Precise and Sustained Gene Silencing in CD4+ Cells Using Designer Epigenome Modifiers As a Therapeutic Approach to Treat HIV Infection
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ggbio: Visualization Toolkits for Genomic Data
ggbio: visualization toolkits for genomic data 1 Tengfei Yin [email protected] May 19, 2021 Contents 1 Getting started............................4 1.1 Citation.................................4 1.2 Introduction ..............................4 2 Case study: building your first tracks..............6 2.1 Add an ideogram track ........................6 2.2 Add a gene model track .......................7 2.2.1 Introduction ...........................7 2.2.2 Make gene model from OrganismDb object ............7 2.2.3 Make gene model from TxDb object ................ 11 2.2.4 Make gene model from EnsDb object ............... 12 2.2.5 Make gene model from GRangesList object ............ 16 2.3 Add a reference track ........................ 19 2.3.1 Semantic zoom.......................... 19 2.4 Add an alignment track ....................... 20 2.5 Add a variants track ......................... 25 2.6 Building your tracks ......................... 28 3 Simple navigation .......................... 30 4 Overview plots............................ 32 4.1 how to make circular plots ...................... 32 4.1.1 Introduction ........................... 32 4.1.2 Buidling circular plot layer by layer................. 32 2 ggbio:visualization toolkits for genomic data 4.1.3 Complex arragnment of plots ................... 39 4.2 How to make grandlinear plots ................... 41 4.2.1 Introduction ........................... 41 4.2.2 Corrdinate genome ........................ 43 4.2.3 Convenient plotGrandLinear function ............... 43 4.2.4 How to highlight some points? .................. 45 4.3 How to make stacked karyogram overview plots ......... 46 4.3.1 Introduction ........................... 46 4.3.2 Create karyogram temlate .................... 46 4.3.3 Add data on karyogram layout................... 48 4.3.4 Add more data using layout_karyogram function .......... 51 4.3.5 More flexible layout of karyogram ................ -

(P -Value<0.05, Fold Change≥1.4), 4 Vs. 0 Gy Irradiation
Table S1: Significant differentially expressed genes (P -Value<0.05, Fold Change≥1.4), 4 vs. 0 Gy irradiation Genbank Fold Change P -Value Gene Symbol Description Accession Q9F8M7_CARHY (Q9F8M7) DTDP-glucose 4,6-dehydratase (Fragment), partial (9%) 6.70 0.017399678 THC2699065 [THC2719287] 5.53 0.003379195 BC013657 BC013657 Homo sapiens cDNA clone IMAGE:4152983, partial cds. [BC013657] 5.10 0.024641735 THC2750781 Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). 4.07 0.04353262 DNAH5 [Source:Uniprot/SWISSPROT;Acc:Q8TE73] [ENST00000382416] 3.81 0.002855909 NM_145263 SPATA18 Homo sapiens spermatogenesis associated 18 homolog (rat) (SPATA18), mRNA [NM_145263] AA418814 zw01a02.s1 Soares_NhHMPu_S1 Homo sapiens cDNA clone IMAGE:767978 3', 3.69 0.03203913 AA418814 AA418814 mRNA sequence [AA418814] AL356953 leucine-rich repeat-containing G protein-coupled receptor 6 {Homo sapiens} (exp=0; 3.63 0.0277936 THC2705989 wgp=1; cg=0), partial (4%) [THC2752981] AA484677 ne64a07.s1 NCI_CGAP_Alv1 Homo sapiens cDNA clone IMAGE:909012, mRNA 3.63 0.027098073 AA484677 AA484677 sequence [AA484677] oe06h09.s1 NCI_CGAP_Ov2 Homo sapiens cDNA clone IMAGE:1385153, mRNA sequence 3.48 0.04468495 AA837799 AA837799 [AA837799] Homo sapiens hypothetical protein LOC340109, mRNA (cDNA clone IMAGE:5578073), partial 3.27 0.031178378 BC039509 LOC643401 cds. [BC039509] Homo sapiens Fas (TNF receptor superfamily, member 6) (FAS), transcript variant 1, mRNA 3.24 0.022156298 NM_000043 FAS [NM_000043] 3.20 0.021043295 A_32_P125056 BF803942 CM2-CI0135-021100-477-g08 CI0135 Homo sapiens cDNA, mRNA sequence 3.04 0.043389246 BF803942 BF803942 [BF803942] 3.03 0.002430239 NM_015920 RPS27L Homo sapiens ribosomal protein S27-like (RPS27L), mRNA [NM_015920] Homo sapiens tumor necrosis factor receptor superfamily, member 10c, decoy without an 2.98 0.021202829 NM_003841 TNFRSF10C intracellular domain (TNFRSF10C), mRNA [NM_003841] 2.97 0.03243901 AB002384 C6orf32 Homo sapiens mRNA for KIAA0386 gene, partial cds. -

