Interferon Alpha-Based Combinations Suppress SARS-Cov-2 Infection In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cytokine Nomenclature
RayBiotech, Inc. The protein array pioneer company Cytokine Nomenclature Cytokine Name Official Full Name Genbank Related Names Symbol 4-1BB TNFRSF Tumor necrosis factor NP_001552 CD137, ILA, 4-1BB ligand receptor 9 receptor superfamily .2. member 9 6Ckine CCL21 6-Cysteine Chemokine NM_002989 Small-inducible cytokine A21, Beta chemokine exodus-2, Secondary lymphoid-tissue chemokine, SLC, SCYA21 ACE ACE Angiotensin-converting NP_000780 CD143, DCP, DCP1 enzyme .1. NP_690043 .1. ACE-2 ACE2 Angiotensin-converting NP_068576 ACE-related carboxypeptidase, enzyme 2 .1 Angiotensin-converting enzyme homolog ACTH ACTH Adrenocorticotropic NP_000930 POMC, Pro-opiomelanocortin, hormone .1. Corticotropin-lipotropin, NPP, NP_001030 Melanotropin gamma, Gamma- 333.1 MSH, Potential peptide, Corticotropin, Melanotropin alpha, Alpha-MSH, Corticotropin-like intermediary peptide, CLIP, Lipotropin beta, Beta-LPH, Lipotropin gamma, Gamma-LPH, Melanotropin beta, Beta-MSH, Beta-endorphin, Met-enkephalin ACTHR ACTHR Adrenocorticotropic NP_000520 Melanocortin receptor 2, MC2-R hormone receptor .1 Activin A INHBA Activin A NM_002192 Activin beta-A chain, Erythroid differentiation protein, EDF, INHBA Activin B INHBB Activin B NM_002193 Inhibin beta B chain, Activin beta-B chain Activin C INHBC Activin C NM005538 Inhibin, beta C Activin RIA ACVR1 Activin receptor type-1 NM_001105 Activin receptor type I, ACTR-I, Serine/threonine-protein kinase receptor R1, SKR1, Activin receptor-like kinase 2, ALK-2, TGF-B superfamily receptor type I, TSR-I, ACVRLK2 Activin RIB ACVR1B -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Antibiotic Fidaxomicin Is an Rdrp Inhibitor As a Potential New Therapeutic Agent Against Zika Virus
Yuan et al. BMC Medicine (2020) 18:204 https://doi.org/10.1186/s12916-020-01663-1 RESEARCH ARTICLE Open Access Antibiotic fidaxomicin is an RdRp inhibitor as a potential new therapeutic agent against Zika virus Jie Yuan1,2,3†, Jianchen Yu2,3,4†, Yun Huang3, Zhenjian He2,5, Jia Luo6, Yun Wu2,4,7, Yingchun Zheng8, Jueheng Wu2,4,7, Xun Zhu2,4, Haihe Wang1 and Mengfeng Li2,3,4* Abstract Background: Zika virus (ZIKV) infection is a global health problem, and its complications, including congenital Zika syndrome and Guillain-Barré syndrome, constitute a continued threat to humans. Unfortunately, effective therapeutics against ZIKV infection are not available thus far. Methods: We screened the compounds collection consisting of 1789 FDA-approved drugs by a computational docking method to obtain anti-ZIKV candidate compounds targeting ZIKV RNA-dependent RNA polymerase (RdRp). SPR (BIAcore) assay was employed to demonstrate the candidate compounds’ direct binding to ZIKV RdRp, and polymerase activity assay was used to determine the inhibitory effect on ZIKV RdRp-catalyzed RNA synthesis. The antiviral effects on ZIKV in vitro and in vivo were detected in infected cultured cells and in Ifnar1−/− mice infected by ZIKV virus using plaque assay, western blotting, tissue immunofluorescence, and immunohistochemistry. Results: Here, we report that a first-in-class macrocyclic antibiotic, which has been clinically used to treat Clostridium difficile infection, fidaxomicin, potently inhibits ZIKV replication in vitro and in vivo. Our data showed that fidaxomicin was effective against African and Asian lineage ZIKV in a wide variety of cell lines of various tissue origins, and prominently suppressed ZIKV infection and significantly improved survival of infected mice. -

