Wild Crop Relatives: Genomic and Breeding Resources: Plantation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Minireview Paper Golden Camellias
1 1 Minireview Paper 2 Golden Camellias: A Review 3 4 ABSTRACT 5 Golden camellias or yellow camellias are species belonging to genus Camellia L., family 6 Theaceae. Fifty two species were described in southern China and Vietnam. Active 7 ingredients such as polysaccharides, polyphenols, tea saponins, and flavonoids are well 8 known characteristics of golden camellias. Its leaves and flowers have been long 9 traditionally used for health improvement. It was found to be able to inhibit the 10 transplanted cancer, lower blood pressure, lower blood lipid, lower cholesterol, and 11 prevent atherosclerosis. Currently, it cost 320-700US$ per one kg of dry flowers. Such 12 price attracts many local ethnic people to plant golden camellias for poverty reduction. 13 This work reviews (1) species and natural distribution, (2) uses and healthcare values, (3) 14 techniques for seedling production, planting and tending, and (4) opportunities and 15 challenges for future development of golden camellias. 16 17 Keywords: Active ingredient; Camellia L.; poverty reduction; shade-tolerant species; yellow flower. 2 18 1. SPECIES AND NATURAL DISTRIBUTION 19 Golden camellias or yellow camellias are shrubs and small-sized trees belonging to genus 20 Camellia L [1], family Theaceae [2-8]. Golden camellias have light to heavy yellow 21 flowers (Fig. 1) and are 3-12 m tall at maturity in natural distribution conditions. The size 22 of flowers are different among species from 1 to 10 cm in diameter (Fig. 1). About 52 23 species (Table 1) of golden camellias have been described in southern China and Vietnam. 24 Of which, nearly 40 species have natural distribution in Vietnam. -

Number 3, Spring 1998 Director’S Letter
Planning and planting for a better world Friends of the JC Raulston Arboretum Newsletter Number 3, Spring 1998 Director’s Letter Spring greetings from the JC Raulston Arboretum! This garden- ing season is in full swing, and the Arboretum is the place to be. Emergence is the word! Flowers and foliage are emerging every- where. We had a magnificent late winter and early spring. The Cornus mas ‘Spring Glow’ located in the paradise garden was exquisite this year. The bright yellow flowers are bright and persistent, and the Students from a Wake Tech Community College Photography Class find exfoliating bark and attractive habit plenty to photograph on a February day in the Arboretum. make it a winner. It’s no wonder that JC was so excited about this done soon. Make sure you check of themselves than is expected to seedling selection from the field out many of the special gardens in keep things moving forward. I, for nursery. We are looking to propa- the Arboretum. Our volunteer one, am thankful for each and every gate numerous plants this spring in curators are busy planting and one of them. hopes of getting it into the trade. preparing those gardens for The magnolias were looking another season. Many thanks to all Lastly, when you visit the garden I fantastic until we had three days in our volunteers who work so very would challenge you to find the a row of temperatures in the low hard in the garden. It shows! Euscaphis japonicus. We had a twenties. There was plenty of Another reminder — from April to beautiful seven-foot specimen tree damage to open flowers, but the October, on Sunday’s at 2:00 p.m. -

