Targeting EZH2 Histone Methyltransferase Activity Alleviates Experimental Intestinal Inflammation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Screening and Identification of Key Biomarkers in Clear Cell Renal Cell Carcinoma Based on Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Screening and identification of key biomarkers in clear cell renal cell carcinoma based on bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Clear cell renal cell carcinoma (ccRCC) is one of the most common types of malignancy of the urinary system. The pathogenesis and effective diagnosis of ccRCC have become popular topics for research in the previous decade. In the current study, an integrated bioinformatics analysis was performed to identify core genes associated in ccRCC. An expression dataset (GSE105261) was downloaded from the Gene Expression Omnibus database, and included 26 ccRCC and 9 normal kideny samples. Assessment of the microarray dataset led to the recognition of differentially expressed genes (DEGs), which was subsequently used for pathway and gene ontology (GO) enrichment analysis. -

Human Catechol O-Methyltransferase Genetic Variation
Molecular Psychiatry (2004) 9, 151–160 & 2004 Nature Publishing Group All rights reserved 1359-4184/04 $25.00 www.nature.com/mp ORIGINAL RESEARCH ARTICLE Human catechol O-methyltransferase genetic variation: gene resequencing and functional characterization of variant allozymes AJ Shield1, BA Thomae1, BW Eckloff2, ED Wieben2 and RM Weinshilboum1 1Department of Molecular Pharmacology and Experimental Therapeutics, Mayo Medical School, Mayo Clinic, Mayo Foundation, Rochester, MN, USA; 2Department of Biochemistry and Molecular Biology, Mayo Medical School, Mayo Clinic, Mayo Foundation, Rochester, MN, USA Catechol O-methyltransferase (COMT) plays an important role in the metabolism of catecholamines, catecholestrogens and catechol drugs. A common COMT G472A genetic polymorphism (Val108/158Met) that was identified previously is associated with decreased levels of enzyme activity and has been implicated as a possible risk factor for neuropsychiatric disease. We set out to ‘resequence’ the human COMT gene using DNA samples from 60 African-American and 60 Caucasian-American subjects. A total of 23 single nucleotide polymorphisms (SNPs), including a novel nonsynonymous cSNP present only in DNA from African-American subjects, and one insertion/deletion were observed. The wild type (WT) and two variant allozymes, Thr52 and Met108, were transiently expressed in COS-1 and HEK293 cells. There was no significant change in level of COMT activity for the Thr52 variant allozyme, but there was a 40% decrease in the level of activity in cells transfected with the Met108 construct. Apparent Km values of the WT and variant allozymes for the two reaction cosubstrates differed slightly, but significantly, for 3,4-dihydroxybenzoic acid but not for S-adenosyl-L-methionine. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Histone Methylases As Novel Drug Targets. Focus on EZH2 Inhibition. Catherine BAUGE1,2,#, Céline BAZILLE 1,2,3, Nicolas GIRARD1
Histone methylases as novel drug targets. Focus on EZH2 inhibition. Catherine BAUGE1,2,#, Céline BAZILLE1,2,3, Nicolas GIRARD1,2, Eva LHUISSIER1,2, Karim BOUMEDIENE1,2 1 Normandie Univ, France 2 UNICAEN, EA4652 MILPAT, Caen, France 3 Service d’Anatomie Pathologique, CHU, Caen, France # Correspondence and copy request: Catherine Baugé, [email protected], EA4652 MILPAT, UFR de médecine, Université de Caen Basse-Normandie, CS14032 Caen cedex 5, France; tel: +33 231068218; fax: +33 231068224 1 ABSTRACT Posttranslational modifications of histones (so-called epigenetic modifications) play a major role in transcriptional control and normal development, and are tightly regulated. Disruption of their control is a frequent event in disease. Particularly, the methylation of lysine 27 on histone H3 (H3K27), induced by the methylase Enhancer of Zeste homolog 2 (EZH2), emerges as a key control of gene expression, and a major regulator of cell