Secretome Screening Reveals Immunomodulating Functions Of
www.nature.com/scientificreports OPEN Secretome screening reveals immunomodulating functions of IFNα‑7, PAP and GDF‑7 on regulatory T‑cells Mei Ding1*, Rajneesh Malhotra2, Tomas Ottosson2, Magnus Lundqvist3, Aman Mebrahtu3, Johan Brengdahl1, Ulf Gehrmann2, Elisabeth Bäck4, Douglas Ross‑Thriepland5, Ida Isaksson6, Björn Magnusson1, Kris F. Sachsenmeier7, Hanna Tegel3, Sophia Hober3, Mathias Uhlén3, Lorenz M. Mayr5, Rick Davies5, Johan Rockberg3 & Lovisa Holmberg Schiavone1* Regulatory T cells (Tregs) are the key cells regulating peripheral autoreactive T lymphocytes. Tregs exert their function by suppressing efector T cells. Tregs have been shown to play essential roles in the control of a variety of physiological and pathological immune responses. However, Tregs are unstable and can lose the expression of FOXP3 and suppressive functions as a consequence of outer stimuli. Available literature suggests that secreted proteins regulate Treg functional states, such as diferentiation, proliferation and suppressive function. Identifcation of secreted proteins that afect Treg cell function are highly interesting for both therapeutic and diagnostic purposes in either hyperactive or immunosuppressed populations. Here, we report a phenotypic screening of a human secretome library in human Treg cells utilising a high throughput fow cytometry technology. Screening a library of 575 secreted proteins allowed us to identify proteins stabilising or destabilising the Treg phenotype as suggested by changes in expression of Treg marker proteins FOXP3 and/or CTLA4. Four proteins including GDF‑7, IL‑10, PAP and IFNα‑7 were identifed as positive regulators that increased FOXP3 and/or CTLA4 expression. PAP is a phosphatase. A catalytic‑dead version of the protein did not induce an increase in FOXP3 expression. -

Affiliations
Supplementary material Proteome-wide survey of the autoimmune target repertoire in autoimmune polyendocrine syndrome type 1 *Nils Landegren1,2, Donald Sharon3,4, Eva Freyhult2,5,6,, Åsa Hallgren1,2, Daniel Eriksson1,2, Per-Henrik Edqvist7, Sophie Bensing8, Jeanette Wahlberg9, Lawrence M. Nelson10, Jan Gustafsson11, Eystein S Husebye12, Mark S Anderson13, Michael Snyder3, Olle Kämpe1,2 Nils Landegren and Donald Sharon contributed equally to the work Affiliations 1Department of Medicine (Solna), Karolinska University Hospital, Karolinska Institutet, Sweden 2Science for Life Laboratory, Department of Medical Sciences, Uppsala University, Sweden 3Department of Genetics, Stanford University, California, USA 4Department of Molecular, Cellular, and Developmental Biology, Yale University, Connecticut, USA 1 5Department of Medical Sciences, Cancer Pharmacology and Computational Medicine, Uppsala University 6Bioinformatics Infrastructure for Life Sciences 7Department of Immunology, Genetics and Pathology, Uppsala University, Sweden and Science for Life Laboratory 8 Department of Molecular Medicine and Surgery, Karolinska Institutet, Stockholm, Sweden 9Department of Endocrinology and Department of Medical and Health Sciences and Department of Clinical and Experimental Medicine, Linköping University, Linköping, Sweden 10Integrative Reproductive Medicine Group, Intramural Research Program on Reproductive and Adult Endocrinology, National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD 20892, USA. 11Department -

A Rational Approach to Identifying Effective Combined Anticoronaviral Therapies Against Feline 2 Coronavirus 3 4 5 S.E
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.09.195016; this version posted July 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 A rational approach to identifying effective combined anticoronaviral therapies against feline 2 coronavirus 3 4 5 S.E. Cook1*, H. Vogel2, D. Castillo3, M. Olsen4, N. Pedersen5, B. G. Murphy3 6 7 1 Graduate Group Integrative Pathobiology, School of Veterinary Medicine, University of 8 California, Davis, CA, USA 9 10 2School of Veterinary Medicine, University of California, Davis, Ca, USA 11 12 3Department of Pathology, Microbiology, and Immunology, School of Veterinary Medicine, 13 University of California, Davis, CA, USA 14 15 4Department of Pharmaceutical Sciences, College of Pharmacy-Glendale, Midwestern 16 University, Glendale, AZ, USA 17 18 5Department of Medicine and Epidemiology, School of Veterinary Medicine, University of 19 California, Davis, CA, USA 20 21 22 *Corresponding author 23 24 E-mail: [email protected] (SEC) 25 26 27 Abstract 28 Feline infectious peritonitis (FIP), caused by a genetic mutant of feline enteric coronavirus 29 known as FIPV, is a highly fatal disease of cats with no currently available vaccine or FDA- 30 approved cure. Dissemination of FIPV in affected cats results in a range of clinical signs 31 including cavitary effusions, anorexia, fever and lesions of pyogranulomatous vasculitis and 32 peri-vasculitis with or without central nervous system and/or ocular involvement. -