Download Camellia Collection.Pdf
List of Taxa TaxonName Genus Species InfraType1 InfraName1 AccCount FamilyEx Camellia × (cf 38) Camellia × 1 Theaceae Camellia × 'Ack-Scent' Camellia × 1 Theaceae Camellia × 'Anne McCulloch Hill' Camellia × 1 Theaceae Camellia × 'April Kiss' Camellia × 1 Theaceae Camellia × 'April Remembered' ICE ANGELS® Camellia × 1 Theaceae Camellia × 'Arctic Dawn' Camellia × 1 Theaceae Camellia × 'Ashton's Cameo' Camellia × 1 Theaceae Camellia × 'Ashton's Pride' Camellia × 1 Theaceae Camellia × 'Autumn Spirit' Camellia × 2 Theaceae Camellia × 'Barbara Clark' Camellia × 1 Theaceae Camellia × 'Bett's Supreme' Camellia × 1 Theaceae Camellia × 'Betty Ridley' Camellia × 1 Theaceae Camellia × 'Big Apple' Camellia × 1 Theaceae Camellia × 'Bill Goertz' Camellia × 1 Theaceae Camellia × 'Black Lace' Camellia × 1 Theaceae Camellia × 'Buddy English' Camellia × 1 Theaceae Camellia × 'Buttermint' Camellia × 1 Theaceae Camellia × 'Cameron Cooper' Camellia × 1 Theaceae Camellia × 'Candle Glow' Camellia × 1 Theaceae Camellia × 'Carolina Moonmist' Camellia × 1 Theaceae Camellia × 'China Girl' Camellia × 1 Theaceae Camellia × 'Cinnamon Cindy' Camellia × 1 Theaceae Camellia × 'Cinnamon Scentsation' Camellia × 1 Theaceae Camellia × 'Coral Bouquet' Camellia × 1 Theaceae Camellia × 'Cornish Snow' Camellia × 1 Theaceae Camellia × 'Crimson Candles' Camellia × 1 Theaceae Camellia × 'Debut' Camellia × 1 Theaceae Camellia × 'Dot Spengler' Camellia × 1 Theaceae Camellia × 'Dr. Clifford Parks' Camellia × 1 Theaceae Camellia × 'Dr. Louis Polizzi' Camellia × 1 Theaceae Camellia × -
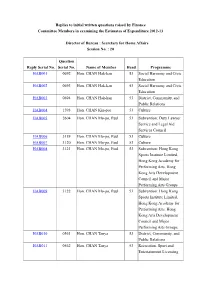
Administration's Replies to Members Initial Written Questions
Replies to initial written questions raised by Finance Committee Members in examining the Estimates of Expenditure 2012-13 Director of Bureau : Secretary for Home Affairs Session No. : 20 Question Reply Serial No. Serial No. Name of Member Head Programme HAB001 0692 Hon. CHAN Hak-kan 53 Social Harmony and Civic Education HAB002 0693 Hon. CHAN Hak-kan 53 Social Harmony and Civic Education HAB003 0694 Hon. CHAN Hak-kan 53 District, Community, and Public Relations HAB004 1593 Hon. CHAN Kin-por 53 Culture HAB005 2604 Hon. CHAN Mo-po, Paul 53 Subvention: Duty Lawyer Service and Legal Aid Services Council HAB006 3119 Hon. CHAN Mo-po, Paul 53 Culture HAB007 3120 Hon. CHAN Mo-po, Paul 53 Culture HAB008 3121 Hon. CHAN Mo-po, Paul 53 Subvention: Hong Kong Sports Institute Limited, Hong Kong Academy for Performing Arts, Hong Kong Arts Development Council and Major Performing Arts Groups HAB009 3122 Hon. CHAN Mo-po, Paul 53 Subvention: Hong Kong Sports Institute Limited, Hong Kong Academy for Performing Arts, Hong Kong Arts Development Council and Major Performing Arts Groups HAB010 0561 Hon. CHAN Tanya 53 District, Community, and Public Relations HAB011 0562 Hon. CHAN Tanya 53 Recreation, Sport and Entertainment Licensing Question Reply Serial No. Serial No. Name of Member Head Programme HAB012 0568 Hon. CHAN Tanya 53 Social Harmony and Civic Education HAB013 0569 Hon. CHAN Tanya 53 Social Harmony and Civic Education HAB014 0570 Hon. CHAN Tanya 53 District, Community, and Public Relations HAB015 0901 Hon. CHAN Tanya 53 Recreation, Sport and Entertainment Licensing HAB016 1677 Hon. FOK Tsun-ting, 53 Social Harmony and Civic Timothy Education HAB017 1678 Hon. -

Camellia Sinensis (L.) Kuntze (7)
“As Primeiras Camélias Asiáticas a Chegarem a Portugal e à Europa”. Armando Oliveira António Sanches (1623), Planisfério. 1 O género Camellia L. está praticamente confinado ao sul da China (80% de todas as espécies) e à região do sul da Ásia que inclui as Filipinas e as zonas do noroeste do arquipélago da Indonésia, com a inclusão do Japão e partes da Coreia. Estima-se que praticamente 20% das espécies de Camellia se encontram no Vietname. A região fitogeográfica do sul da Ásia é composta pela China, Laos, Mianmar (ex-Birmânia), Tailândia, Camboja e Vietname. 1 (Huang et al., 2016) 106 • A proposta taxonómica de Linnaeus (1835), “Sistema Natura”, permitiu-nos obter uma mais fácil e rápida identificação das espécies. • Baseia-se numa classificação dita binomial que atribui nomes compostos por duas palavras, quase sempre recorrendo ao latim. Adaptado de Fairy Lake Botanical Garden Flora (2018) 2 Reino Filo Classe Ordem Família Género Espécies/Variedades Cultivares Camellia caudata Wall. (11) Camellia drupifera Lour. (4) Dicotiledóneas Antófitas Camellia euryoides Lindl. (7) Vegetal (a semente Ericales (25) Theaceaes (12) Camellia (102+40) (que dão flor) contém 2 ou mais Camellia japonica L. cotilédones) Camellia kissi Wall. (11) Camellia oleifera Abel (6) Camellia rosaeflora Hook. (1) Camellia sasanqua Thunb. Camellia sinensis (L.) Kuntze (7) • A 1ª parte do nome é referente ao género da espécie em causa e a 2ª parte identifica a espécie dentro de um determinado género. Adaptado de Fairy Lake Botanical Garden Flora (2018) 2 Ordem Família -