physiology. The identification of driver mutations in EZH2 has already led to new prognostic and therapeutic advances, and new classes of potent and specific inhibitors for EZH2 show promising results in preclinical trials. This review examines roles of histone lysine methylases and demetylases in cells, and focuses on the recent knowledge and developments about EZH2. Key-terms: epigenetic, histone methylation, EZH2, cancerology, tumors, apoptosis, cell death, inhibitor, stem cells, H3K27 2 Histone modifications and histone code Epigenetic has been defined as inheritable changes in gene expression that occur without a change in DNA sequence. Key components of epigenetic processes are DNA methylation, histone modifications and variants, non-histone chromatin proteins, small interfering RNA (siRNA) and micro RNA (miRNA). -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Histone Deacetylase Inhibitors Synergizes with Catalytic Inhibitors of EZH2 to Exhibit Anti-Tumor Activity in Small Cell Carcinoma of the Ovary, Hypercalcemic Type
Author Manuscript Published OnlineFirst on September 19, 2018; DOI: 10.1158/1535-7163.MCT-18-0348 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Histone deacetylase inhibitors synergizes with catalytic inhibitors of EZH2 to exhibit anti- tumor activity in small cell carcinoma of the ovary, hypercalcemic type Yemin Wang1,2,*, Shary Yuting Chen1,2, Shane Colborne3, Galen Lambert2, Chae Young Shin2, Nancy Dos Santos4, Krystal A. Orlando5, Jessica D. Lang6, William P.D. Hendricks6, Marcel B. Bally4, Anthony N. Karnezis1,2, Ralf Hass7, T. Michael Underhill8, Gregg B. Morin3,9, Jeffrey M. Trent6, Bernard E. Weissman5, David G. Huntsman1,2,10,* 1Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, BC, Canada 2Department of Molecular Oncology, British Columbia Cancer Research Centre, Vancouver, BC, Canada. 3Michael Smith Genome Science Centre, British Columbia Cancer Agency, Vancouver, BC, Canada. 4Department of Experimental Therapeutics, British Columbia Cancer Research Centre, Vancouver, BC, Canada. 5Department of Pathology and Laboratory Medicine and Lineberger Comprehensive Cancer Center, University of North Carolina, Chapel Hill, NC, USA. 6Division of Integrated Cancer Genomics, Translational Genomics Research Institute (TGen), Phoenix, AZ, USA. 7Department of Obstetrics and Gynecology, Hannover Medical School, D-30625 Hannover, Germany. 8Department of Cellular and Physiological Sciences and Biomedical Research Centre, University 1 Downloaded from mct.aacrjournals.org on September 26, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on September 19, 2018; DOI: 10.1158/1535-7163.MCT-18-0348 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

EZH2 in Normal Hematopoiesis and Hematological Malignancies
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 3 EZH2 in normal hematopoiesis and hematological malignancies Laurie Herviou2, Giacomo Cavalli2, Guillaume Cartron3,4, Bernard Klein1,2,3 and Jérôme Moreaux1,2,3 1 Department of Biological Hematology, CHU Montpellier, Montpellier, France 2 Institute of Human Genetics, CNRS UPR1142, Montpellier, France 3 University of Montpellier 1, UFR de Médecine, Montpellier, France 4 Department of Clinical Hematology, CHU Montpellier, Montpellier, France Correspondence to: Jérôme Moreaux, email: [email protected] Keywords: hematological malignancies, EZH2, Polycomb complex, therapeutic target Received: August 07, 2015 Accepted: October 14, 2015 Published: October 20, 2015 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Enhancer of zeste homolog 2 (EZH2), the catalytic subunit of the Polycomb repressive complex 2, inhibits gene expression through methylation on lysine 27 of histone H3. EZH2 regulates normal hematopoietic stem cell self-renewal and differentiation. EZH2 also controls normal B cell differentiation. EZH2 deregulation has been described in many cancer types including hematological malignancies. Specific small molecules have been recently developed to exploit the oncogenic addiction of tumor cells to EZH2. Their therapeutic potential is currently under evaluation. This review summarizes the roles of EZH2 in normal and pathologic hematological processes and recent advances in the development of EZH2 inhibitors for the personalized treatment of patients with hematological malignancies. PHYSIOLOGICAL FUNCTIONS OF EZH2 state through tri-methylation of lysine 27 on histone H3 (H3K27me3) [6]. -