Proteome-Wide Survey of the Autoimmune Target Repertoire In
www.nature.com/scientificreports OPEN Proteome-wide survey of the autoimmune target repertoire in autoimmune polyendocrine Received: 28 October 2015 Accepted: 23 December 2015 syndrome type 1 Published: 01 February 2016 Nils Landegren1,2,*, Donald Sharon3,4,*, Eva Freyhult2,5,6, Åsa Hallgren1,2, Daniel Eriksson1,2, Per-Henrik Edqvist7, Sophie Bensing8, Jeanette Wahlberg9, Lawrence M. Nelson10, Jan Gustafsson11, Eystein S Husebye12, Mark S Anderson13, Michael Snyder3 & Olle Kämpe1,2 Autoimmune polyendocrine syndrome type 1 (APS1) is a monogenic disorder that features multiple autoimmune disease manifestations. It is caused by mutations in the Autoimmune regulator (AIRE) gene, which promote thymic display of thousands of peripheral tissue antigens in a process critical for establishing central immune tolerance. We here used proteome arrays to perform a comprehensive study of autoimmune targets in APS1. Interrogation of established autoantigens revealed highly reliable detection of autoantibodies, and by exploring the full panel of more than 9000 proteins we further identified MAGEB2 and PDILT as novel major autoantigens in APS1. Our proteome-wide assessment revealed a marked enrichment for tissue-specific immune targets, mirroringAIRE ’s selectiveness for this category of genes. Our findings also suggest that only a very limited portion of the proteome becomes targeted by the immune system in APS1, which contrasts the broad defect of thymic presentation associated with AIRE-deficiency and raises novel questions what other factors are needed for break of tolerance. Autoimmune responses can ultimately be defined at the molecular level by the specific interaction between T- or B-cell receptors and a distinct self-molecule. In tissue-specific autoimmune disorders the immune system typically target molecules that are exclusively expressed in the affected tissue and involve a combined cellular and humoral response with cognate specificities1–3. -

Heatmaps - the Gene Expression Edition
Heatmaps - the gene expression edition Jeff Oliver 20 July, 2020 An application of heatmap visualization to investigate differential gene expression. Learning objectives 1. Manipulate data into a ‘tidy’ format 2. Visualize data in a heatmap 3. Become familiar with ggplot syntax for customizing plots Heatmaps for differential gene expression Heatmaps are a great way of displaying three-dimensional data in only two dimensions. But how can we easily translate tabular data into a format for heatmap plotting? By taking advantage of “data munging” and graphics packages, heatmaps are relatively easy to produce in R. Getting started We are going to start by isolating different types of information by imposing structure in our file managment. That is, we are going to put our input data in one folder and any output such as plots or analytical results in a different folder. We can use the dir.create to create these two folders: dir.create("data") dir.create("output") For this lesson, we will use a subset of data on a study of gene expression in cells infected with the influenza virus (doi: 10.4049/jimmunol.1501557). The study infected human plasmacytoid dendritic cells with the influenza virus, and compared gene expression in those cells to gene expression in uninfected cells. Thegoal was to see how the flu virus affected the function of these immune system cells. The data for this lesson is available at: http://tinyurl.com/flu-expression-data or https://jcoliver.github.io/ learn-r/data/GSE68849-expression.csv. Download this comma separated file and put it in the data folder. -

Microarray Analysis of Colorectal Cancer Stromal Tissue Reveals Upregulation of Two Oncogenic Mirna Clusters
Published OnlineFirst March 27, 2012; DOI: 10.1158/1078-0432.CCR-11-1078 Clinical Cancer Human Cancer Biology Research Microarray Analysis of Colorectal Cancer Stromal Tissue Reveals Upregulation of Two Oncogenic miRNA Clusters Naohiro Nishida1,4, Makoto Nagahara2, Tetsuya Sato3, Koshi Mimori1, Tomoya Sudo1, Fumiaki Tanaka1, Kohei Shibata1, Hideshi Ishii1,4, Kenichi Sugihara2, Yuichiro Doki4, and Masaki Mori1,4 Abstract Purpose: Cancer stroma plays an important role in the progression of cancer. Although alterations in miRNA expression have been explored in various kinds of cancers, the expression of miRNAs in cancer stroma has not been explored in detail. Experimental Design: Using a laser microdissection technique, we collected RNA samples specific for epithelium or stroma from 13 colorectal cancer tissues and four normal tissues, and miRNA microarray and gene expression microarray were carried out. The expression status of miRNAs was confirmed by reverse transcriptase PCR. Furthermore, we investigated whether miRNA expression status in stromal tissue could influence the clinicopathologic factors. Results: Oncogenic miRNAs, including two miRNA clusters, miR-17-92a and miR-106b-25 cluster, were upregulated in cancer stromal tissues compared with normal stroma. Gene expression profiles from cDNA microarray analyses of the same stromal tissue samples revealed that putative targets of these miRNA clusters, predicted by Target Scan, such as TGFBR2, SMAD2, and BMP family genes, were significantly downregulated in cancer stromal tissue. Downregulated putative targets were also found to be involved in cytokine interaction and cellular adhesion. Importantly, expression of miR-25 and miR-92a in stromal tissues was associated with a variety of clinicopathologic factors. Conclusions: Oncogenic miRNAs were highly expressed in cancer stroma. -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