Anglemetry of Neural Axis Cell Differentiation Genes by Structural Pressurotopy of DNA Loop Strand Segment Tropy in Reference to Tissue Macro-Compliance Hemant Sarin
Sarin Translational Medicine Communications (2019) 4:13 Translational Medicine https://doi.org/10.1186/s41231-019-0045-4 Communications RESEARCH Open Access Anglemetry of neural axis cell differentiation genes by structural pressurotopy of DNA loop strand segment tropy in reference to tissue macro-compliance Hemant Sarin Abstract Background: Even as the eukaryotic stranded DNA is known to heterochromatinize at the nuclear envelope in response to mechanical strain, the precise mechanistic basis for alterations in chromatin gene transcription in differentiating cell lineages has been difficult to determine due to limited spatial resolution for detection of shifts in reference to a specific gene in vitro. In this study, heterochromatin shift during euchromatin gene transcription has been studied by parallel determinations of DNA strand loop segmentation tropy nano-compliance (esebssiwaagoTQ units, linear nl), gene positioning angulation in linear normal two- 0 dimensional (2-D) z, y-vertical plane (anglemetry, ), horizontal alignment to the z, x-plane (vectormetry; mA, mM,a.u.),andby pressuromodulation mapping of differentiated neuron cell sub-class operating range for neuroaxis gene expression in reference to tissue macro-compliance (Peff). Methods: The esebssiwaagoTQ effectivepressureunit(Peff) maxima and minima for horizontal gene intergene base segment tropy loop alignment were determined (n = 224); the Peff esebssiwaagoTQ quotient were determined (n =28)foranalysisof gene intergene base loop segment tropy structure nano-compliance (n = 28; n = 188); and gene positioning anglemetry and vectormetry was performed (n = 42). The sebs intercept-to-sebssiwa intercept quotient for linear normalization was determined (bsebs/bsebssiwa) by exponential plotting of sub-episode block sum (sebs) (x1, y1; x2, y2) and sub-episode block sum split integrated weighted average (sebssiwa) functions was performed, and the sebs – sebssiwa function residuals were determined. -

Plasma Fatty Acid Ratios Affect Blood Gene Expression Profiles - a Cross-Sectional Study of the Norwegian Women and Cancer Post-Genome Cohort
Plasma Fatty Acid Ratios Affect Blood Gene Expression Profiles - A Cross-Sectional Study of the Norwegian Women and Cancer Post-Genome Cohort Karina Standahl Olsen1*, Christopher Fenton2, Livar Frøyland3, Marit Waaseth4, Ruth H. Paulssen2, Eiliv Lund1 1 Department of Community Medicine, University of Tromsø, Tromsø, Norway, 2 Department of Clinical Medicine, University of Tromsø, Tromsø, Norway, 3 National Institute of Nutrition and Seafood Research (NIFES), Bergen, Norway, 4 Department of Pharmacy, University of Tromsø, Tromsø, Norway Abstract High blood concentrations of n-6 fatty acids (FAs) relative to n-3 FAs may lead to a ‘‘physiological switch’’ towards permanent low-grade inflammation, potentially influencing the onset of cardiovascular and inflammatory diseases, as well as cancer. To explore the potential effects of FA ratios prior to disease onset, we measured blood gene expression profiles and plasma FA ratios (linoleic acid/alpha-linolenic acid, LA/ALA; arachidonic acid/eicosapentaenoic acid, AA/EPA; and total n-6/n-3) in a cross-section of middle-aged Norwegian women (n = 227). After arranging samples from the highest values to the lowest for all three FA ratios (LA/ALA, AA/EPA and total n-6/n-3), the highest and lowest deciles of samples were compared. Differences in gene expression profiles were assessed by single-gene and pathway-level analyses. The LA/ALA ratio had the largest impact on gene expression profiles, with 135 differentially expressed genes, followed by the total n-6/ n-3 ratio (125 genes) and the AA/EPA ratio (72 genes). All FA ratios were associated with genes related to immune processes, with a tendency for increased pro-inflammatory signaling in the highest FA ratio deciles. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