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Peter Midford Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_003855395Cyc: Shewanella livingstonensis LMG 19866 Cellular Overview Connections between pathways are omitted for legibility. -
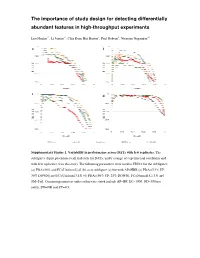
The Importance of Study Design Abundant Features in High
The importance of study design for detecting differentially abundant features in high -throughput experiments Luo Huaien 1* , Li Juntao 1*, C hia Kuan Hui Burton 1, Paul Robson 2, Niranjan Nagarajan 1# a b c d Supplementary Figure 1. Variability in performance across DATs with few repreplicates.licates. The subfigures depict precision-recall tradeoffs for DATs, under a range of experimexperimenentaltal conditions and with few replicates (2 in this case). The following parameters were used in EDDA for the sub figures: (a) PDA=30% and FC=Uniform[2,6] (b) as in subfigure (a) but with AP=HBR (c) PDA= [15% UP, 30% DOWN] and FC=Uniform[3,15] (d) PDA=[50% UP, 25% DOWN], FC=Normal[5,1.33] and SM=Full. Common parameters unless otherwise stated include AP=BP, EC=1000, ND=500 per entity, SM=NB and SV=0.5. a b c d Supplementary Figure 2 . Variability in performance across DATs with few data -points. Note that as in Supplementary Figure 1 , the performance of DATs varies widely merely by changing the number of replicates from 1 to 4 in sub figures (a)-(d) in a situation where the number data -points generated is limited (only 50 on average per entity). Results reported are based on data generated in EDDA with the following parameters (roughly mimicking a microbial RNA-seq application) : EC=1000, PDA=26%, FC=Uniform[3,7 ], AP=BP, SM=Full and SV=0.85. Supplementary Figure 3. Distribution of relative abundance in different empirically derived profiles. Note that both axes are plotted on a log-scale and that while BP does not have entities with relative abundance < 1e-5, the profile from Wu et al is more enriched for entities with relative abundance < 1e-5 than the HBR profile. -

Expanding the Structural Diversity of DNA Methyltransferase Inhibitors
pharmaceuticals Article Expanding the Structural Diversity of DNA Methyltransferase Inhibitors K. Eurídice Juárez-Mercado 1 , Fernando D. Prieto-Martínez 1 , Norberto Sánchez-Cruz 1 , Andrea Peña-Castillo 1, Diego Prada-Gracia 2 and José L. Medina-Franco 1,* 1 DIFACQUIM Research Group, Department of Pharmacy, School of Chemistry, National Autonomous University of Mexico, Avenida Universidad 3000, Mexico City 04510, Mexico; [email protected] (K.E.J.-M.); [email protected] (F.D.P.-M.); [email protected] (N.S.-C.); [email protected] (A.P.-C.) 2 Research Unit on Computational Biology and Drug Design, Children’s Hospital of Mexico Federico Gomez, Mexico City 06720, Mexico; [email protected] * Correspondence: [email protected] Abstract: Inhibitors of DNA methyltransferases (DNMTs) are attractive compounds for epigenetic drug discovery. They are also chemical tools to understand the biochemistry of epigenetic processes. Herein, we report five distinct inhibitors of DNMT1 characterized in enzymatic inhibition assays that did not show activity with DNMT3B. It was concluded that the dietary component theaflavin is an inhibitor of DNMT1. Two additional novel inhibitors of DNMT1 are the approved drugs glyburide and panobinostat. The DNMT1 enzymatic inhibitory activity of panobinostat, a known pan inhibitor of histone deacetylases, agrees with experimental reports of its ability to reduce DNMT1 activity in liver cancer cell lines. Molecular docking of the active compounds with DNMT1, and re-scoring with the recently developed extended connectivity interaction features approach, led to an excellent agreement between the experimental IC50 values and docking scores. Citation: Juárez-Mercado, K.E.; Keywords: dietary component; epigenetics; enzyme inhibition; focused library; epi-informatics; Prieto-Martínez, F.D.; Sánchez-Cruz, multitarget epigenetic agent; natural products; chemoinformatics N.; Peña-Castillo, A.; Prada-Gracia, D.; Medina-Franco, J.L. -