Isolation and Characterization of S-Rnase-Homologous Genes Expressed in Styles in ‘Hyuganatsu’ (Citrus Tamurana Hort
This article is an Advance Online Publication of the authors’ corrected proof. Note that minor changes may be made before final version publication. The Horticulture Journal Preview e Japanese Society for doi: 10.2503/hortj.UTD-032 JSHS Horticultural Science http://www.jshs.jp/ Isolation and Characterization of S-RNase-homologous Genes Expressed in Styles in ‘Hyuganatsu’ (Citrus tamurana hort. ex Tanaka) Chitose Honsho1*, Shingo Umegatani2, Dai Furukawa1, Shuji Ishimura1 and Takuya Tetsumura1 1Faculty of Agriculture, University of Miyazaki, Miyazaki 889-2192, Japan 2Graduate School of Agriculture, University of Miyazaki, Miyazaki 889-2192, Japan In this study, S-RNase-homologous genes expressed in styles were isolated from a gametophytic self- incompatible citrus cultivar, ‘Hyuganatsu’. Sweet orange and clementine genome databases were searched to identify 13 ribonuclease T2 (T2 RNase) genes. Further blast searches using citrus EST databases were conducted with these 13 sequences as queries to obtain five additional EST sequences. Known T2 RNase genes, including the S-RNases of Rosaceae, Solanaceae, and Plantaginaceae were retrieved from the public database. All data collected from the databases were combined to make a dataset for phylogenetic analysis. From the phylogenetic tree, 10 citrus sequences were found to be monophyletic to the clade of S-RNases. Degenerate primers were designed from these genes to identify similar sequences in cDNA derived from ‘Hyuganatsu’ styles. RT-PCR and 3' and 5' RACE resulted in the isolation of three S-RNase-like sequences; these sequences were further characterized. Pistil-specific gene expression was confirmed for all sequences by RT-PCR; Citrus tamurana RNase1 (CtRNS1) and RNase3 (CtRNS3) had basic pI values (> 8) and molecular weights of approximately 25 kDa, consistent with S-RNase features of other families. -

And Cross-Pollination in Pollen Tube Growth, Early Ovule Development and Fruit Set of Camellia Grijsii
INTERNATIONAL JOURNAL OF AGRICULTURE & BIOLOGY ISSN Print: 1560–8530; ISSN Online: 1814–9596 18–1478/2019/21–4–819–826 DOI: 10.17957/IJAB/15.0960 http://www.fspublishers.org Full Length Article Comparison of Self- and Cross-Pollination in Pollen Tube Growth, Early Ovule Development and Fruit Set of Camellia grijsii Huan Xiong1, Feng Zou1*, Deyi Yuan1*, Xiaofeng Tan1, Jun Yuan1, Ting Liao1 and Genhua Niu2 1Key Laboratory of Cultivation and Protection for Non-Wood Forest Trees, Ministry of Education Central South University of Forestry and Technology, Changsha 410004, Hunan, China 2Texas AgriLife Research at El Paso, Texas A&M University System, 1380 A&M Circle, El Paso, TX 79927, USA *For correspondence: [email protected]; [email protected] Abstract Camellia grijsii Hance is one of the most important woody edible oil tree species in Southern China; however, it often has a low fruit set rate. To elucidate the causes of poor fruit set in C. grijsii, self-pollination (SP) with C. grijsii and cross-pollination (CP) of C. grijsii × C. villosa tests were conducted. Pollen germination and pollen tube growth into pistils, and early ovule development after SP and CP, were examined using a paraffin section and fluorescence microscopy. The fruit set percentage in SP and CP was also investigated. The results showed that pollen germinated normally on the stigma, and the pollen tubes both reached the style base after SP and CP, but the growth rates of pollen differed significantly between SP and CP, being faster for CP. The pollen tubes arrived at the style base 48 h after SP, but only 24 h after CP. -