MIKULASOVA Et Al. MMEJ DRIVES 8Q24 REARRANGEMENTS in MYELOMA
MIKULASOVA et al. MMEJ DRIVES 8q24 REARRANGEMENTS IN MYELOMA Microhomology-mediated end joining drives complex rearrangements and over-expression of MYC and PVT1 in multiple myeloma Authors Aneta Mikulasova1,2, Cody Ashby1, Ruslana G. Tytarenko1, Pingping Qu3, Adam Rosenthal3, Judith A. Dent1, Katie R. Ryan1, Michael A. Bauer1, Christopher P. Wardell1, Antje Hoering3, Konstantinos Mavrommatis4, Matthew Trotter5, Shayu Deshpande1, Erming Tian1, Jonathan Keats6, Daniel Auclair7, Graham H. Jackson8, Faith E. Davies1, Anjan Thakurta4, Gareth J. Morgan1, and Brian A. Walker1 Affiliations 1Myeloma Center, University of Arkansas for Medical Sciences, Little Rock, AR, USA 2Institute of Cellular Medicine, Newcastle University, Newcastle upon Tyne, United Kingdom 3Cancer Research and Biostatistics, Seattle, WA, USA 4Celgene Corporation, Summit, NJ, USA 5Celgene Institute for Translational Research Europe, Seville, Spain 6Translational Genomics Research Institute, Phoenix, AZ, USA. 7Multiple Myeloma Research Foundation, Norwalk, CT, USA. 8Northern Institute for Cancer Research, Newcastle University, Newcastle upon Tyne, United Kingdom MIKULASOVA et al. MMEJ DRIVES 8q24 REARRANGEMENTS IN MYELOMA Contents Supplementary Methods ............................................................................................................... 2 Data Analysis .............................................................................................................................. 2 External Datasets ...................................................................................................................... -

Elucidating the Genetic Architecture of Reproductive Ageing in the Japanese Population
bioRxiv preprint doi: https://doi.org/10.1101/182717; this version posted August 30, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Title: Elucidating the genetic architecture of reproductive ageing in the Japanese population Momoko Horikoshi*1, Felix R. Day*2, Yoichiro Kamatani3,4, Makoto Hirata5, Masato Akiyama3, Koichi Matsuda5, Hollis Wright6, Carlos A. Toro6, Sergio R. Ojeda7, Alejandro Lomniczi6, Michiaki Kubo*8, Ken K. Ong*2 and John R. B. Perry*2 1. Laboratory for Endocrinology, Metabolism and Kidney Diseases, RIKEN Centre for Integrative Medical Sciences, Yokohama 230-0045, Japan 2. MRC Epidemiology Unit, University of Cambridge School of Clinical Medicine, Institute of Metabolic Science, Cambridge Biomedical Campus, Cambridge, UK. 3. Laboratory for Statistical Analysis, RIKEN Center for Integrative Medical Sciences, Yokohama 230-0045, Japan 4. Center for Genomic Medicine, Kyoto University Graduate School of Medicine, Kyoto 606- 8507, Japan. 5. Laboratory of Genome Technology, Human Genome Center, Institute of Medical Science, The University of Tokyo, Tokyo 108-8639, Japan 6. Primate Genetics Section / Division of Neuroscience, Oregon National Primate Research Center, Oregon Health and Sciences University, Beaverton, OR 97006, USA 7. Division of Neuroscience, Oregon National Primate Research Center, Oregon Health and Sciences University, Beaverton, OR 97006, USA 8. RIKEN Center for Integrative Medical Sciences, Yokohama 230-0045, Japan * denotes equal contributions Correspondence to Momoko Horikoshi ([email protected]) and John R.B. -