TAZVERIK (Tazemetostat) RATIONALE for INCLUSION in PA PROGRAM
TAZVERIK (tazemetostat) RATIONALE FOR INCLUSION IN PA PROGRAM Background Tazverik (tazemetostat) is an inhibitor of the methyltransferase, EZH2, and some EZH2 gain-of- function mutations including Y646X and A687V. The most well-characterized function of EZH2 is as the catalytic subunit of the polycomb repressive complex 2 (PRC2), catalyzing mono-, di-, and trimethylation of the lysine 27 of the histone H3. Trimethylation of histone H3 leads to transcriptional repression. SWItch/Sucrose Non-Fermentable (SWI/SNF) complexes can antagonize PRC2 function in the regulation of the expression of certain genes. The loss or dysfunction of certain SWI/SNF complex members can lead to aberrant EZH2 activity or expression and a resulting oncogenic dependence on EZH2 (1). Regulatory Status FDA -approved indication: Tazverik is a methyltransferase inhibitor indicated for the treatment of: (1) 1. Adults and pediatric patients aged 16 years and older with metastatic or locally advanced epithelioid sarcoma not eligible for complete resection. 2. Adult patients with relapsed or refractory follicular lymphoma whose tumors are positive for an EZH2 mutation as detected by an FDA-approved test and who have received at least 2 prior systemic therapies. 3. Adult patients with relapsed or refractory follicular lymphoma who have no satisfactory alternative treatment options. The risk of developing secondary malignancies is increased following treatment with Tazverik. Across clinical trials of 668 adults who received Tazverik 800 mg twice daily, myelodysplastic syndrome (MDS) or acute myeloid leukemia (AML) occurred in 0.6% of patients (1). Tazverik can cause fetal harm when administered to pregnant women. Pregnant women should be advised of the potential risk to a fetus. -

Catechol-O-Methyltransferase Inhibitor, on the Motor Response to Acute Treatment with Levodopa in Patients with Parkinson's Disease
18616ournal ofNeurology, Neurosurgery, and Psychiatry 1994;57:186-189 J Neurol Neurosurg Psychiatry: first published as 10.1136/jnnp.57.2.186 on 1 February 1994. Downloaded from Effect of entacapone, a peripherally acting catechol-O-methyltransferase inhibitor, on the motor response to acute treatment with levodopa in patients with Parkinson's disease Marcelo Merello, Andrew J Lees, Roy Webster, Michael Bovingdon, Ariel Gordin Abstract treatment.7-'0 The oral bioavailability of lev- Catechol-O-methyltransferase (COMT) odopa is low due to first pass metabolism inhibitors may be useful in the treatment which is at least partly due to the high activity of Parkinson's disease by improving the of COMT in the gut and liver." 12 The com- bioavailability of levodopa and by pro- bination of a peripheral COMT inhibitor longing its effects. Entacapone (OR-611), with levodopa/DCI therapy might therefore a novel COMT inhibitor, which does not produce a smoother and more prolonged cross the blood brain barrier, was motor response in patients with Parkinson's assessed in 12 patients with Parkinson's disease."3 14 As early as in 1971 Ericsson'5 disease and motor fluctuations in a reported beneficial effects with a COMT randomised, double-blind, cross-over, inhibitor with concomitant reduction in single dose study. The magnitude and levodopa-induced dyskinesias. More recently, duration of the therapeutic response to a a new generation of COMT inhibitors have single dose of 200 mg levodopa/50 mg been developed, which are much more potent carbidopa was evaluated after concomi- and specific."'3 1620 Fluorodopa positron tant placebo, or 200 or 800 mg enta- emission tomography (PET) studies in mon- capone.