Vascular Plants of Santa Cruz County, California
ANNOTATED CHECKLIST of the VASCULAR PLANTS of SANTA CRUZ COUNTY, CALIFORNIA SECOND EDITION Dylan Neubauer Artwork by Tim Hyland & Maps by Ben Pease CALIFORNIA NATIVE PLANT SOCIETY, SANTA CRUZ COUNTY CHAPTER Copyright © 2013 by Dylan Neubauer All rights reserved. No part of this publication may be reproduced without written permission from the author. Design & Production by Dylan Neubauer Artwork by Tim Hyland Maps by Ben Pease, Pease Press Cartography (peasepress.com) Cover photos (Eschscholzia californica & Big Willow Gulch, Swanton) by Dylan Neubauer California Native Plant Society Santa Cruz County Chapter P.O. Box 1622 Santa Cruz, CA 95061 To order, please go to www.cruzcps.org For other correspondence, write to Dylan Neubauer [email protected] ISBN: 978-0-615-85493-9 Printed on recycled paper by Community Printers, Santa Cruz, CA For Tim Forsell, who appreciates the tiny ones ... Nobody sees a flower, really— it is so small— we haven’t time, and to see takes time, like to have a friend takes time. —GEORGIA O’KEEFFE CONTENTS ~ u Acknowledgments / 1 u Santa Cruz County Map / 2–3 u Introduction / 4 u Checklist Conventions / 8 u Floristic Regions Map / 12 u Checklist Format, Checklist Symbols, & Region Codes / 13 u Checklist Lycophytes / 14 Ferns / 14 Gymnosperms / 15 Nymphaeales / 16 Magnoliids / 16 Ceratophyllales / 16 Eudicots / 16 Monocots / 61 u Appendices 1. Listed Taxa / 76 2. Endemic Taxa / 78 3. Taxa Extirpated in County / 79 4. Taxa Not Currently Recognized / 80 5. Undescribed Taxa / 82 6. Most Invasive Non-native Taxa / 83 7. Rejected Taxa / 84 8. Notes / 86 u References / 152 u Index to Families & Genera / 154 u Floristic Regions Map with USGS Quad Overlay / 166 “True science teaches, above all, to doubt and be ignorant.” —MIGUEL DE UNAMUNO 1 ~ACKNOWLEDGMENTS ~ ANY THANKS TO THE GENEROUS DONORS without whom this publication would not M have been possible—and to the numerous individuals, organizations, insti- tutions, and agencies that so willingly gave of their time and expertise. -

Common Bean. from the United States
26 PLANT INVENTORY NO. 170 278662. GOSSYPIUM HIRSUTUM L. Malvaceae. Upland cotton. From the Republic of Ivory Coast. Seeds presented by the Cotton Research Station, Bouake. Received Jan. 15, 1962. Selfed seeds of a monosomic plant found in 'Allen 333' at Bouake, 1960. 278663. CAMELLIA HONGKONGENSIS Seem. Theaceae. From Hong Kong. Seeds presented by the Urban Services Department, Vic- toria. Received Jan. 15, 1962. 278664 to 278695. PHASEOLUS VULGARIS L. Fabaceae. Common bean. From the United States. Seeds held in storage at the United States Regional Plant Introduction Station, Geneva, N.Y. Numbered Jan. 15, 1962. 278664. No. G 11341. Bush to 12 inches high; flowers white; pods 4.5 by 0.5 inch, flat, light green, clusters above and below foliage, midseason to late maturing; seeds medium size, plump, oval, tan with red stripes. 278665. No. G 11342. Pole bean to 9 feet high; flowers whitish purple; pods 4 by 0.37 inch, flat, dark green to purple, clusters above and below foliage, ripening early midseason; seeds small, black, plump, rectangu- lar. 278666. No. G 11343. Pole bean to 9 feet high; flowers whitish purple; pods 4 by 0.37 inch, flat, dark green to purple, clusters above and below foliage, seeds medium size, long, plump, oval, red. 278667. No. G 11344. Pole bean to 9 feet high; flowers whitish purple; pods 4 by 0.37 inch, flat, dark green to purple, clusters above and below foliage; seeds medium size, flat, rectangular, tan. 278668. No. G 11345. Bush to 10 inches high; flowers pink; pods 4.5 by 0.5 inch, flat, light green, maturing midseason to late; seeds large, ob- long, buff, many aborted; hilum sunken, brown. -

