both derived from distal tubules, are likely distinctive in their histogenesis.