Nº Ref Uniprot Proteína Péptidos Identificados Por MS/MS 1 P01024
Document downloaded from http://www.elsevier.es, day 26/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Nº Ref Uniprot Proteína Péptidos identificados 1 P01024 CO3_HUMAN Complement C3 OS=Homo sapiens GN=C3 PE=1 SV=2 por 162MS/MS 2 P02751 FINC_HUMAN Fibronectin OS=Homo sapiens GN=FN1 PE=1 SV=4 131 3 P01023 A2MG_HUMAN Alpha-2-macroglobulin OS=Homo sapiens GN=A2M PE=1 SV=3 128 4 P0C0L4 CO4A_HUMAN Complement C4-A OS=Homo sapiens GN=C4A PE=1 SV=1 95 5 P04275 VWF_HUMAN von Willebrand factor OS=Homo sapiens GN=VWF PE=1 SV=4 81 6 P02675 FIBB_HUMAN Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 78 7 P01031 CO5_HUMAN Complement C5 OS=Homo sapiens GN=C5 PE=1 SV=4 66 8 P02768 ALBU_HUMAN Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 66 9 P00450 CERU_HUMAN Ceruloplasmin OS=Homo sapiens GN=CP PE=1 SV=1 64 10 P02671 FIBA_HUMAN Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 58 11 P08603 CFAH_HUMAN Complement factor H OS=Homo sapiens GN=CFH PE=1 SV=4 56 12 P02787 TRFE_HUMAN Serotransferrin OS=Homo sapiens GN=TF PE=1 SV=3 54 13 P00747 PLMN_HUMAN Plasminogen OS=Homo sapiens GN=PLG PE=1 SV=2 48 14 P02679 FIBG_HUMAN Fibrinogen gamma chain OS=Homo sapiens GN=FGG PE=1 SV=3 47 15 P01871 IGHM_HUMAN Ig mu chain C region OS=Homo sapiens GN=IGHM PE=1 SV=3 41 16 P04003 C4BPA_HUMAN C4b-binding protein alpha chain OS=Homo sapiens GN=C4BPA PE=1 SV=2 37 17 Q9Y6R7 FCGBP_HUMAN IgGFc-binding protein OS=Homo sapiens GN=FCGBP PE=1 SV=3 30 18 O43866 CD5L_HUMAN CD5 antigen-like OS=Homo -
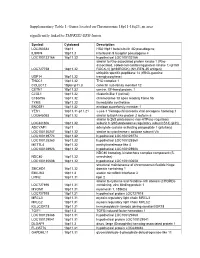
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Transcriptional Regulators Are Upregulated in the Substantia Nigra
Journal of Emerging Investigators Transcriptional Regulators are Upregulated in the Substantia Nigra of Parkinson’s Disease Patients Marianne Cowherd1 and Inhan Lee2 1Community High School, Ann Arbor, MI 2miRcore, Ann Arbor, MI Summary neurological conditions is an established practice (3). Parkinson’s disease (PD) affects approximately 10 Significant gene expression dysregulation in the SN and million people worldwide with tremors, bradykinesia, in the striatum has been described, particularly decreased apathy, memory loss, and language issues. Though such expression in PD synapses. Protein degradation has symptoms are due to the loss of the substantia nigra (SN) been found to be upregulated (4). Mutations in SNCA brain region, the ultimate causes and complete pathology are unknown. To understand the global gene expression (5), LRRK2 (6), and GBA (6) have also been identified changes in SN, microarray expression data from the SN as familial markers of PD. SNCA encodes alpha- tissue of 9 controls and 16 PD patients were compared, synuclein, a protein found in presynaptic terminals that and significantly upregulated and downregulated may regulate vesicle presence and dopamine release. genes were identified. Among the upregulated genes, Eighteen SNCA mutations have been associated with a network of 33 interacting genes centered around the PD and, although the exact pathogenic mechanism is cAMP-response element binding protein (CREBBP) was not confirmed, mutated alpha-synuclein is the major found. The downstream effects of increased CREBBP- component of protein aggregates, called Lewy bodies, related transcription and the resulting protein levels that are often found in PD brains and may contribute may result in PD symptoms, making CREBBP a potential therapeutic target due to its central role in the interactive to cell death. -

(12) Patent Application Publication (10) Pub. No.: US 2016/0264934 A1 GALLOURAKIS Et Al
US 20160264934A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2016/0264934 A1 GALLOURAKIS et al. (43) Pub. Date: Sep. 15, 2016 (54) METHODS FOR MODULATING AND Publication Classification ASSAYING MI6AIN STEM CELL POPULATIONS (51) Int. Cl. CI2N5/0735 (2006.01) (71) Applicants: THE GENERAL, HOSPITAL AOIN I/02 (2006.01) CORPORATION, Boston, MA (US); CI2O I/68 (2006.01) The Regents of the University of GOIN 33/573 (2006.01) California, Oakland, CA (US) CI2N 5/077 (2006.01) CI2N5/0793 (2006.01) (72) Inventors: Cosmas GIALLOURAKIS, Boston, (52) U.S. Cl. MA (US); Alan C. MULLEN, CPC ............ CI2N5/0606 (2013.01); CI2N5/0657 Brookline, MA (US); Yi XING, (2013.01); C12N5/0619 (2013.01); C12O Torrance, CA (US) I/6888 (2013.01); G0IN33/573 (2013.01); A0IN I/0226 (2013.01); C12N 2501/72 (73) Assignees: THE GENERAL, HOSPITAL (2013.01); C12N 2506/02 (2013.01); C12O CORPORATION, Boston, MA (US); 2600/158 (2013.01); C12Y 201/01062 The Regents of the University of (2013.01); C12Y 201/01 (2013.01) California, Oakland, CA (US) (57) ABSTRACT (21) Appl. No.: 15/067,780 The present invention generally relates to methods, assays and kits to maintain a human stem cell population in an (22) Filed: Mar 11, 2016 undifferentiated state by inhibiting the expression or function of METTL3 and/or METTL4, and mA fingerprint methods, assays, arrays and kits to assess the cell state of a human stem Related U.S. Application Data cell population by assessing mA levels (e.g. mA peak inten (60) Provisional application No. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19