International Camellia Journal 2017
International Camellia Journal 2017 An official publication of the International Camellia Society Journal Number 49 ISSN 0159-656X International Camellia Journal 2017 No. 49 International Camellia Society Congress 2018 Nantes, France, March 25 to 29 Pre-Congress Tour March 21-25 Gardens in Brittany Aims of the International Camellia Society Congress Registration March 25 in Nantes To foster the love of camellias throughout the world and maintain and increase their popularity Post-Congress Tours To undertake historical, scientific and horticultural research in connection with camellias Tour A March 29 to 31 Normandy, including World War II To co-operate with all national and regional camellia societies and with other horticultural societies sites To disseminate information concerning camellias by means of bulletins and other publications To encourage a friendly exchange between camellia enthusiasts of all nationalities Tour B March 29 to 31 Gardens and nurseries in southwest France Major dates in the International Camellia Society calendar Reassemble in Paris April 1 to 2, including visit to Versailles International Camellia Society Congresses 2018 - Nantes, Brittany, France. 2020 - Goto City, Japan. 2022 - Italy ISSN 0159-656X Published in 2017 by the International Camellia Society. © The International Camellia Society unless otherwise stated 3 Contents Camellia research A transcriptomic database of petal blight-resistant Camellia lutchuensis 47 Nikolai Kondratev1, Matthew Denton-Giles1,2, Cade D Fulton1, President’s message 6 Paul P Dijkwel1 -

* Correspondence To: Yujing Yan, Email: [email protected] Or Charles C
Supplementary Materials for Phytogeographic history of the Tea family inferred through high-resolution phylogeny and fossils *Yujing Yan1,2, *Charles C. Davis2, Dimitar Dimitrov1,3, Zhiheng Wang4, Carsten Rahbek5,1,4,6,7, Michael Krabbe Borregaard1 1. Center for Macroecology, Evolution and Climate, GLOBE Institute, University of Copenhagen, Universitetsparken 15, 2100, Copenhagen, Denmark 2. Department of Organismic and Evolutionary Biology, Harvard University Herbaria, 22 Divinity Ave, Cambridge, MA 02138, USA 3. Department of Natural History, University Museum of Bergen, University of Bergen, P.O. Box 7800, 5020 Bergen, Norway 4. Institute of Ecology, College of Urban and Environmental Sciences, Key Laboratory of Earth Surface Processes of Ministry of Education, Peking University, Beijing 100871, China 5. Center for Global Mountain Biodiversity, GLOBE Institute, University of Copenhagen, Universitetsparken 15, 2100 Copenhagen, Denmark 6. Department of Life Sciences, Imperial College London, Silkwood Park campus, Ascot SL5 7PY, UK 7. Danish Institute for Advanced Study, University of Southern Denmark, Odense, Denmark. * Correspondence to: Yujing Yan, email: [email protected] or Charles C. Davis, email: [email protected] This PDF file includes: Appendix 1 (p.4) Supplementary Notes 1 (p. 2-4) Supplementary Tables S1 to S11 (p.7-42) Supplementary Figures S1 to S10 (p.43-52) 1 Supplementary Note 1. Inference of biogeographical patterns using a fully Bayesian method in RevBayes We applied an alternative biogeographic method proposed by Landis et al. (2020) to a subset of our Theaceae empirical dataset and compared its performance to our method. This method uses a hierarchical Bayesian approach to account for the uncertainty in the position of fossils (lack of characters), phylogenetic relationships, and the geological/biogeographic template at once. -

Els Herbaris, Fonts Per Al Coneixement De La Flora. Aplicacions En Conservació I Taxonomia
Els herbaris, fonts per al coneixement de la flora. Aplicacions en conservació i taxonomia Neus Nualart Dexeus ADVERTIMENT. La consulta d’aquesta tesi queda condicionada a l’acceptació de les següents condicions d'ús: La difusió d’aquesta tesi per mitjà del servei TDX (www.tdx.cat) i a través del Dipòsit Digital de la UB (diposit.ub.edu) ha estat autoritzada pels titulars dels drets de propietat intel·lectual únicament per a usos privats emmarcats en activitats d’investigació i docència. No s’autoritza la seva reproducció amb finalitats de lucre ni la seva difusió i posada a disposició des d’un lloc aliè al servei TDX ni al Dipòsit Digital de la UB. No s’autoritza la presentació del seu contingut en una finestra o marc aliè a TDX o al Dipòsit Digital de la UB (framing). Aquesta reserva de drets afecta tant al resum de presentació de la tesi com als seus continguts. En la utilització o cita de parts de la tesi és obligat indicar el nom de la persona autora. ADVERTENCIA. La consulta de esta tesis queda condicionada a la aceptación de las siguientes condiciones de uso: La difusión de esta tesis por medio del servicio TDR (www.tdx.cat) y a través del Repositorio Digital de la UB (diposit.ub.edu) ha sido autorizada por los titulares de los derechos de propiedad intelectual únicamente para usos privados enmarcados en actividades de investigación y docencia. No se autoriza su reproducción con finalidades de lucro ni su difusión y puesta a disposición desde un sitio ajeno al servicio TDR o al Repositorio Digital de la UB.