DNA/RNA Synthesis
DNA/RNA Synthesis RNA synthesis, which is also called DNA transcription, is a highly selective process. Transcription by RNA polymerase II extends beyond RNA synthesis, towards a more active role in mRNA maturation, surveillance and export to the cytoplasm. Single-strand breaks are repaired by DNA ligase using the complementary strand of the double helix as a template, with DNA ligase creating the final phosphodiester bond to fully repair the DNA.DNA ligases discriminate against substrates containing RNA strands or mismatched base pairs at positions near the ends of the nickedDNA. Bleomycin (BLM) exerts its genotoxicity by generating free radicals, whichattack C-4′ in the deoxyribose backbone of DNA, leading to opening of the ribose ring and strand breakage; it is an S-independentradiomimetic agent that causes double-strand breaks in DNA. First strand cDNA is synthesized using random hexamer primers and M-MuLV Reverse Transcriptase (RNase H). Second strand cDNA synthesis is subsequently performed using DNA Polymerase I and RNase H. The remaining overhangs are converted into blunt ends using exonuclease/polymerase activity. After adenylation of the 3′ ends of DNA fragments, NEBNext Adaptor with hairpin loop structure is ligated to prepare the samples for hybridization. Cell cycle and DNA replication are the top two pathways regulated by BET bromodomain inhibition. Cycloheximide blocks the translation of mRNA to protein. www.MedChemExpress.com 1 DNA/RNA Synthesis Inhibitors, Agonists, Activators, Modulators & Chemicals (+)-TK216 (-)-TK216 Cat. No.: HY-122903B Cat. No.: HY-122903A (+)-TK216 is an enantiomer of TK216 (HY-122903). (-)-TK216 is an enantiomer of TK216 (HY-122903). TK216 is an orally active and potent E26 TK216 is an orally active and potent E26 transformation specific (ETS) inhibitor. -
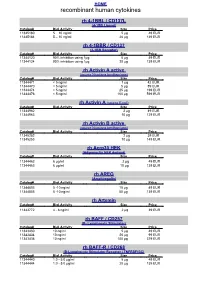
Recombinant Human Cytokines
HOME recombinant human cytokines rh 4-1BBL / CD137L (4-1BB Ligand) Catalog# Biol.Activity Size Price 11345180 5 – 10 ng/ml 5 µg 49 EUR 11345184 5 – 10 ng/ml 20 µg 139 EUR rh 4-1BBR / CD137 (4-1BB Receptor) Catalog# Biol.Activity Size Price 11344120 90% inhibition using 1µg 5 µg 49 EUR 11344124 90% inhibition using 1µg 20 µg 139 EUR rh Activin A active (source Nicotiana benthamiana) Catalog# Biol.Activity Size Price 11344471 < 5 ng/ml 1 µg 42 EUR 11344470 < 5 ng/ml 5 µg 89 EUR 11344474 < 5 ng/ml 25 µg 199 EUR 11344476 < 5 ng/ml 100 µg 599 EUR rh Activin A (source E.coli) Catalog# Biol.Activity Size Price 11344962 2 µg 49 EUR 11344963 10 µg 129 EUR rh Activin B active (source Nicotiana benthamiana) Catalog# Biol.Activity Size Price 11345252 2 µg 39 EUR 11345253 10 µg 149 EUR rh Acrp30 HEK (Adiponectin HEK derived) Catalog# Biol.Activity Size Price 11344462 6 µg/ml 2 µg 49 EUR 11344463 6 µg/ml 10 µg 139 EUR rh AREG (Amphiregulin) Catalog# Biol.Activity Size Price 11344803 5 -10 ng/ml 10 µg 49 EUR 11344805 5 -10 ng/ml 50 µg 139 EUR rh Artemin Catalog# Biol.Activity Size Price 11343772 4 - 8 ng/ml 2 µg 39 EUR rh BAFF / CD257 (B- Lymphocyte Stimulator) Catalog# Biol.Activity Size Price 11343430 10 ng/ml 5 µg 49 EUR 11343434 10 ng/ml 20 µg 99 EUR 11343436 10 ng/ml 100 µg 379 EUR rh BAFF-R / CD268 (B-Lymphocyte Stimulator Receptor / TNFRSF13C) Catalog# Biol.Activity Size Price 11344440 1.0 - 5.0 µg/ml 5 µg 49 EUR 11344444 1.0 - 5.0 µg/ml 20 µg 139 EUR HOME recombinant human cytokines rh BCA-1 (CXCL13) Catalog# Biol.Activity Size Price 11344180